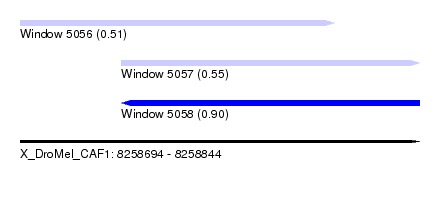
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 8,258,694 – 8,258,844 |
| Length | 150 |
| Max. P | 0.901541 |

| Location | 8,258,694 – 8,258,812 |
|---|---|
| Length | 118 |
| Sequences | 3 |
| Columns | 118 |
| Reading direction | forward |
| Mean pairwise identity | 96.05 |
| Mean single sequence MFE | -26.40 |
| Consensus MFE | -23.91 |
| Energy contribution | -23.47 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.68 |
| Structure conservation index | 0.91 |
| SVM decision value | -0.04 |
| SVM RNA-class probability | 0.511200 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 8258694 118 + 22224390 CUUCCUUUGCCUUUAAAUUAAGCCCAUUAAAAUUAAUUAAGAUAUUGACCCAGUUCGCUGGCAGUCGUCGGAAAAGCGUCGGCACAGAAACAAAAUUGUUUCAAGUGCCUGACAUUUU .((((..........((((((...........))))))..(((((((..((((....)))))))).)))))))....((((((((.((((((....))))))..))))).)))..... ( -29.30) >DroSec_CAF1 8956 118 + 1 CUUCCUUUGCCUUUAAAUUAAGCCCAUUAAAAUUAAUUAAGAUAUUGACCCAGUUCGCUGGCAGUCGUCGGAAAACCGCCGGUACAGAAAUAAAAUUGUUUCAAGUGCCUGACAUUUU .((((..........((((((...........))))))..(((((((..((((....)))))))).))))))).......(((((.((((((....))))))..)))))......... ( -21.70) >DroYak_CAF1 13550 118 + 1 CUUCCUUUGUCUUUAAAUUAAGCCCAUUAAAAUUAAUUAAGAUAUUGACCCAGUUCGCUGGCAGUCUACGGAAAACCGCCGGCACAGAAACAAAAUUGUUUCAAGUGCCUGACAUUUU .((((..((((((..((((((...........))))))))))))..(((((((....))))..)))...)))).......(((((.((((((....))))))..)))))......... ( -28.20) >consensus CUUCCUUUGCCUUUAAAUUAAGCCCAUUAAAAUUAAUUAAGAUAUUGACCCAGUUCGCUGGCAGUCGUCGGAAAACCGCCGGCACAGAAACAAAAUUGUUUCAAGUGCCUGACAUUUU .((((.....(((..((((((...........))))))))).....(((((((....))))..)))...)))).......(((((.((((((....))))))..)))))......... (-23.91 = -23.47 + -0.44)

