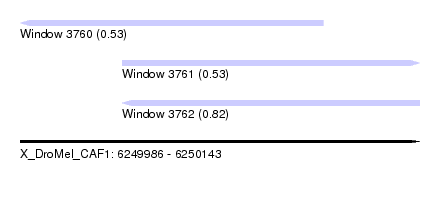
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 6,249,986 – 6,250,143 |
| Length | 157 |
| Max. P | 0.820806 |

| Location | 6,249,986 – 6,250,105 |
|---|---|
| Length | 119 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 95.82 |
| Mean single sequence MFE | -28.00 |
| Consensus MFE | -21.62 |
| Energy contribution | -22.38 |
| Covariance contribution | 0.75 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.56 |
| Structure conservation index | 0.77 |
| SVM decision value | -0.01 |
| SVM RNA-class probability | 0.529311 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 6249986 119 - 22224390 AGU-GUGUGCGUGUAAGCUUUUAGCUAGCUAAUUGCGGCCAAUUGGCCAAAACAAAACCACCGCAAAAAUUGCACAAAAAUUCAAAAUGUACUGCAAUGUAUGCAUUUAACUCAACCAGA (((-..((((((((((((..((((....))))..))((((....)))).....................)))))).............((((......))))))))...)))........ ( -29.10) >DroSec_CAF1 16773 119 - 1 AUA-GUGUGCGUGUAAGCUUUUAGCUAGCUAAUUGCGGCCAAUUGGCCAAAACAAAACCACCGCAAAAAUUGCACAAAAAUUCAACAUGUACUGCAAUGUAUGCAUUUAACUCAACCAGA ...-..((((((((..((..((((....))))..))((((....))))..............(((.....)))...........))))))))((((.....))))............... ( -28.70) >DroEre_CAF1 11442 119 - 1 ACU-GUGUGCGUGUAAGCUUUUAGCUAGCUAAUUGCGUCCAAUUGGCCAAAACAAAACCACCGCAAAAAUUGCACAAAAAUUCAGCAUGUACUGCAAUGUAUGCAUUUAACUCAACCGGA .((-(.((((((((.(((.....))).((((((((....))))))))...............(((.....)))...........))))))))((((.....))))...........))). ( -25.60) >DroYak_CAF1 23907 120 - 1 ACUGGUGUGCGUGUAAGCUUUUAGCUAGCUAAUUGCGUCCAAUUGGCCAAAACAAAACCACCGCAAAAAUUUCACAAAAAUUCAACAUGUACCGCAAUGUAUGCAUUUAACUCAACCAGA .(((((((((((((.(((.....))).((((((((....)))))))).....................................)))))))).(((.....)))..........))))). ( -28.60) >consensus ACU_GUGUGCGUGUAAGCUUUUAGCUAGCUAAUUGCGGCCAAUUGGCCAAAACAAAACCACCGCAAAAAUUGCACAAAAAUUCAACAUGUACUGCAAUGUAUGCAUUUAACUCAACCAGA ......((((((((((((..((((....))))..))((((....)))).....................)))))).............((((......)))))))).............. (-21.62 = -22.38 + 0.75)

