
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 5,799,708 – 5,799,819 |
| Length | 111 |
| Max. P | 0.879996 |

| Location | 5,799,708 – 5,799,819 |
|---|---|
| Length | 111 |
| Sequences | 4 |
| Columns | 111 |
| Reading direction | forward |
| Mean pairwise identity | 74.54 |
| Mean single sequence MFE | -32.53 |
| Consensus MFE | -21.01 |
| Energy contribution | -22.82 |
| Covariance contribution | 1.81 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.79 |
| Structure conservation index | 0.65 |
| SVM decision value | 0.91 |
| SVM RNA-class probability | 0.879996 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
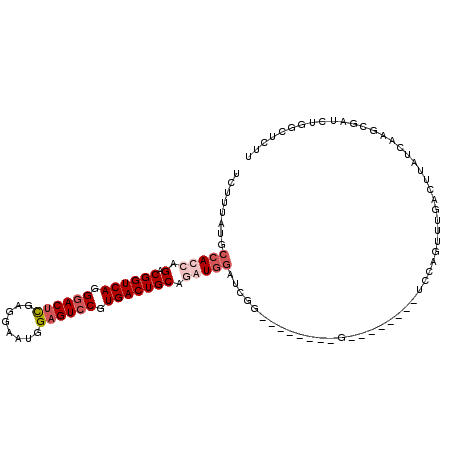
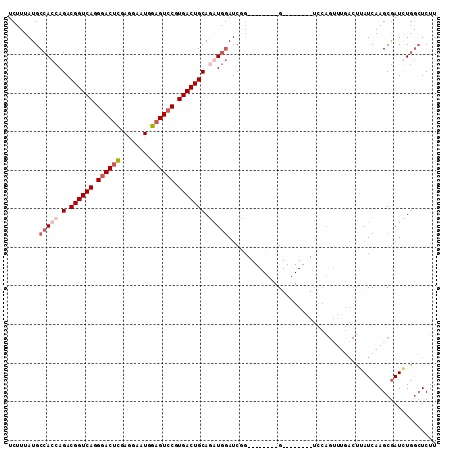
>X_DroMel_CAF1 5799708 111 + 22224390 UCUUUAUGCCACCAGACGGUCAGGGACUCAAGGAAUGGAGUCCGUGACUGCAUAUGGAUCGGUAUACUCCGUAAGUAAAUCCAGUUUGACUUAUCAAGCGAUGUGGCUCUU .......(((((..(.((((((.((((((........)))))).)))))))...((((((((......))).......)))))((((((....))))))...))))).... ( -40.01) >DroSim_CAF1 5400 110 + 1 UCUUUAUGCCACCAGACGGUCAGGGACUCGAGGAAUGGAGUCCGUGACUGCAUAUGGAUCGGUAUACU-CGUAAGUAAAUCCAGUUUGACUUAUCAAGCGAUGUGGCUCUU .......(((((((..((((((.((((((.(....).)))))).))))))....)))(((((..((((-....))))...)).((((((....))))))))).)))).... ( -39.70) >DroEre_CAF1 5120 84 + 1 UCUUUAUGCCACCAGACGGUCAGGGACUCGAGGAAUGGAGUGCGUGACUGCAGGUGGAUCUG--------G-------------------CGAUCAAGCGAUCUGGCUCUU .......((((((.(.((((((.(.((((.(....).)))).).))))))).)))(((((..--------(-------------------(......)))))))))).... ( -30.70) >DroYak_CAF1 5911 95 + 1 UCUUUAUGCCACCAGACGGUCAGGGACUUGAAGAAUGGCGUCCGUGACUGCAGGUAAAUCCC--------G--------UCCAGACCAACUAAUCAAGCAAUCUAAGUCUU ......(((.(((.(.((((((.((((............)))).))))))).))).......--------(--------(....))...........)))........... ( -19.70) >consensus UCUUUAUGCCACCAGACGGUCAGGGACUCGAGGAAUGGAGUCCGUGACUGCAGAUGGAUCGG________G________UCCAGUUUGACUUAUCAAGCGAUCUGGCUCUU ........(((((.(.((((((.((((((........)))))).))))))).)))))...................................................... (-21.01 = -22.82 + 1.81)

