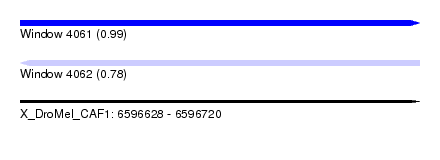

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 6,596,628 – 6,596,720 |
| Length | 92 |
| Max. P | 0.988385 |

| Location | 6,596,628 – 6,596,720 |
|---|---|
| Length | 92 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 65.12 |
| Mean single sequence MFE | -33.48 |
| Consensus MFE | -17.06 |
| Energy contribution | -17.05 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.15 |
| Structure conservation index | 0.51 |
| SVM decision value | 2.12 |
| SVM RNA-class probability | 0.988385 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 6596628 92 + 22224390 UC---------CUCCCCACUCGAUUCGAGUUUUGUGUUUCGAG-------UUGAGUUUCGAGUUU------------GCGAUCGAGUUGGAGGCAACCAAGCCUGACCGUUCUUGGUCUA ..---------(((...((((((((((((........))))))-------))))))...)))...------------..(((((((.(((((((......))))..)))..))))))).. ( -33.30) >DroSim_CAF1 8717 87 + 1 UC--------------CACUCGAUUCGAGUUUUGAGUUUCGAG-------UUGAGUUUCGAGUUU------------GAGUUCGGGUUGGAGGCAGCCAAGCCUGACCGUUCUUGGUCUA ..--------------.((((((((((((........))))))-------)))))).(((((..(------------(.(((.((.((((......)))).)).)))))..))))).... ( -31.00) >DroEre_CAF1 12156 120 + 1 UCAACCCUCAACUGUCCACUCGAUUCGAGUUCUGAGUCUGAGUCUGGGUCUGGAGUCUCGACUCCGACUCCGAGUUCGAGUUCCAGCGCGAGGCAGCCAAGCCUGACCGUUCUUGGUCCA ...(((...(((.(((.....(((((((..((.(((((.(((((.(((.(....).)))))))).))))).))..)))))))........((((......))))))).)))...)))... ( -41.00) >DroYak_CAF1 11499 97 + 1 UCAACUCUCGACUGUCCACUUGAUUCGAGUU-----------UCAAAGUUUGGAGUCUCGAUUCU------------AAGCUCUAGCUCGAGGCAGCCAAGCCUGACCGUUCUUGGUCUA ........................(((((((-----------....((((((((((....)))))------------)))))..)))))))(((......))).(((((....))))).. ( -28.60) >consensus UC_________CUGUCCACUCGAUUCGAGUUUUGAGUUUCGAG_______UGGAGUCUCGAGUCU____________GAGUUCGAGCUCGAGGCAGCCAAGCCUGACCGUUCUUGGUCUA .....................((((((((...........................))))))))..........................((((......))))(((((....))))).. (-17.06 = -17.05 + -0.00)