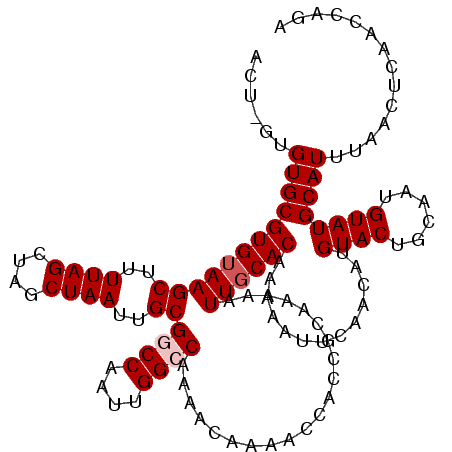
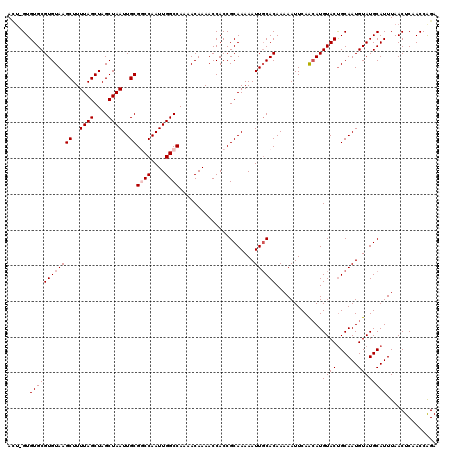
| Location | 6,250,026 – 6,250,143 |
|---|---|
| Length | 117 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 87.11 |
| Mean single sequence MFE | -33.53 |
| Consensus MFE | -23.24 |
| Energy contribution | -25.30 |
| Covariance contribution | 2.06 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.49 |
| Structure conservation index | 0.69 |
| SVM decision value | -0.01 |
| SVM RNA-class probability | 0.529027 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
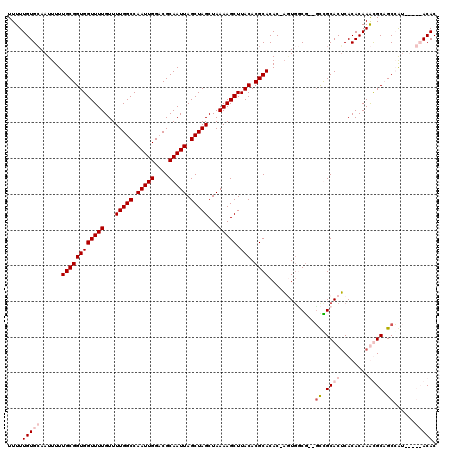
>X_DroMel_CAF1 6250026 117 + 22224390 UUUUUGUGCAAUUUUUGCGGUGGUUUUGUUUUGGCCAAUUGGCCGCAAUUAGCUAGCUAAAAGCUUACACGCACAC-ACUUUCG--GCCGCACUCACACAAGUGCCGCCGUCUAGCACAC ....(((((......((((((((.........((((....))))((..((((....))))..)))))).))))...-.....((--((.(((((......))))).))))....))))). ( -40.10) >DroSec_CAF1 16813 119 + 1 UUUUUGUGCAAUUUUUGCGGUGGUUUUGUUUUGGCCAAUUGGCCGCAAUUAGCUAGCUAAAAGCUUACACGCACAC-UAUGGCAGCGCUGCACGCACACAAGUGCCGCCGACUAGCACAC ....(((((......((((((((.........((((....))))((..((((....))))..)))))).))))...-..((((.(((((...........))))).))))....))))). ( -38.60) >DroEre_CAF1 11482 112 + 1 UUUUUGUGCAAUUUUUGCGGUGGUUUUGUUUUGGCCAAUUGGACGCAAUUAGCUAGCUAAAAGCUUACACGCACAC-AGUGGCG--CCCGCACUCACACAAACGCAGCGAU-----ACAC ..((((((.......((((((((((((...(((((.(((((....))))).)))))...))))).))).))))...-.((((..--.)))).....)))))).........-----.... ( -26.60) >DroYak_CAF1 23947 113 + 1 UUUUUGUGAAAUUUUUGCGGUGGUUUUGUUUUGGCCAAUUGGACGCAAUUAGCUAGCUAAAAGCUUACACGCACACCAGUGCCG--GUUGCACUCACACAAACGCAGCGAU-----ACAC ..((((((.......((((((((((((...(((((.(((((....))))).)))))...))))).))).))))....(((((..--...)))))..)))))).........-----.... ( -28.80) >consensus UUUUUGUGCAAUUUUUGCGGUGGUUUUGUUUUGGCCAAUUGGACGCAAUUAGCUAGCUAAAAGCUUACACGCACAC_AGUGGCG__GCCGCACUCACACAAACGCAGCCAU_____ACAC ....(((((......((((((((((((...(((((.(((((....))))).)))))...))))).))).)))).............((.((((........)))).))......))))). (-23.24 = -25.30 + 2.06)

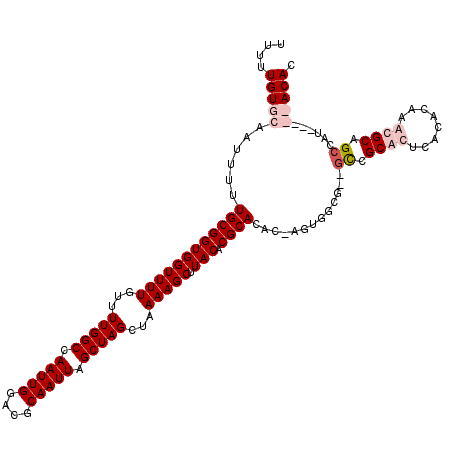
