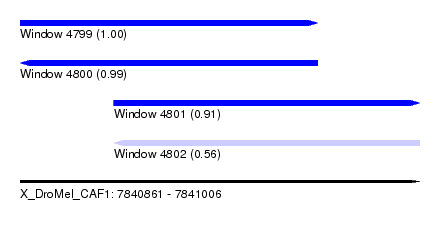

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 7,840,861 – 7,841,006 |
| Length | 145 |
| Max. P | 0.998392 |

| Location | 7,840,861 – 7,840,969 |
|---|---|
| Length | 108 |
| Sequences | 3 |
| Columns | 113 |
| Reading direction | forward |
| Mean pairwise identity | 73.65 |
| Mean single sequence MFE | -31.30 |
| Consensus MFE | -20.40 |
| Energy contribution | -21.07 |
| Covariance contribution | 0.67 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.26 |
| Structure conservation index | 0.65 |
| SVM decision value | 3.09 |
| SVM RNA-class probability | 0.998392 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 7840861 108 + 22224390 CGUGCUAUUCGUUACGCGAUUUCUCUGACGAAAGCGUAGAAGCGCGCCAAAAAAAGCGCGCGCAAAAACAAAAAGCCAA---GCUAACGGGACAAAAGUCGCGCGCGACUC-- (((((...((((((.((...((((.((.(....))).))))((((((........))))))..................---))))))))(((....)))..)))))....-- ( -34.80) >DroEre_CAF1 26275 110 + 1 CCUGCUCUCCGUCACGCGAUUUCCCUGCCGAAAGCGUAGAAACGCGCCAAAAAAAGCGCGCGCAAAAACAAAAAGCCAA---GCUAACGGGACAAAAGUCGCGCGCGACUCGC ..........(((.((((.((((..((((....).)))))))))))...........((((((............((..---......))(((....)))))))))))).... ( -31.80) >DroAna_CAF1 13030 90 + 1 -CGGCUCUCCA--ACGCG-UCUCAUUGU------------UUGGCGCGGUAAAAAGCGCGCGCAAAAACAAAAAGCCAACUAACUAACGGGACAAAAGUCGCGCGC------- -..(((.....--.((((-((.......------------..))))))......)))((((((............((...........))(((....)))))))))------- ( -27.30) >consensus CCUGCUCUCCGU_ACGCGAUUUCACUGACGAAAGCGUAGAAACGCGCCAAAAAAAGCGCGCGCAAAAACAAAAAGCCAA___GCUAACGGGACAAAAGUCGCGCGCGACUC__ ..............(((..........................((((........))))((((..........(((......))).....(((....))))))))))...... (-20.40 = -21.07 + 0.67)

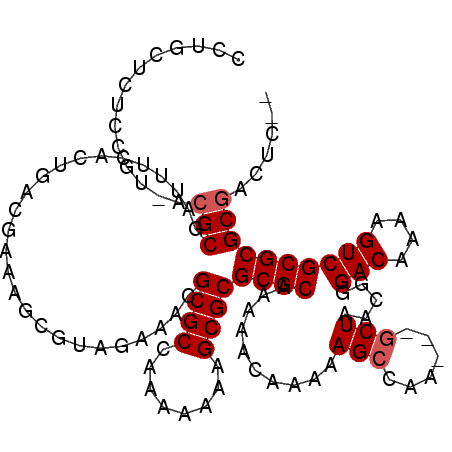
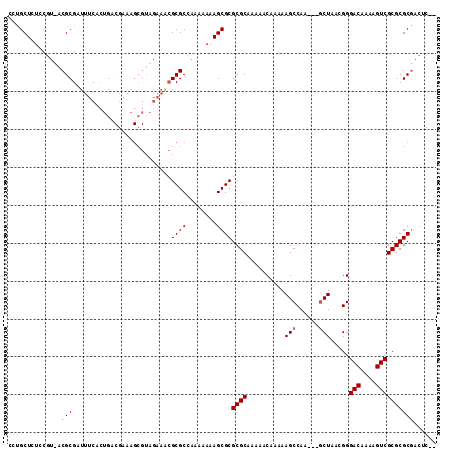
| Location | 7,840,861 – 7,840,969 |
|---|---|
