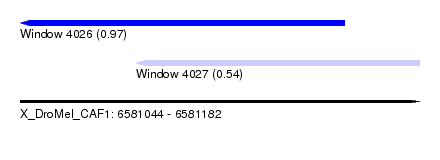
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 6,581,044 – 6,581,182 |
| Length | 138 |
| Max. P | 0.970759 |

| Location | 6,581,044 – 6,581,156 |
|---|---|
| Length | 112 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 89.38 |
| Mean single sequence MFE | -31.30 |
| Consensus MFE | -26.45 |
| Energy contribution | -27.09 |
| Covariance contribution | 0.64 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.98 |
| Structure conservation index | 0.85 |
| SVM decision value | 1.66 |
| SVM RNA-class probability | 0.970759 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
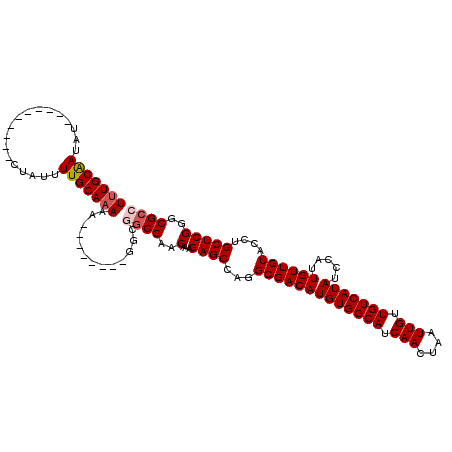
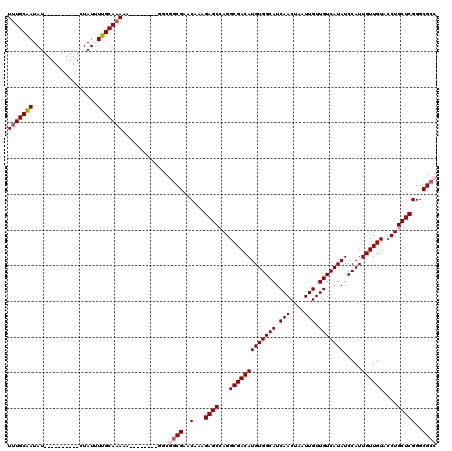
>X_DroMel_CAF1 6581044 112 - 22224390 UUUGCGAUAUAUGUAUGUAUAUAUUUUGCAAAAA--------GGCGGCGAACAAAGAGCCAGGCGACAUGUGGCAUCAACUAAUUGUUGUCAUAUCCAUUGUUGUACCUGCUCGGGCGCC (((((((.((((((....)))))).)))))))..--------...((((..(...((((.(((((((((((((((.(((....))).))))))).....)))))..))))))))..)))) ( -35.60) >DroSec_CAF1 25423 102 - 1 UUUGCAAUAU----------CUAUUUUGCAAAAA--------GACGGCGAACAAAGAGCCAGGCGACAUGUGGCAUCAACUAAUUGUUGUCAUAUCCAUUGUUGUACCUGCUCGGGCGCC (((((((...----------.....)))))))..--------...((((..(...((((.(((((((((((((((.(((....))).))))))).....)))))..))))))))..)))) ( -32.10) >DroSim_CAF1 37468 102 - 1 UUUGCAAUAU----------CUAUUUUGCAAAAA--------GGCGGCGAACAAAGAGCCAGGCGACAUGUGGCAUCAACUAAUUGUUGUCAUAUCCAUUGUUGUACCUGCUCGGGCGCC (((((((...----------.....)))))))..--------...((((..(...((((.(((((((((((((((.(((....))).))))))).....)))))..))))))))..)))) ( -32.10) >DroEre_CAF1 24751 101 - 1 UUUGCAAUAU----------CUAUUUUGCACAA---------GGCAGCGAACAAAGAGCGAGGCGACAUGUGGCAUCAACUAAUUGUUGUCAUAUCCAUUGUUGUACCUGCUCGGGCGCC ..(((((...----------.....)))))...---------....(((..(...(((((..(((((((((((((.(((....))).))))))).....))))))...))))))..))). ( -28.00) >DroYak_CAF1 36055 110 - 1 UUUGCAAUAU----------CUAUUUUGCAAAUAACGACAGCGACAACGAACAAAGAGCCAGGCGACAUGUGGCAUCAACUAAUUGUUGUCAUAUCCAUUGUUGUAACUGCUCGGGCGCC (((((((...----------.....)))))))...((((((..(((((((...........(....)((((((((.(((....))).))))))))...)))))))..))).)))...... ( -28.70) >consensus UUUGCAAUAU__________CUAUUUUGCAAAAA________GGCGGCGAACAAAGAGCCAGGCGACAUGUGGCAUCAACUAAUUGUUGUCAUAUCCAUUGUUGUACCUGCUCGGGCGCC (((((((..................))))))).............((((..(...((((...(((((((((((((.(((....))).))))))).....))))))....)))))..)))) (-26.45 = -27.09 + 0.64)