| Location | 5,799,708 – 5,799,819 |
|---|---|
| Length | 111 |
| Sequences | 4 |
| Columns | 111 |
| Reading direction | reverse |
| Mean pairwise identity | 74.54 |
| Mean single sequence MFE | -28.95 |
| Consensus MFE | -17.96 |
| Energy contribution | -18.65 |
| Covariance contribution | 0.69 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.55 |
| Structure conservation index | 0.62 |
| SVM decision value | -0.04 |
| SVM RNA-class probability | 0.515672 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
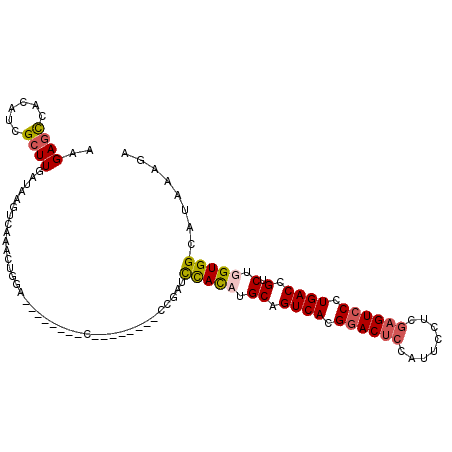
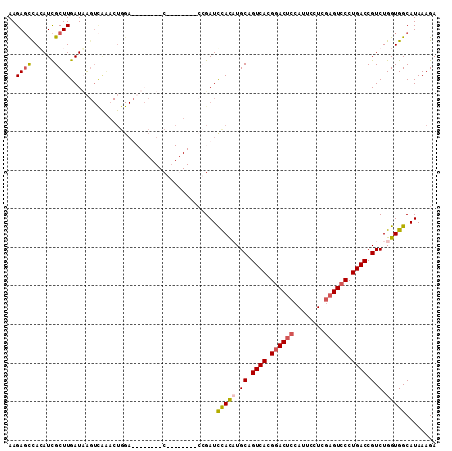
>X_DroMel_CAF1 5799708 111 - 22224390 AAGAGCCACAUCGCUUGAUAAGUCAAACUGGAUUUACUUACGGAGUAUACCGAUCCAUAUGCAGUCACGGACUCCAUUCCUUGAGUCCCUGACCGUCUGGUGGCAUAAAGA ....(((((...((((((....))))..(((((((((((...)))))....))))))...)).((((.((((((........)))))).))))......)))))....... ( -33.50) >DroSim_CAF1 5400 110 - 1 AAGAGCCACAUCGCUUGAUAAGUCAAACUGGAUUUACUUACG-AGUAUACCGAUCCAUAUGCAGUCACGGACUCCAUUCCUCGAGUCCCUGACCGUCUGGUGGCAUAAAGA ....(((((...((((((....))))..((((((((((....-))))....))))))...)).((((.((((((........)))))).))))......)))))....... ( -33.40) >DroEre_CAF1 5120 84 - 1 AAGAGCCAGAUCGCUUGAUCG-------------------C--------CAGAUCCACCUGCAGUCACGCACUCCAUUCCUCGAGUCCCUGACCGUCUGGUGGCAUAAAGA ....(((.((((....))))(-------------------(--------(((((.........((((.(.((((........)))).).)))).))))))))))....... ( -23.60) >DroYak_CAF1 5911 95 - 1 AAGACUUAGAUUGCUUGAUUAGUUGGUCUGGA--------C--------GGGAUUUACCUGCAGUCACGGACGCCAUUCUUCAAGUCCCUGACCGUCUGGUGGCAUAAAGA ....(((....((((..(((((.(((((.(((--------(--------((......)).((.(((...)))))..........))))..))))).)))))))))..))). ( -25.30) >consensus AAGAGCCACAUCGCUUGAUAAGUCAAACUGGA________C________CCGAUCCACAUGCAGUCACGGACUCCAUUCCUCGAGUCCCUGACCGUCUGGUGGCAUAAAGA ..((((......))))......................................(((((.((.((((.((((((........)))))).)))).).).)))))........ (-17.96 = -18.65 + 0.69)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:27:05 2006