
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 5,514,857 – 5,514,950 |
| Length | 93 |
| Max. P | 0.996230 |

| Location | 5,514,857 – 5,514,950 |
|---|---|
| Length | 93 |
| Sequences | 6 |
| Columns | 96 |
| Reading direction | forward |
| Mean pairwise identity | 89.48 |
| Mean single sequence MFE | -30.72 |
| Consensus MFE | -24.11 |
| Energy contribution | -24.08 |
| Covariance contribution | -0.03 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.43 |
| Structure conservation index | 0.78 |
| SVM decision value | 0.11 |
| SVM RNA-class probability | 0.590766 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 5514857 93 + 22224390 GGCAGCUGCAUCUGCUGGCAG---UAGCGCAGAUGCUGCCGCUGCGAACAUCAAAAAGGAUGAGAAUGGUGCAAAGGUGUAUGAUGAAAAACGGCG ((((((...((((((..((..---..))))))))))))))((((....((((......))))...((.(((((....))))).))......)))). ( -32.00) >DroVir_CAF1 6988 90 + 1 UGCAGCCGCAUCUGCGGGCAG------CGCAGAUGCUGCCGCGACAAACACCAAAAAGGAUGAGAAUGGUGCAAAGGUGCAUGAUGAAAAACGCCG .((.((.(((((((((.....------))))))))).)).))...............((......((.(((((....))))).))........)). ( -30.54) >DroYak_CAF1 4882 93 + 1 GGCAGCUGCAUCUGCCGGCGG---UAGCGGAGAUGCUGCCGCUGCAAACACCAAAAAGGAUGAGAAUGGUGCAAAGGUGUAUGAUGAAAAACGGCG (((((......)))))(((((---(((((....))))))))))((...((((.....)).))...((.(((((....))))).))........)). ( -32.30) >DroMoj_CAF1 5872 90 + 1 UGCAGCUGCAUCUGCGGGUGG------CGCAGAUGCCGCCGCGUCAAACACCAAAAAGGAUGAGAAUGGUGCAAAGGUGCAUGAUGAAAAACGGCG .((.((.(((((((((.....------))))))))).)).))(((...((((.....)).))...((.(((((....))))).)).......))). ( -32.90) >DroAna_CAF1 4667 96 + 1 UGCAGCUGCAUCUGCGAGCAGUAGUAGCGCAGAUGCUGCCGCUACAAACACCAAAAAGGAUGAGAAUGGUGCAAAGGUGUAUGAUGAAAAACGGCG .......(((((((((.((....))..))))))))).((((.......((((.....)).))...((.(((((....))))).))......)))). ( -28.30) >DroPer_CAF1 22250 93 + 1 UGCAGCUGCAUCUGCGACCAA---CAGCGCAGAUGCUGCCGCGACAAACACCAAAAAGGAUGAGAAUGGUGCAAAGGUGUAUGAUGAAAAACGGCG .((.((.(((((((((.....---...))))))))).)).))..((.(((((.......................))))).))............. ( -28.30) >consensus UGCAGCUGCAUCUGCGGGCAG___UAGCGCAGAUGCUGCCGCGACAAACACCAAAAAGGAUGAGAAUGGUGCAAAGGUGUAUGAUGAAAAACGGCG (((.((.(((((((((...........))))))))).)).))).....((((.....)).))...((.(((((....))))).))........... (-24.11 = -24.08 + -0.03)

