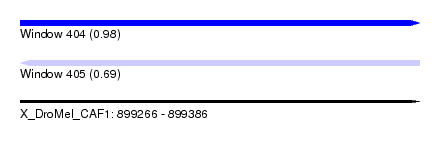

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 899,266 – 899,386 |
| Length | 120 |
| Max. P | 0.978079 |

| Location | 899,266 – 899,386 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 85.14 |
| Mean single sequence MFE | -39.52 |
| Consensus MFE | -31.19 |
| Energy contribution | -32.07 |
| Covariance contribution | 0.87 |
| Combinations/Pair | 1.19 |
| Mean z-score | -2.57 |
| Structure conservation index | 0.79 |
| SVM decision value | 1.81 |
| SVM RNA-class probability | 0.978079 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 899266 120 + 22224390 UGGUCUGGUAGUACCUCCAGAUCCAAAGAACCGGGCAGGGGUGUCGAUAUACUUUACUUGGUUUCGAAACUCCAGCAAUCUCUUCUUCGAAUGUAAAGACGAAGAGAAGAGAUUUUUGGA ((((((((........))))).))).....((((((.(((((.((((...(((......))).)))).))))).))(((((((((((((..(....)..)))..)))))))))).)))). ( -40.40) >DroSec_CAF1 14289 120 + 1 UGGUCUGGUAGUACCUCCAGAUACAAACAUCCGGGCAGGGGUGUCGAUAUACUUUACUUGGUUUCGAAAUUCGAGCGAUCUCUUCUUCGAAUGUGAAAAAGAAGAGAAGAGAUUUCUGGA ..((((((........)))))).......(((((((..((((.((((...(((......))).)))).))))..))(((((((((((......(....)....))))))))))).))))) ( -35.10) >DroSim_CAF1 14419 120 + 1 UGGUCUGGUAGUACCUCCAGAUACAAACAUCCGGGCUGGGGUGUCGAUAUACUUUACAUGGUUUCGAAAUUCGAGCGAUCUCUUCUUCGAAUGUGAAAAAGAAGAGAAGAGAUUUCUGGA ..((((((........)))))).......((((((((.((((.((((...(((......))).)))).)))).)))(((((((((((......(....)....))))))))))).))))) ( -39.20) >DroEre_CAF1 8294 114 + 1 UGGGCUGGUAGUACCUUCAGAUCCAACGAUCCGGGCAGGGGUGUCGAUAUUCUUGAUUUGGUGUCGAGACCCCAGGGAUCUCUGUUCUGAAUGUGG------AUGGAAGAGAUUUCUGGA ((((((((........)))).))))....(((.....(((((.((((((((........)))))))).)))))((..((((((..(((......))------)....))))))..))))) ( -43.40) >consensus UGGUCUGGUAGUACCUCCAGAUACAAACAUCCGGGCAGGGGUGUCGAUAUACUUUACUUGGUUUCGAAACUCCAGCGAUCUCUUCUUCGAAUGUGAAAAAGAAGAGAAGAGAUUUCUGGA .(((((((........))))))).......((((((.(((((.((((...(((......))).)))).))))).))(((((((((((................))))))))))).)))). (-31.19 = -32.07 + 0.87)

