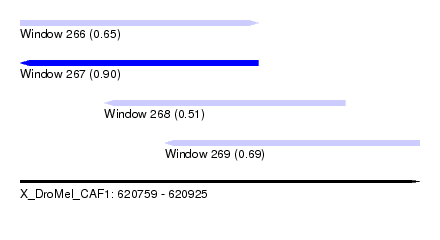
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 620,759 – 620,925 |
| Length | 166 |
| Max. P | 0.902085 |

| Location | 620,759 – 620,858 |
|---|---|
| Length | 99 |
| Sequences | 6 |
| Columns | 106 |
| Reading direction | forward |
| Mean pairwise identity | 79.31 |
| Mean single sequence MFE | -27.97 |
| Consensus MFE | -17.52 |
| Energy contribution | -16.83 |
| Covariance contribution | -0.69 |
| Combinations/Pair | 1.25 |
| Mean z-score | -1.55 |
| Structure conservation index | 0.63 |
| SVM decision value | 0.25 |
| SVM RNA-class probability | 0.653311 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 620759 99 + 22224390 ----UAGCGUUGUUCCAGAAGUUGUUAACCGCCGGCGGCACGGGA---AAGUCCGCUGGCAGUUUAUCGCAAAAUGGAGUGAACUCACUAUAACCACAACUUUUUA ----....(((((((((....((((.(((.((((((((.((....---..)))))))))).)))....))))..))))(((....))).......)))))...... ( -33.60) >DroPse_CAF1 9950 101 + 1 UG--UGAACAUGUUCCAAGAGUUGUUAACCGCUAGCGGAACGGAUAUUGAAUCCGCAAGCAGUUUAAUCGCAAAUGAA---AGUUCGCUAUAACCACAACUUUUUA .(--.(((....))))(((((((((.(((.(((.(((((............))))).))).))).....((.(((...---.))).)).......))))))))).. ( -23.20) >DroSec_CAF1 5002 98 + 1 ----UAGCCUUGUUCCAGAAGUUGUUAACCGCCGGCGGCACGGGA---AAGUCCGCUGGCAGUUUAUCGCCAAAUGGAGUCAAUUCACUAUAACCACA-CUUUUUA ----..(..((((((((...((.((.(((.((((((((.((....---..)))))))))).))).)).))....))))).)))..)............-....... ( -28.10) >DroEre_CAF1 5367 104 + 1 GGUGUACAGUUGUUCCAGAAGUUGUUAACCGCCUGCGGCACGGGAA--AAGUCCGCUGGCAGUUUAUCGCCAAAUGGAGUGACCUCACUGUAACCACAACUUUUUA (((.((((((..(((((...((.((.(((.(((.((((.((.....--..)))))).))).))).)).))....)))))..)).....)))))))........... ( -31.80) >DroYak_CAF1 4165 103 + 1 GCU-UACUGUUGUUCCAGAAGUUGUUAACCGCCGGCGGCACGGGAA--AAGUCCGCCAGCAGUUUAUCGCCAAAUGGAGUGAACUUACUUUAACCACAACUUUUUA ...-............(((((((((.....((.(((((.((.....--..))))))).))(((((((..((....)).)))))))..........))))))))).. ( -27.90) >DroPer_CAF1 17920 101 + 1 UG--UGAACAUGUUCCAAGAGUUGUUAACCGCUAGCGGAACGGAUAUUGAAUCCGCAAGCAGUUUAAUCGCAAAUGAA---AGUUCGCUAUAACCACAACUUUUUA .(--.(((....))))(((((((((.(((.(((.(((((............))))).))).))).....((.(((...---.))).)).......))))))))).. ( -23.20) >consensus _G__UAAACUUGUUCCAGAAGUUGUUAACCGCCGGCGGCACGGGAA__AAGUCCGCUAGCAGUUUAUCGCCAAAUGGAGUGAACUCACUAUAACCACAACUUUUUA ................(((((((((.(((.(((.((((.((.........)))))).))).)))..........(((.......)))........))))))))).. (-17.52 = -16.83 + -0.69)