| Length | 108 |
| Sequences | 3 |
| Columns | 113 |
| Reading direction | reverse |
| Mean pairwise identity | 73.65 |
| Mean single sequence MFE | -36.67 |
| Consensus MFE | -27.01 |
| Energy contribution | -27.46 |
| Covariance contribution | 0.45 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.71 |
| Structure conservation index | 0.74 |
| SVM decision value | 2.02 |
| SVM RNA-class probability | 0.985905 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 7840861 108 - 22224390 --GAGUCGCGCGCGACUUUUGUCCCGUUAGC---UUGGCUUUUUGUUUUUGCGCGCGCUUUUUUUGGCGCGCUUCUACGCUUUCGUCAGAGAAAUCGCGUAACGAAUAGCACG --((((...(((.(((....))).)))..))---)).(((.((((((..((((((((((......)))))))(((((((....))).)))).....))).)))))).)))... ( -36.20) >DroEre_CAF1 26275 110 - 1 GCGAGUCGCGCGCGACUUUUGUCCCGUUAGC---UUGGCUUUUUGUUUUUGCGCGCGCUUUUUUUGGCGCGUUUCUACGCUUUCGGCAGGGAAAUCGCGUGACGGAGAGCAGG ((...((((((((((.((((.(.(((..(((---...((...........))(((((((......)))))))......)))..))).).)))).))))))).)))...))... ( -42.70) >DroAna_CAF1 13030 90 - 1 -------GCGCGCGACUUUUGUCCCGUUAGUUAGUUGGCUUUUUGUUUUUGCGCGCGCUUUUUACCGCGCCAA------------ACAAUGAGA-CGCGU--UGGAGAGCCG- -------(((((((((....)))..(((((....)))))...........))))))((((((((.(((((((.------------....)).).-)))).--))))))))..- ( -31.10) >consensus __GAGUCGCGCGCGACUUUUGUCCCGUUAGC___UUGGCUUUUUGUUUUUGCGCGCGCUUUUUUUGGCGCGAUUCUACGCUUUCGACAGAGAAAUCGCGU_ACGGAGAGCAGG .......(((((((((....)))..(((((....)))))...........))))))(((((((...(((((........................)))))...)))))))... (-27.01 = -27.46 + 0.45)

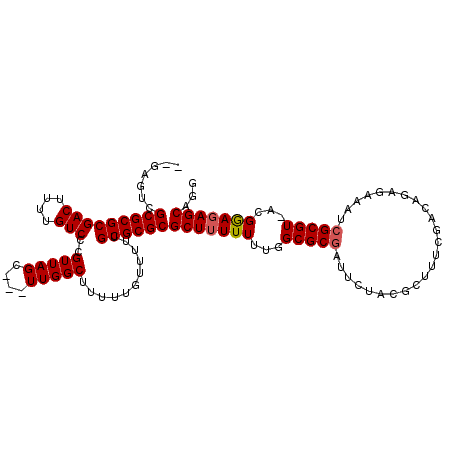
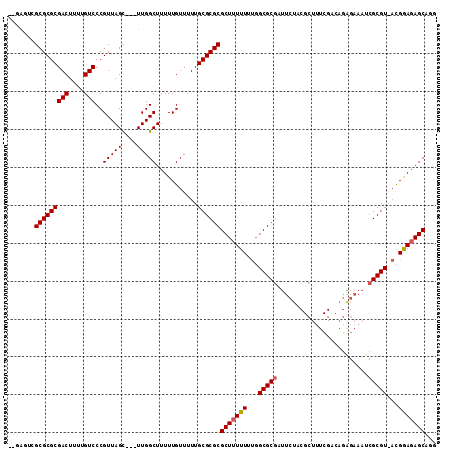
| Location | 7,840,895 – 7,841,006 |
|---|---|
| Length | 111 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 88.02 |
| Mean single sequence MFE | -33.58 |
| Consensus MFE | -25.42 |
| Energy contribution | -26.17 |