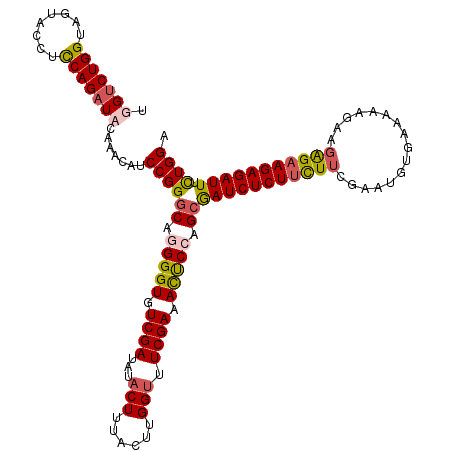
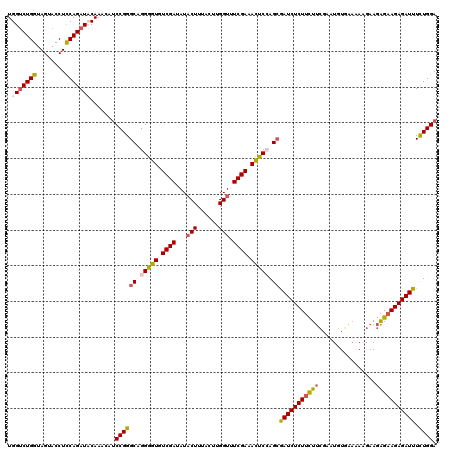
| Location | 899,266 – 899,386 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 85.14 |
| Mean single sequence MFE | -29.00 |
| Consensus MFE | -20.05 |
| Energy contribution | -21.36 |
| Covariance contribution | 1.31 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.76 |
| Structure conservation index | 0.69 |
| SVM decision value | 0.33 |
| SVM RNA-class probability | 0.692978 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 899266 120 - 22224390 UCCAAAAAUCUCUUCUCUUCGUCUUUACAUUCGAAGAAGAGAUUGCUGGAGUUUCGAAACCAAGUAAAGUAUAUCGACACCCCUGCCCGGUUCUUUGGAUCUGGAGGUACUACCAGACCA ((((((((((((((((..(((..........)))))))))))))((.((.((.((((.((........))...)))).)).)).)).......))))))(((((........)))))... ( -32.10) >DroSec_CAF1 14289 120 - 1 UCCAGAAAUCUCUUCUCUUCUUUUUCACAUUCGAAGAAGAGAUCGCUCGAAUUUCGAAACCAAGUAAAGUAUAUCGACACCCCUGCCCGGAUGUUUGUAUCUGGAGGUACUACCAGACCA ((((((.......((((((((((.........))))))))))....(((.....)))..............(((.((((.((......)).)))).)))))))))(((........))). ( -27.60) >DroSim_CAF1 14419 120 - 1 UCCAGAAAUCUCUUCUCUUCUUUUUCACAUUCGAAGAAGAGAUCGCUCGAAUUUCGAAACCAUGUAAAGUAUAUCGACACCCCAGCCCGGAUGUUUGUAUCUGGAGGUACUACCAGACCA ((((((.......((((((((((.........))))))))))....(((.....)))..............(((.((((.((......)).)))).)))))))))(((........))). ( -27.60) >DroEre_CAF1 8294 114 - 1 UCCAGAAAUCUCUUCCAU------CCACAUUCAGAACAGAGAUCCCUGGGGUCUCGACACCAAAUCAAGAAUAUCGACACCCCUGCCCGGAUCGUUGGAUCUGAAGGUACUACCAGCCCA ..................------.....((((((.(((.(((((..(((((.((((................)))).))))).....))))).)))..))))))((.....))...... ( -28.69) >consensus UCCAGAAAUCUCUUCUCUUCUUUUUCACAUUCGAAGAAGAGAUCGCUCGAAUUUCGAAACCAAGUAAAGUAUAUCGACACCCCUGCCCGGAUCUUUGGAUCUGGAGGUACUACCAGACCA (((((..(((((((((..................)))))))))..)))))...((((................)))).........(((((((....))))))).((.....))...... (-20.05 = -21.36 + 1.31)

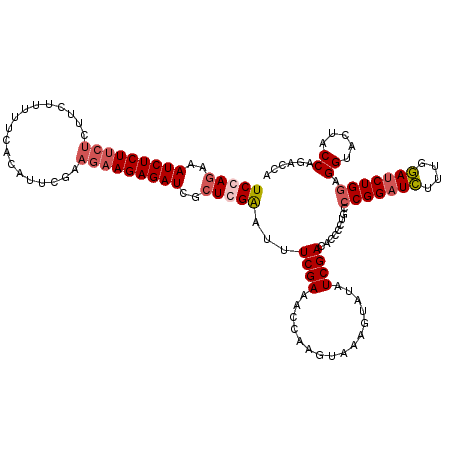
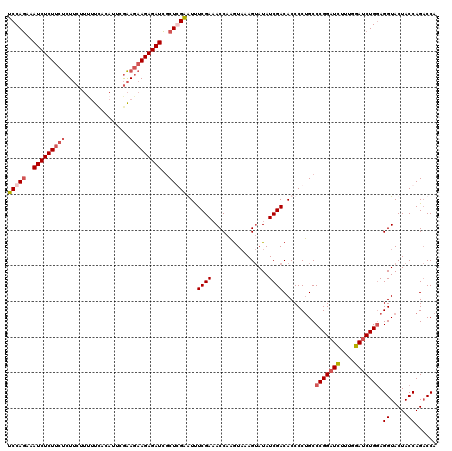
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:40:23 2006