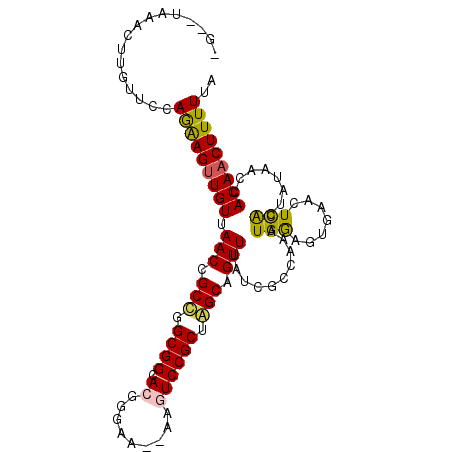
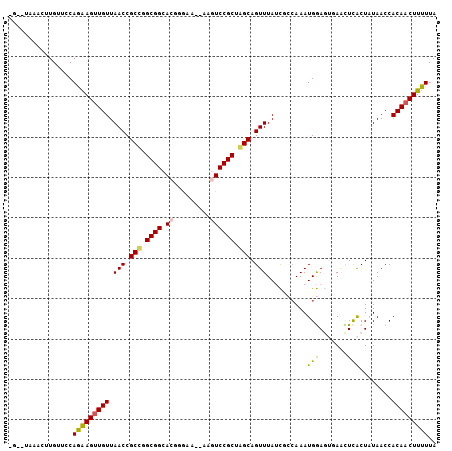
| Location | 620,759 – 620,858 |
|---|---|
| Length | 99 |
| Sequences | 6 |
| Columns | 106 |
| Reading direction | reverse |
| Mean pairwise identity | 79.31 |
| Mean single sequence MFE | -29.60 |
| Consensus MFE | -20.95 |
| Energy contribution | -20.54 |
| Covariance contribution | -0.41 |
| Combinations/Pair | 1.18 |
| Mean z-score | -1.88 |
| Structure conservation index | 0.71 |
| SVM decision value | 1.02 |
| SVM RNA-class probability | 0.902085 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 620759 99 - 22224390 UAAAAAGUUGUGGUUAUAGUGAGUUCACUCCAUUUUGCGAUAAACUGCCAGCGGACUU---UCCCGUGCCGCCGGCGGUUAACAACUUCUGGAACAACGCUA---- ......(((((.......(((....)))((((..........(((((((.((((((..---....)).)))).))))))).........)))))))))....---- ( -29.31) >DroPse_CAF1 9950 101 - 1 UAAAAAGUUGUGGUUAUAGCGAACU---UUCAUUUGCGAUUAAACUGCUUGCGGAUUCAAUAUCCGUUCCGCUAGCGGUUAACAACUCUUGGAACAUGUUCA--CA .....((((((.......(((((..---....))))).....(((((((.(((((............))))).))))))).))))))....(((....))).--.. ( -27.20) >DroSec_CAF1 5002 98 - 1 UAAAAAG-UGUGGUUAUAGUGAAUUGACUCCAUUUGGCGAUAAACUGCCAGCGGACUU---UCCCGUGCCGCCGGCGGUUAACAACUUCUGGAACAAGGCUA---- ....(((-((..((((........))))..)))))(((....(((((((.((((((..---....)).)))).)))))))........((......))))).---- ( -28.90) >DroEre_CAF1 5367 104 - 1 UAAAAAGUUGUGGUUACAGUGAGGUCACUCCAUUUGGCGAUAAACUGCCAGCGGACUU--UUCCCGUGCCGCAGGCGGUUAACAACUUCUGGAACAACUGUACACC ...........(((((((((...((...((((...((.....(((((((.((((((..--.....)).)))).))))))).....))..)))))).)))))).))) ( -30.60) >DroYak_CAF1 4165 103 - 1 UAAAAAGUUGUGGUUAAAGUAAGUUCACUCCAUUUGGCGAUAAACUGCUGGCGGACUU--UUCCCGUGCCGCCGGCGGUUAACAACUUCUGGAACAACAGUA-AGC ....(((((((.((((((...((....))...))))))....((((((((((((((..--.....)).)))))))))))).)))))))(((......)))..-... ( -34.40) >DroPer_CAF1 17920 101 - 1 UAAAAAGUUGUGGUUAUAGCGAACU---UUCAUUUGCGAUUAAACUGCUUGCGGAUUCAAUAUCCGUUCCGCUAGCGGUUAACAACUCUUGGAACAUGUUCA--CA .....