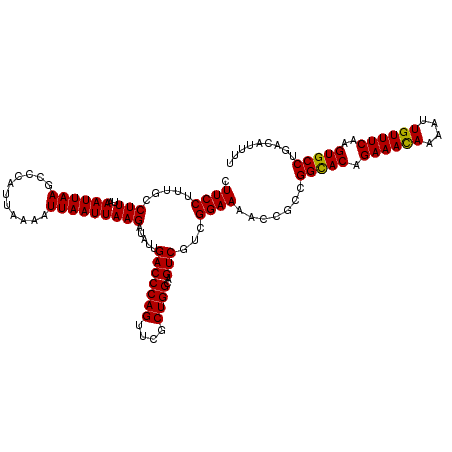
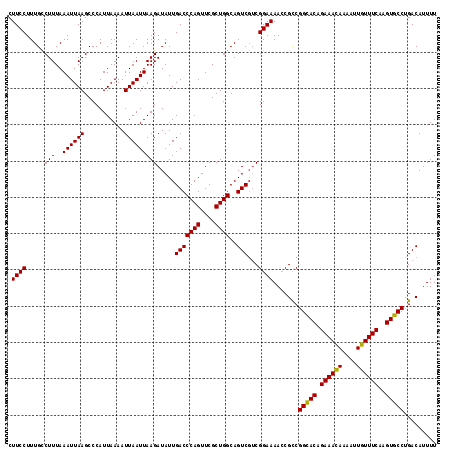
| Location | 8,258,732 – 8,258,844 |
|---|---|
| Length | 112 |
| Sequences | 3 |
| Columns | 112 |
| Reading direction | forward |
| Mean pairwise identity | 95.83 |
| Mean single sequence MFE | -28.67 |
| Consensus MFE | -25.51 |
| Energy contribution | -25.73 |
| Covariance contribution | 0.23 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.80 |
| Structure conservation index | 0.89 |
| SVM decision value | 0.03 |
| SVM RNA-class probability | 0.550455 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 8258732 112 + 22224390 AAGAUAUUGACCCAGUUCGCUGGCAGUCGUCGGAAAAGCGUCGGCACAGAAACAAAAUUGUUUCAAGUGCCUGACAUUUUUACAUUAAAGCGACAUGACGUCAUUAGAUGAA ....((.((((((((....))))..(((((..((((((.((((((((.((((((....))))))..))))).))).))))).)......))))).....)))).))...... ( -32.00) >DroSec_CAF1 8994 112 + 1 AAGAUAUUGACCCAGUUCGCUGGCAGUCGUCGGAAAACCGCCGGUACAGAAAUAAAAUUGUUUCAAGUGCCUGACAUUUUUACAUUAAAGCGACAUGACGUCAUUAGAUGAA ....((.((((((((....))))..((((((((....)))..(((((.((((((....))))))..)))))..................))))).....)))).))...... ( -26.50) >DroYak_CAF1 13588 112 + 1 AAGAUAUUGACCCAGUUCGCUGGCAGUCUACGGAAAACCGCCGGCACAGAAACAAAAUUGUUUCAAGUGCCUGACAUUUUUACAUUAAAACGACAUGACGUCAUUAGAUGAA ....((.((((((((....))))..(((..(((....)))..(((((.((((((....))))))..)))))....................))).....)))).))...... ( -27.50) >consensus AAGAUAUUGACCCAGUUCGCUGGCAGUCGUCGGAAAACCGCCGGCACAGAAACAAAAUUGUUUCAAGUGCCUGACAUUUUUACAUUAAAGCGACAUGACGUCAUUAGAUGAA ....((.((((((((....))))..(((((((......))..(((((.((((((....))))))..)))))..................))))).....)))).))...... (-25.51 = -25.73 + 0.23)

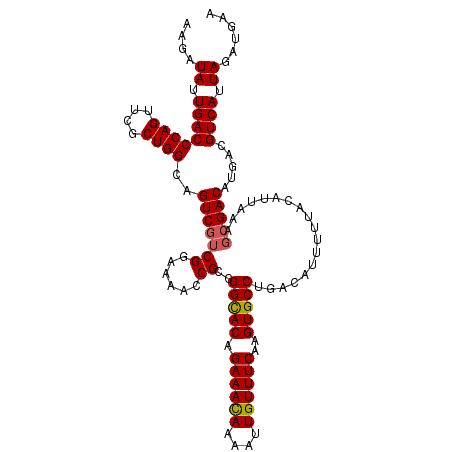
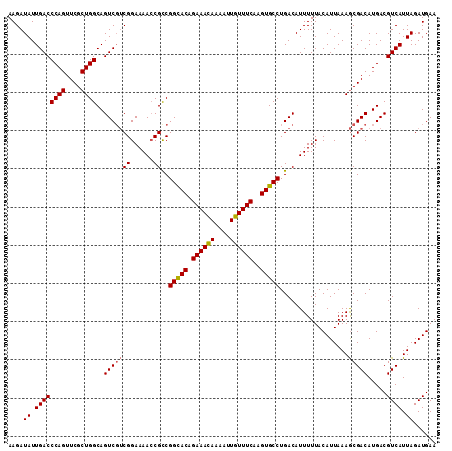