| Location | 6,581,084 – 6,581,182 |
|---|---|
| Length | 98 |
| Sequences | 4 |
| Columns | 100 |
| Reading direction | reverse |
| Mean pairwise identity | 89.22 |
| Mean single sequence MFE | -22.10 |
| Consensus MFE | -15.76 |
| Energy contribution | -16.32 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.59 |
| Structure conservation index | 0.71 |
| SVM decision value | 0.01 |
| SVM RNA-class probability | 0.539457 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
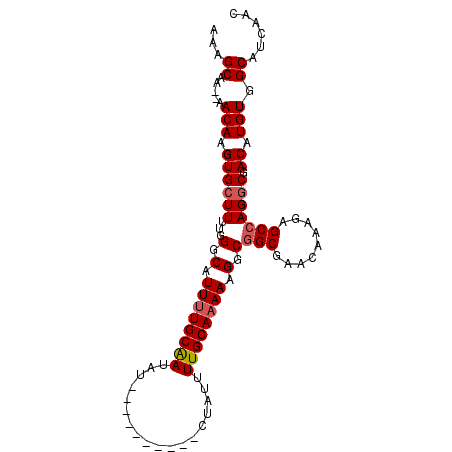
>X_DroMel_CAF1 6581084 98 - 22224390 AAAGCAA--AACAAGUGCUUUUGGGCAUUUUGCGAUAUAUGUAUGUAUAUAUUUUGCAAAAAGGCGGCGAACAAAGAGCCAGGCGACAUGUGGCAUCAAC ...((..--.(((.((((((....((.((((((((.((((((....)))))).))))))))..))(((.........))))))).)).))).))...... ( -26.60) >DroSec_CAF1 25463 89 - 1 AAAGCAAA-AACAAGUGCUUUUGGGCAUUUUGCAAUAU----------CUAUUUUGCAAAAAGACGGCGAACAAAGAGCCAGGCGACAUGUGGCAUCAAC ...((...-.(((.((((((.......((((((((...----------.....))))))))....(((.........))))))).)).))).))...... ( -20.40) >DroSim_CAF1 37508 90 - 1 AAAGCAAAAAACAAGUGCUUUUGGGCAUUUUGCAAUAU----------CUAUUUUGCAAAAAGGCGGCGAACAAAGAGCCAGGCGACAUGUGGCAUCAAC ...((.....(((.((((((....((.((((((((...----------.....))))))))..))(((.........))))))).)).))).))...... ( -21.80) >DroEre_CAF1 24791 87 - 1 AAAGCAA--AACAAGUGGUUUUGGGCAUUUUGCAAUAU----------CUAUUUUGCACAA-GGCAGCGAACAAAGAGCGAGGCGACAUGUGGCAUCAAC ...((..--.(((.((.(((((..((.((.(((((...----------.....))))).))-.)).((.........))))))).)).))).))...... ( -19.60) >consensus AAAGCAA__AACAAGUGCUUUUGGGCAUUUUGCAAUAU__________CUAUUUUGCAAAAAGGCGGCGAACAAAGAGCCAGGCGACAUGUGGCAUCAAC ...((.....(((.((((((...(.(.((((((((..................)))))))).).)(((.........))))))).)).))).))...... (-15.76 = -16.32 + 0.56)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:35:25 2006