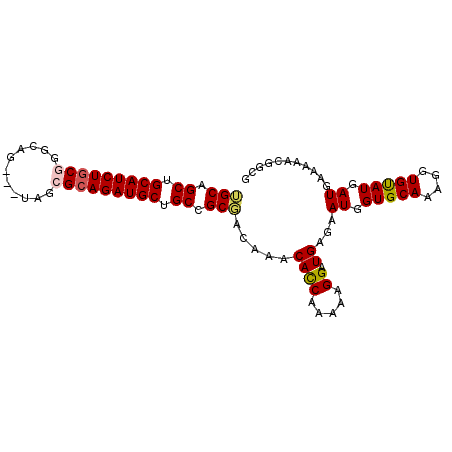
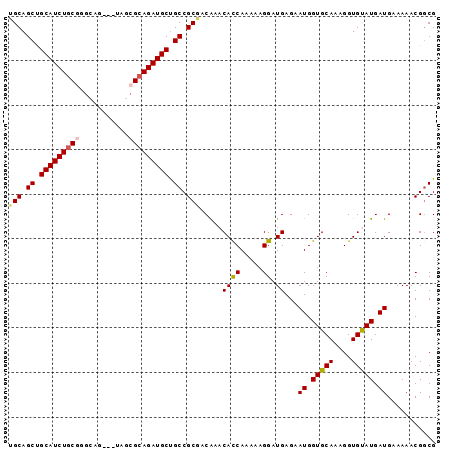
| Location | 5,514,857 – 5,514,950 |
|---|---|
| Length | 93 |
| Sequences | 6 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 89.48 |
| Mean single sequence MFE | -31.38 |
| Consensus MFE | -27.02 |
| Energy contribution | -27.36 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -3.58 |
| Structure conservation index | 0.86 |
| SVM decision value | 2.67 |
| SVM RNA-class probability | 0.996230 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 5514857 93 - 22224390 CGCCGUUUUUCAUCAUACACCUUUGCACCAUUCUCAUCCUUUUUGAUGUUCGCAGCGGCAGCAUCUGCGCUA---CUGCCAGCAGAUGCAGCUGCC .((((((...(((((............................))))).....)))))).((((((((((..---..))..))))))))....... ( -27.59) >DroVir_CAF1 6988 90 - 1 CGGCGUUUUUCAUCAUGCACCUUUGCACCAUUCUCAUCCUUUUUGGUGUUUGUCGCGGCAGCAUCUGCG------CUGCCCGCAGAUGCGGCUGCA ..((((........))))......((((((.............)))))).....(((((.(((((((((------.....))))))))).))))). ( -34.72) >DroYak_CAF1 4882 93 - 1 CGCCGUUUUUCAUCAUACACCUUUGCACCAUUCUCAUCCUUUUUGGUGUUUGCAGCGGCAGCAUCUCCGCUA---CCGCCGGCAGAUGCAGCUGCC .((.((..........))......((((((.............))))))..)).(((((.((((((((((..---..).))).)))))).))))). ( -27.42) >DroMoj_CAF1 5872 90 - 1 CGCCGUUUUUCAUCAUGCACCUUUGCACCAUUCUCAUCCUUUUUGGUGUUUGACGCGGCGGCAUCUGCG------CCACCCGCAGAUGCAGCUGCA ............(((.(((((.......................))))).))).(((((.(((((((((------.....))))))))).))))). ( -33.80) >DroAna_CAF1 4667 96 - 1 CGCCGUUUUUCAUCAUACACCUUUGCACCAUUCUCAUCCUUUUUGGUGUUUGUAGCGGCAGCAUCUGCGCUACUACUGCUCGCAGAUGCAGCUGCA ...............((((.....((((((.............)))))).))))(((((.((((((((((.......))..)))))))).))))). ( -32.42) >DroPer_CAF1 22250 93 - 1 CGCCGUUUUUCAUCAUACACCUUUGCACCAUUCUCAUCCUUUUUGGUGUUUGUCGCGGCAGCAUCUGCGCUG---UUGGUCGCAGAUGCAGCUGCA ........................((((((.............)))))).....(((((.((((((((((..---...).))))))))).))))). ( -32.32) >consensus CGCCGUUUUUCAUCAUACACCUUUGCACCAUUCUCAUCCUUUUUGGUGUUUGUAGCGGCAGCAUCUGCGCUA___CUGCCCGCAGAUGCAGCUGCA ........................((((((.............)))))).....(((((.((((((((.............)))))))).))))). (-27.02 = -27.36 + 0.33)

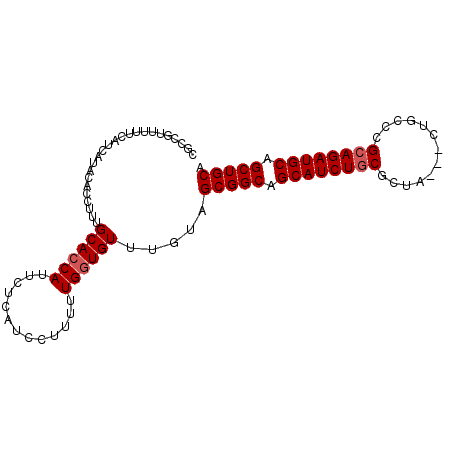
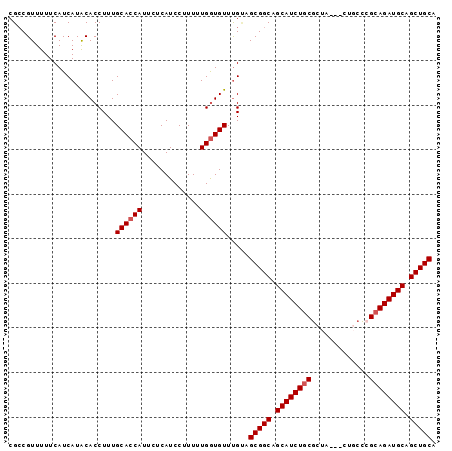
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:24:03 2006