
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 5,521,586 – 5,521,685 |
| Length | 99 |
| Max. P | 0.999597 |

| Location | 5,521,586 – 5,521,685 |
|---|---|
| Length | 99 |
| Sequences | 4 |
| Columns | 100 |
| Reading direction | forward |
| Mean pairwise identity | 80.84 |
| Mean single sequence MFE | -29.75 |
| Consensus MFE | -19.55 |
| Energy contribution | -19.55 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.15 |
| Mean z-score | -3.14 |
| Structure conservation index | 0.66 |
| SVM decision value | 2.82 |
| SVM RNA-class probability | 0.997222 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
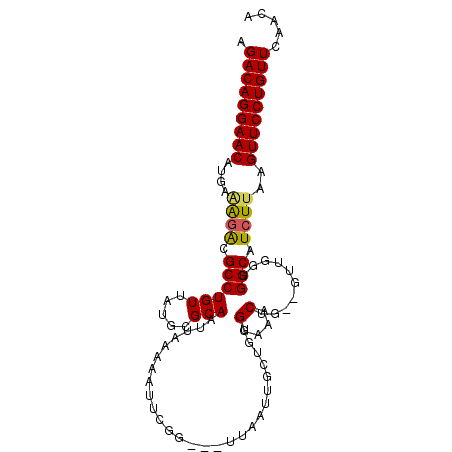
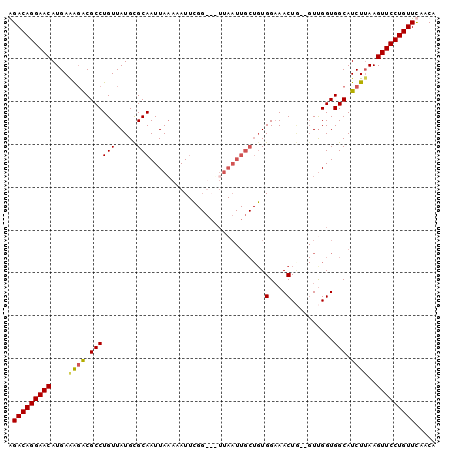
>X_DroMel_CAF1 5521586 99 + 22224390 GGACAGGAACAUGAGAGACGCCUGUUAUGCGCAAUUAAAAAUUUGG-GUUUAAUUGCUGUGGAAACUGUUGUUGGUGGCAUCUUAAGUUCCUGUUUAACA ((((((((((.(((((..((((......(((((((((((.......-.)))))))))...(....).))....))))...))))).)))))))))).... ( -34.80) >DroSec_CAF1 11664 80 + 1 GGACAGGAACAUGAAAGACGCCUGUUAUGCGCAAUUAAAAAU-----------------UGGAAACUG---UUGGUGGCAUCUUAAGUUCCUGUUUAACA ((((((((((....((((.((((((.....)))......(((-----------------.(....).)---))...))).))))..)))))))))).... ( -22.10) >DroEre_CAF1 11796 95 + 1 AGACAGGAACAUGAAGCACGCCUGUUAUGUGCAAUUAAACAUUCGGU--UUAAUUGCUGUGGAAACU---GUUGGUGGCAUCUAAAGUUCCUGUUCAACA .(((((((((..((.((.((((........((((((((((.....))--))))))))...(....).---...)))))).))....)))))))))..... ( -33.80) >DroYak_CAF1 11721 100 + 1 AGACAGGAACAUGAAAGGCGCCUGUUAUGUGCAAUUACAAAUUCGGUGGUUAAUUGCUGUGGCAACUAAAGUCGGUGGCAUCUUAAGUUCCUGUUCAACA .(((((((((....(((..((((((((((.(((((((.............))))))))))))))(((......))))))..)))..)))))))))..... ( -28.32) >consensus AGACAGGAACAUGAAAGACGCCUGUUAUGCGCAAUUAAAAAUUCGG___UUAAUUGCUGUGGAAACUG__GUUGGUGGCAUCUUAAGUUCCUGUUCAACA .(((((((((....((((.((((((.....)))...........................(....)..........))).))))..)))))))))..... (-19.55 = -19.55 + 0.00)