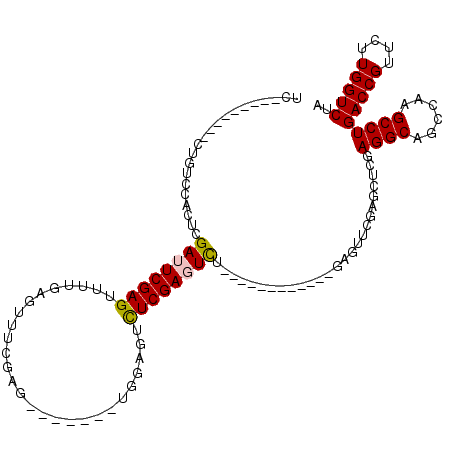
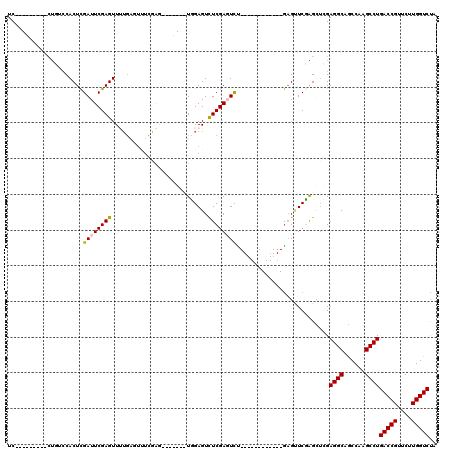
| Location | 6,596,628 – 6,596,720 |
|---|---|
| Length | 92 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 65.12 |
| Mean single sequence MFE | -30.62 |
| Consensus MFE | -13.60 |
| Energy contribution | -13.85 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.92 |
| Structure conservation index | 0.44 |
| SVM decision value | 0.56 |
| SVM RNA-class probability | 0.781108 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 6596628 92 - 22224390 UAGACCAAGAACGGUCAGGCUUGGUUGCCUCCAACUCGAUCGC------------AAACUCGAAACUCAA-------CUCGAAACACAAAACUCGAAUCGAGUGGGGAG---------GA ..(((((((..(.....).))))))).(((((.(((((((((.------------.....))).......-------.((((..........)))).))))))..))))---------). ( -27.90) >DroSim_CAF1 8717 87 - 1 UAGACCAAGAACGGUCAGGCUUGGCUGCCUCCAACCCGAACUC------------AAACUCGAAACUCAA-------CUCGAAACUCAAAACUCGAAUCGAGUG--------------GA ..((((......)))).(((......)))(((....(((....------------....))).......(-------(((((...((.......)).)))))))--------------)) ( -20.00) >DroEre_CAF1 12156 120 - 1 UGGACCAAGAACGGUCAGGCUUGGCUGCCUCGCGCUGGAACUCGAACUCGGAGUCGGAGUCGAGACUCCAGACCCAGACUCAGACUCAGAACUCGAAUCGAGUGGACAGUUGAGGGUUGA ..((((......))))...((..((((..((.((((.((..((((..((.(((((.(((((...............))))).))))).))..)))).)).))))))))))..))...... ( -42.96) >DroYak_CAF1 11499 97 - 1 UAGACCAAGAACGGUCAGGCUUGGCUGCCUCGAGCUAGAGCUU------------AGAAUCGAGACUCCAAACUUUGA-----------AACUCGAAUCAAGUGGACAGUCGAGAGUUGA ..((((......))))...((((((((((.(((((....))))------------.((.(((((..((........))-----------..))))).))..).)).))))))))...... ( -31.60) >consensus UAGACCAAGAACGGUCAGGCUUGGCUGCCUCCAACUCGAACUC____________AAACUCGAAACUCAA_______AUCGAAACUCAAAACUCGAAUCGAGUGGACAG_________GA ..((((......))))((((......))))............................................................(((((...)))))................. (-13.60 = -13.85 + 0.25)

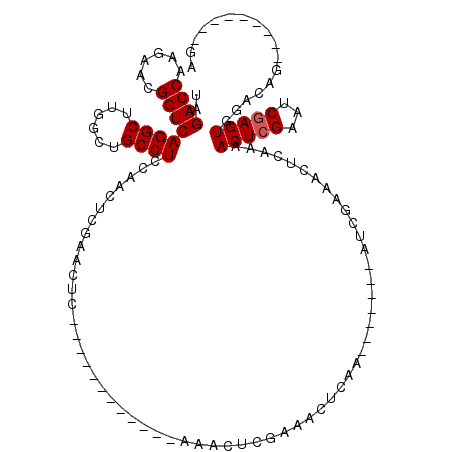
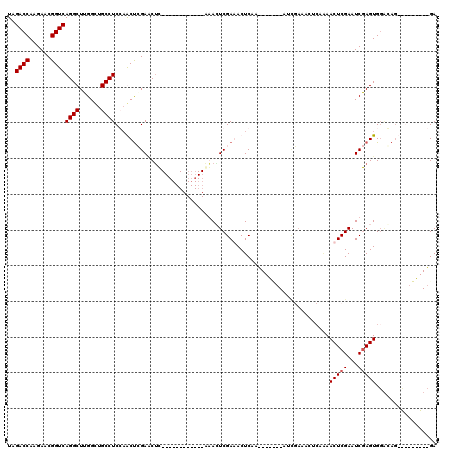
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:35:56 2006