| Location | 8,258,732 – 8,258,844 |
|---|---|
| Length | 112 |
| Sequences | 3 |
| Columns | 112 |
| Reading direction | reverse |
| Mean pairwise identity | 95.83 |
| Mean single sequence MFE | -31.70 |
| Consensus MFE | -28.72 |
| Energy contribution | -29.83 |
| Covariance contribution | 1.11 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.45 |
| Structure conservation index | 0.91 |
| SVM decision value | 1.02 |
| SVM RNA-class probability | 0.901541 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 8258732 112 - 22224390 UUCAUCUAAUGACGUCAUGUCGCUUUAAUGUAAAAAUGUCAGGCACUUGAAACAAUUUUGUUUCUGUGCCGACGCUUUUCCGACGACUGCCAGCGAACUGGGUCAAUAUCUU .........(((((((..(((((......))((((.((((.(((((..((((((....)))))).))))))))).))))..))))))..((((....))))))))....... ( -32.20) >DroSec_CAF1 8994 112 - 1 UUCAUCUAAUGACGUCAUGUCGCUUUAAUGUAAAAAUGUCAGGCACUUGAAACAAUUUUAUUUCUGUACCGGCGGUUUUCCGACGACUGCCAGCGAACUGGGUCAAUAUCUU .........((((.(((..(((((.................((.((..((((........)))).)).))(((((((.......))))))))))))..)))))))....... ( -26.30) >DroYak_CAF1 13588 112 - 1 UUCAUCUAAUGACGUCAUGUCGUUUUAAUGUAAAAAUGUCAGGCACUUGAAACAAUUUUGUUUCUGUGCCGGCGGUUUUCCGUAGACUGCCAGCGAACUGGGUCAAUAUCUU .........((((.(((..(((((.................(((((..((((((....)))))).)))))((((((((.....)))))))))))))..)))))))....... ( -36.60) >consensus UUCAUCUAAUGACGUCAUGUCGCUUUAAUGUAAAAAUGUCAGGCACUUGAAACAAUUUUGUUUCUGUGCCGGCGGUUUUCCGACGACUGCCAGCGAACUGGGUCAAUAUCUU .........((((.(((..(((((.................(((((..((((((....)))))).)))))(((((((.......))))))))))))..)))))))....... (-28.72 = -29.83 + 1.11)

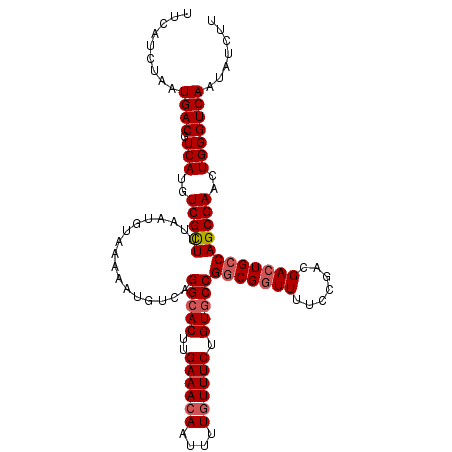
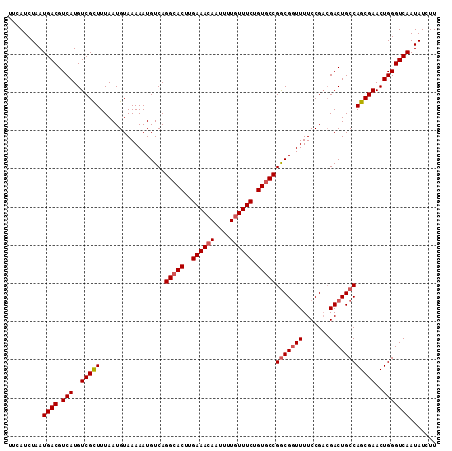
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:50:46 2006