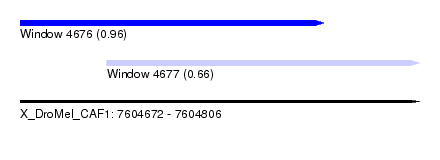

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 7,604,672 – 7,604,806 |
| Length | 134 |
| Max. P | 0.962934 |

| Location | 7,604,672 – 7,604,774 |
|---|---|
| Length | 102 |
| Sequences | 4 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 92.48 |
| Mean single sequence MFE | -36.52 |
| Consensus MFE | -29.92 |
| Energy contribution | -31.54 |
| Covariance contribution | 1.62 |
| Combinations/Pair | 1.07 |
| Mean z-score | -3.31 |
| Structure conservation index | 0.82 |
| SVM decision value | 1.55 |
| SVM RNA-class probability | 0.962934 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
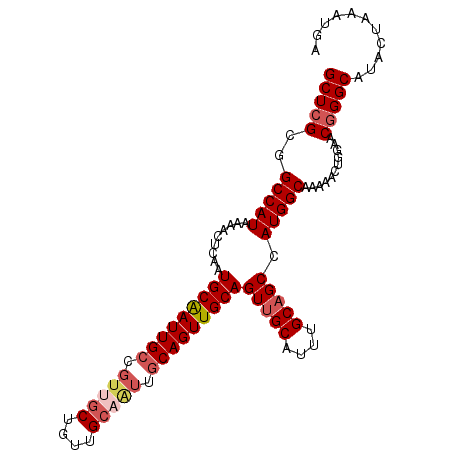
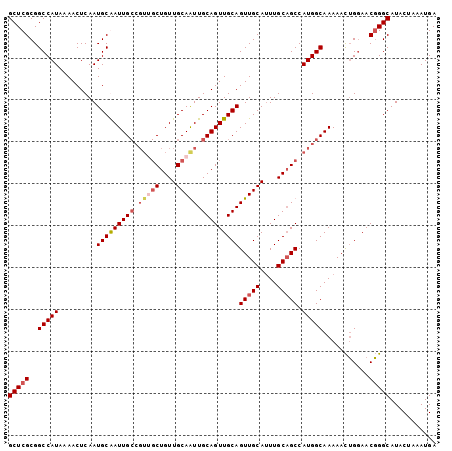
>X_DroMel_CAF1 7604672 102 + 22224390 GCUCGCGGCCAUAAAACUCAAUGCAAUUGCCGUCGCUGUUGCAGUUGCAGUUGCAGUUGCAUUUGCAGCCAUGGCAAAAACUGGAACGGGCAUACUAAAUGA (((((...(((.....(.....)...(((((((.(((((((((..(((....)))..))))...))))).)))))))....)))..)))))........... ( -35.50) >DroSec_CAF1 21711 102 + 1 GCUCGCGGCCAUAAAACUCAAUGCGAUUGCCGUUGCUGUUGCAAUUGCAGUUGCAGUUGCAUUUGCAGCCAUGGCAAAAACUGGAACGGGCAUACUAAAUGA (((((...(((......((.....))(((((((.((((((((((((((....)))))))))...))))).)))))))....)))..)))))........... ( -39.50) >DroSim_CAF1 22516 102 + 1 GCUCGCGGCCAUAAAACUCAAUGCGAUUGCCGUUGCUGUUGCAAUUGCAGUUGCAGUUGCAUUUGCAGCCAUGGCAAAAACUGGAACGGGCAUACUAAAUGA (((((...(((......((.....))(((((((.((((((((((((((....)))))))))...))))).)))))))....)))..)))))........... ( -39.50) >DroYak_CAF1 23128 96 + 1 GCUCGCGGCCAUAAAACUCAAUGCAAUUG------CUGUUGCAGUUGCAGUUGCAGUUGCAGUUGCCGCCAUGGCAAAAAUGGAAACUGGCAUACUAAAUGA (((.(...((((.........((((((((------(..((((....))))..))))))))).(((((.....)))))..))))...).)))........... ( -31.60) >consensus GCUCGCGGCCAUAAAACUCAAUGCAAUUGCCGUUGCUGUUGCAAUUGCAGUUGCAGUUGCAUUUGCAGCCAUGGCAAAAACUGGAACGGGCAUACUAAAUGA (((((..(((((.........(((((((((.(((((....))))).)))))))))(((((....))))).)))))...........)))))........... (-29.92 = -31.54 + 1.62)