| Location | 5,521,586 – 5,521,685 |
|---|---|
| Length | 99 |
| Sequences | 4 |
| Columns | 100 |
| Reading direction | reverse |
| Mean pairwise identity | 80.84 |
| Mean single sequence MFE | -27.41 |
| Consensus MFE | -19.23 |
| Energy contribution | -22.30 |
| Covariance contribution | 3.06 |
| Combinations/Pair | 1.04 |
| Mean z-score | -4.24 |
| Structure conservation index | 0.70 |
| SVM decision value | 3.77 |
| SVM RNA-class probability | 0.999597 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
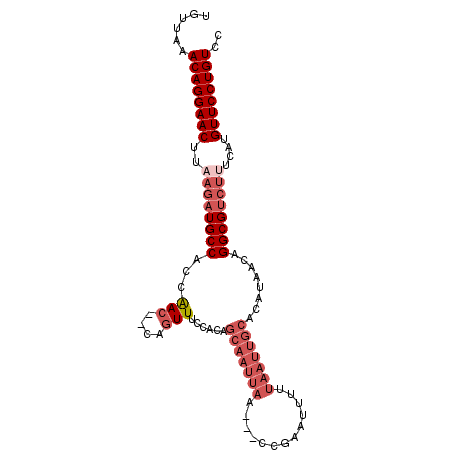
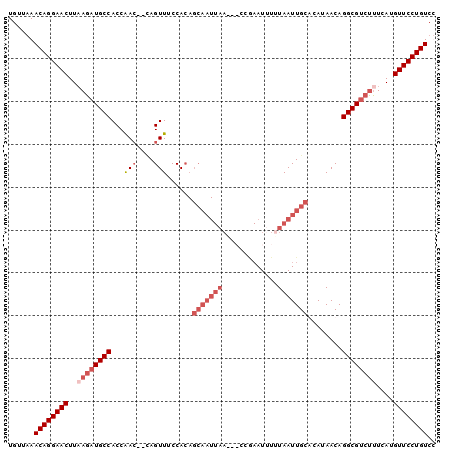
>X_DroMel_CAF1 5521586 99 - 22224390 UGUUAAACAGGAACUUAAGAUGCCACCAACAACAGUUUCCACAGCAAUUAAAC-CCAAAUUUUUAAUUGCGCAUAACAGGCGUCUCUCAUGUUCCUGUCC ......((((((((...(((((((...(((....)))......(((((((((.-.......)))))))))........))))))).....)))))))).. ( -30.00) >DroSec_CAF1 11664 80 - 1 UGUUAAACAGGAACUUAAGAUGCCACCAA---CAGUUUCCA-----------------AUUUUUAAUUGCGCAUAACAGGCGUCUUUCAUGUUCCUGUCC ......((((((((..((((((((.....---(((((....-----------------......))))).(.....).))))))))....)))))))).. ( -22.70) >DroEre_CAF1 11796 95 - 1 UGUUGAACAGGAACUUUAGAUGCCACCAAC---AGUUUCCACAGCAAUUAA--ACCGAAUGUUUAAUUGCACAUAACAGGCGUGCUUCAUGUUCCUGUCU ......((((((((...(((((((......---.((....)).((((((((--((.....))))))))))........))))).))....)))))))).. ( -30.00) >DroYak_CAF1 11721 100 - 1 UGUUGAACAGGAACUUAAGAUGCCACCGACUUUAGUUGCCACAGCAAUUAACCACCGAAUUUGUAAUUGCACAUAACAGGCGCCUUUCAUGUUCCUGUCU ......((((((((..(((.((((..((((....)))).....(((((((.............)))))))........)))).)))....)))))))).. ( -26.92) >consensus UGUUAAACAGGAACUUAAGAUGCCACCAAC__CAGUUUCCACAGCAAUUAA___CCGAAUUUUUAAUUGCACAUAACAGGCGUCUUUCAUGUUCCUGUCC ......((((((((..((((((((...(((....)))......(((((((.............)))))))........))))))))....)))))))).. (-19.23 = -22.30 + 3.06)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:24:10 2006