| Covariance contribution | 0.75 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.33 |
| Structure conservation index | 0.76 |
| SVM decision value | 1.10 |
| SVM RNA-class probability | 0.914519 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 7840895 111 + 22224390 CGUAGAAGCGCGCCAAA-AAAAGCGCGCGCAAAAACAAAAAGCCAA---GCUAACGGGACAAAAGUCGCGCGCGACUC--GUAAGCUUGUUUACUUUGCCAUUAUAAAUCC---GGCGGG .(((((.((((((....-....))))))...............(((---(((.(((.......(((((....))))))--)).))))))))))).(((((...........---))))). ( -34.31) >DroEre_CAF1 26309 113 + 1 CGUAGAAACGCGCCAAA-AAAAGCGCGCGCAAAAACAAAAAGCCAA---GCUAACGGGACAAAAGUCGCGCGCGACUCGCGUAAGCUUGUUUACUUUGCCAUUAUAAAUCC---GGCGGG .(((((..(((((....-....)))))................(((---(((.(((.((.....((((....)))))).))).))))))))))).(((((...........---))))). ( -32.50) >DroYak_CAF1 21601 112 + 1 CGUAGAAGCGCGCCAAAAAAAAGCGCGCGCAAAAACAAAAAGCCAA---GCUAACGGGACAAAAGUCGCGCGCGACUC--GUAAGCUUGUUUACUUUGCCAUUAUAAAUCC---GGCGGG .(((((.(((((((........).)))))).............(((---(((.(((.......(((((....))))))--)).))))))))))).(((((...........---))))). ( -34.31) >DroAna_CAF1 13054 106 + 1 ------UUGGCGCGGUA-AAAAGCGCGCGCAAAAACAAAAAGCCAACUAACUAACGGGACAAAAGUCGCGCGC-------GUAAGCUUGUUUACUUUGCCAUUAUAAAUCCGACGGCGGG ------.(((((.((((-((((((((((((............((...........))(((....)))))))))-------....)))).)))))).)))))........(((....))). ( -33.20) >consensus CGUAGAAGCGCGCCAAA_AAAAGCGCGCGCAAAAACAAAAAGCCAA___GCUAACGGGACAAAAGUCGCGCGCGACUC__GUAAGCUUGUUUACUUUGCCAUUAUAAAUCC___GGCGGG ..........((((.........(((((((..........(((......))).....(((....))))))))))......(((((....)))))....................)))).. (-25.42 = -26.17 + 0.75)

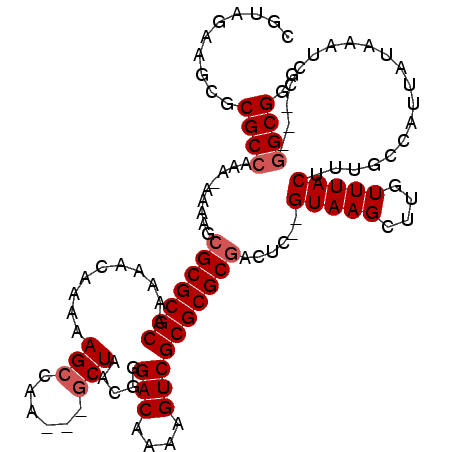
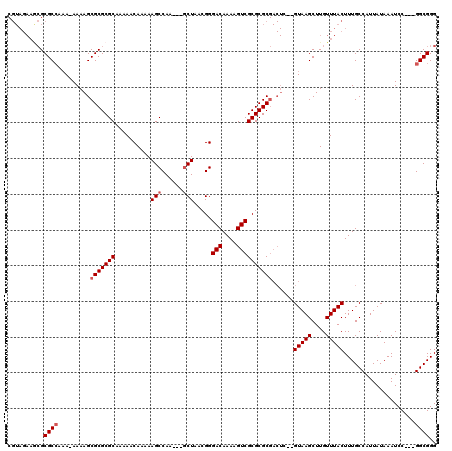
| Location | 7,840,895 – 7,841,006 |
|---|---|
| Length | 111 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 88.02 |
| Mean single sequence MFE | -34.16 |