| Location | 7,604,701 – 7,604,806 |
|---|---|
| Length | 105 |
| Sequences | 4 |
| Columns | 113 |
| Reading direction | forward |
| Mean pairwise identity | 88.53 |
| Mean single sequence MFE | -30.04 |
| Consensus MFE | -21.48 |
| Energy contribution | -23.10 |
| Covariance contribution | 1.62 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.78 |
| Structure conservation index | 0.71 |
| SVM decision value | 0.26 |
| SVM RNA-class probability | 0.657834 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
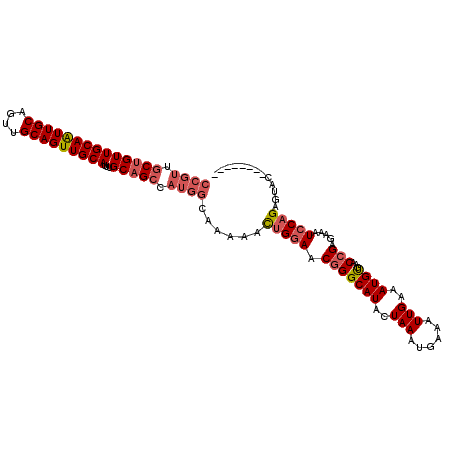
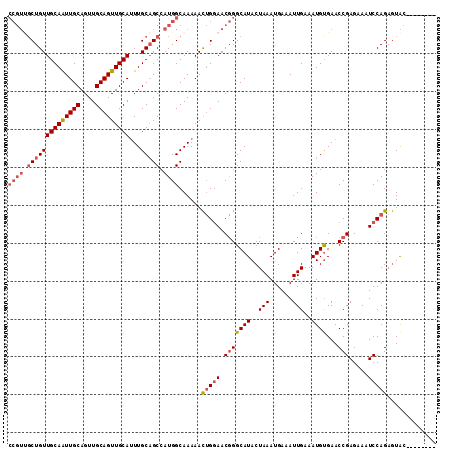
>X_DroMel_CAF1 7604701 105 + 22224390 CCGUCGCUGUUGCAGUUGCAGUUGCAGUUGCAUUUGCAGCCAUGGCAAAAACUGGAACGGGCAUACUAAAUGAAAUUGAAAUGCGAACCGAGAAAUCCAGAGUAC-------- ((((.(((((((((..(((....)))..))))...))))).))))......(((((.(((((((..(((......)))..))))...))).....))))).....-------- ( -30.80) >DroSec_CAF1 21740 105 + 1 CCGUUGCUGUUGCAAUUGCAGUUGCAGUUGCAUUUGCAGCCAUGGCAAAAACUGGAACGGGCAUACUAAAUGAAAUUGAAAUGUGAACCGAGAAAUCCAGAGUAC-------- ((((.((((((((((((((....)))))))))...))))).))))......(((((.(((...(((................)))..))).....))))).....-------- ( -32.29) >DroSim_CAF1 22545 105 + 1 CCGUUGCUGUUGCAAUUGCAGUUGCAGUUGCAUUUGCAGCCAUGGCAAAAACUGGAACGGGCAUACUAAAUGAAAUUGAAAUGUGAACCGAGAAAUCCAGAGUAC-------- ((((.((((((((((((((....)))))))))...))))).))))......(((((.(((...(((................)))..))).....))))).....-------- ( -32.29) >DroYak_CAF1 23157 107 + 1 ------CUGUUGCAGUUGCAGUUGCAGUUGCAGUUGCCGCCAUGGCAAAAAUGGAAACUGGCAUACUAAAUGAAAUUGAAAUGUGAACCGAGAAAUCCAGAAUGCCUCCCCCC ------...(((((..(((....)))..)))))(((((.....)))))....(((....(((((.((...............(....).((....)).)).)))))))).... ( -24.80) >consensus CCGUUGCUGUUGCAAUUGCAGUUGCAGUUGCAUUUGCAGCCAUGGCAAAAACUGGAACGGGCAUACUAAAUGAAAUUGAAAUGUGAACCGAGAAAUCCAGAGUAC________ ((((.((((((((((((((....)))))))))...))))).))))......(((((.(((((((..(((......)))..))))...))).....)))))............. (-21.48 = -23.10 + 1.62)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:45:03 2006