| Location | 6,250,026 – 6,250,143 |
|---|---|
| Length | 117 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 87.11 |
| Mean single sequence MFE | -37.88 |
| Consensus MFE | -25.56 |
| Energy contribution | -27.00 |
| Covariance contribution | 1.44 |
| Combinations/Pair | 1.10 |
| Mean z-score | -2.35 |
| Structure conservation index | 0.67 |
| SVM decision value | 0.68 |
| SVM RNA-class probability | 0.820806 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
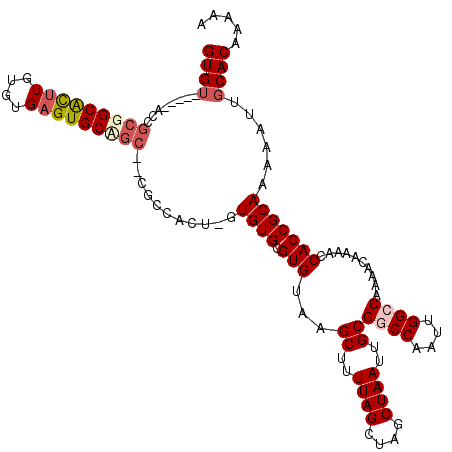
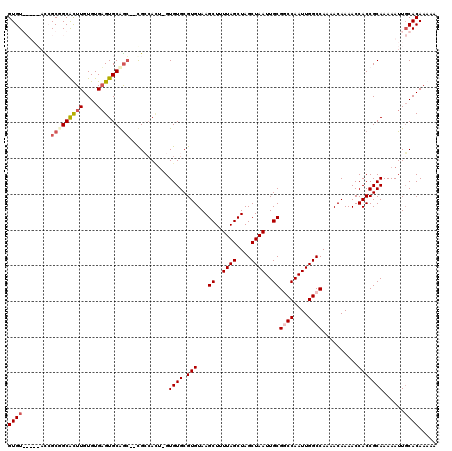
>X_DroMel_CAF1 6250026 117 - 22224390 GUGUGCUAGACGGCGGCACUUGUGUGAGUGCGGC--CGAAAGU-GUGUGCGUGUAAGCUUUUAGCUAGCUAAUUGCGGCCAAUUGGCCAAAACAAAACCACCGCAAAAAUUGCACAAAAA .(((((....((((.((((((....)))))).))--))...((-(.(((.((....((..((((....))))..))((((....))))........)))))))).......))))).... ( -46.90) >DroSec_CAF1 16813 119 - 1 GUGUGCUAGUCGGCGGCACUUGUGUGCGUGCAGCGCUGCCAUA-GUGUGCGUGUAAGCUUUUAGCUAGCUAAUUGCGGCCAAUUGGCCAAAACAAAACCACCGCAAAAAUUGCACAAAAA .(((((.....((((((.((.((......)))).))))))...-.((((.(((...((..((((....))))..))((((....))))..........)))))))......))))).... ( -42.20) >DroEre_CAF1 11482 112 - 1 GUGU-----AUCGCUGCGUUUGUGUGAGUGCGGG--CGCCACU-GUGUGCGUGUAAGCUUUUAGCUAGCUAAUUGCGUCCAAUUGGCCAAAACAAAACCACCGCAAAAAUUGCACAAAAA ....-----.........((((((..(.((((((--(((....-..)))).(((.(((.....))).((((((((....))))))))....)))......)))))....)..)))))).. ( -33.30) >DroYak_CAF1 23947 113 - 1 GUGU-----AUCGCUGCGUUUGUGUGAGUGCAAC--CGGCACUGGUGUGCGUGUAAGCUUUUAGCUAGCUAAUUGCGUCCAAUUGGCCAAAACAAAACCACCGCAAAAAUUUCACAAAAA (((.-----.....(((.........(((((...--..)))))((((....(((.(((.....))).((((((((....))))))))....)))....))))))).......)))..... ( -29.12) >consensus GUGU_____ACCGCGGCACUUGUGUGAGUGCAGC__CGCCACU_GUGUGCGUGUAAGCUUUUAGCUAGCUAAUUGCGGCCAAUUGGCCAAAACAAAACCACCGCAAAAAUUGCACAAAAA ((((........(((((((((....)))))))))...........((((.(((...((..((((....))))..))((((....))))..........)))))))......))))..... (-25.56 = -27.00 + 1.44)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:31:30 2006