| Consensus MFE | -25.64 |
| Energy contribution | -26.26 |
| Covariance contribution | 0.62 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.46 |
| Structure conservation index | 0.75 |
| SVM decision value | 0.04 |
| SVM RNA-class probability | 0.555805 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 7840895 111 - 22224390 CCCGCC---GGAUUUAUAAUGGCAAAGUAAACAAGCUUAC--GAGUCGCGCGCGACUUUUGUCCCGUUAGC---UUGGCUUUUUGUUUUUGCGCGCGCUUUU-UUUGGCGCGCUUCUACG ...(((---(.........))))...(((((((((((.((--((((((....))))).......))).)))---)))((.....)).)))))(((((((...-...)))))))....... ( -36.31) >DroEre_CAF1 26309 113 - 1 CCCGCC---GGAUUUAUAAUGGCAAAGUAAACAAGCUUACGCGAGUCGCGCGCGACUUUUGUCCCGUUAGC---UUGGCUUUUUGUUUUUGCGCGCGCUUUU-UUUGGCGCGUUUCUACG ...(((---(.........))))...(((((((((((.(((.((((((....))))))......))).)))---)))((.....)).)))))(((((((...-...)))))))....... ( -34.80) >DroYak_CAF1 21601 112 - 1 CCCGCC---GGAUUUAUAAUGGCAAAGUAAACAAGCUUAC--GAGUCGCGCGCGACUUUUGUCCCGUUAGC---UUGGCUUUUUGUUUUUGCGCGCGCUUUUUUUUGGCGCGCUUCUACG ...(((---(.........))))...(((((((((((.((--((((((....))))).......))).)))---)))((.....)).)))))(((((((.......)))))))....... ( -35.81) >DroAna_CAF1 13054 106 - 1 CCCGCCGUCGGAUUUAUAAUGGCAAAGUAAACAAGCUUAC-------GCGCGCGACUUUUGUCCCGUUAGUUAGUUGGCUUUUUGUUUUUGCGCGCGCUUUU-UACCGCGCCAA------ .(((....)))........((((...(((((.((((....-------(((((((((....)))..(((((....)))))...........))))))))))))-)))...)))).------ ( -29.70) >consensus CCCGCC___GGAUUUAUAAUGGCAAAGUAAACAAGCUUAC__GAGUCGCGCGCGACUUUUGUCCCGUUAGC___UUGGCUUUUUGUUUUUGCGCGCGCUUUU_UUUGGCGCGCUUCUACG .((......))..........((((((..((.(((((..........(((.(.(((....))))))).........))))).))..))))))(((((((.......)))))))....... (-25.64 = -26.26 + 0.62)

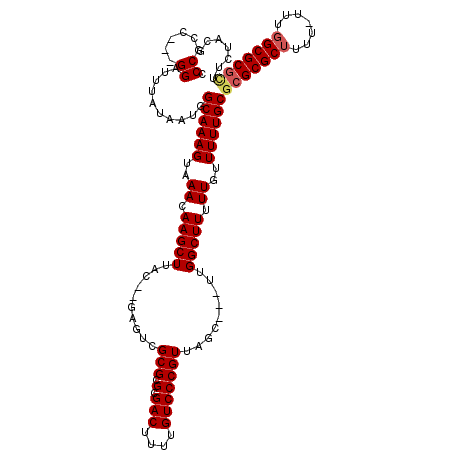
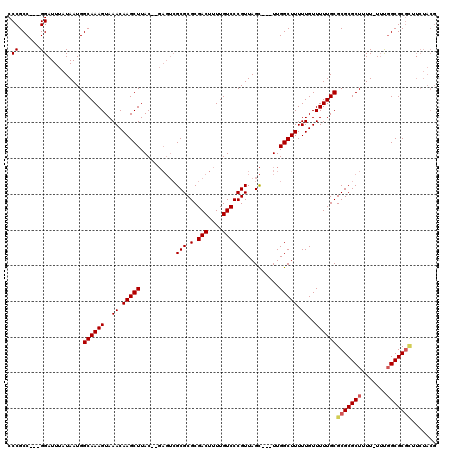
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:46:56 2006