((((((.......(((((..---....))))).....(((((((.(((((............))))).))))))).))))))....(((....))).--.. ( -27.20) >consensus UAAAAAGUUGUGGUUAUAGUGAAUUCACUCCAUUUGGCGAUAAACUGCCAGCGGACUU__UUCCCGUGCCGCCGGCGGUUAACAACUUCUGGAACAACUCUA__C_ ....(((((((((................))...........(((((((.((((((.........)).)))).))))))).))))))).................. (-20.95 = -20.54 + -0.41)

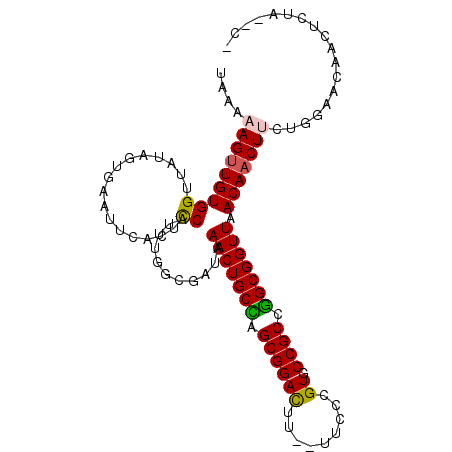
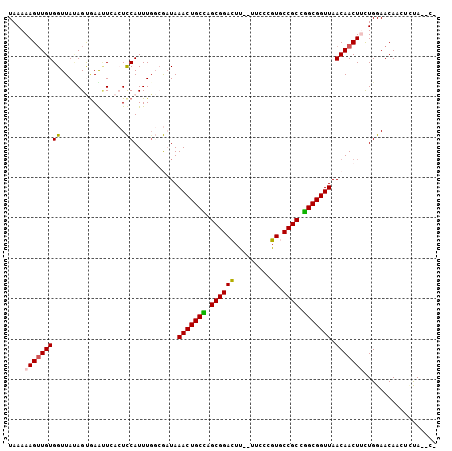
| Location | 620,794 – 620,894 |
|---|---|
| Length | 100 |
| Sequences | 6 |
| Columns | 103 |
| Reading direction | reverse |
| Mean pairwise identity | 81.82 |
| Mean single sequence MFE | -27.70 |
| Consensus MFE | -18.70 |
| Energy contribution | -18.48 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.39 |
| Structure conservation index | 0.68 |
| SVM decision value | -0.04 |
| SVM RNA-class probability | 0.513613 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 620794 100 - 22224390 UUUCCACUGCCCUCCAAGUUAACCAGGGGCUCUGGAUAAAAAGUUGUGGUUAUAGUGAGUUCACUCCAUUUUGCGAUAAACUGCCAGCGGACUU---UCCCGU ..((((..(((((............)))))..))))......((((..(....((((....)))).....)..)))).........((((....---..)))) ( -24.70) >DroPse_CAF1 9987 100 - 1 UUUCCAUUGCCCCCAAUCAUAGCCUGGGGCUUUGGAUAAAAAGUUGUGGUUAUAGCGAACU---UUCAUUUGCGAUUAAACUGCUUGCGGAUUCAAUAUCCGU ..((((..(((((............)))))..)))).........(..(((...(((((..---....))))).....)))..)..((((((.....)))))) ( -29.50) >DroSec_CAF1 5037 99 - 1 UUUCCACUGCCCUGCAAGUUAACCAGGGGCUCUGGAUAAAAAG-UGUGGUUAUAGUGAAUUGACUCCAUUUGGCGAUAAACUGCCAGCGGACUU---UCCCGU ..((((..(((((............)))))..))))....(((-((..((((........))))..)))))((((......)))).((((....---..)))) ( -29.50) >DroEre_CAF1 5406 101 - 1 UUUCCAUUGCCCUCCAAGUUAACUAGGGGCUCUGGAUAAAAAGUUGUGGUUACAGUGAGGUCACUCCAUUUGGCGAUAAACUGCCAGCGGACUU--UUCCCGU ..((((..(((((...((....)).)))))..))))..(((((((((((....((((....))))))))((((((......))))))..)))))--))..... ( -29.30) >DroYak_CAF1 4203 101 - 1 UUUCCACUGCCCUCCAAGUUAACCAGGGGCUCUGGAUAAAAAGUUGUGGUUAAAGUAAGUUCACUCCAUUUGGCGAUAAACUGCUGGCGGACUU--UUCCCGU ..((((..(((((............)))))..)))).....((((((.((((((...((....))...)))))).)).))))....((((....--...)))) ( -23.70) >DroPer_CAF1 17957 100 - 1 UUUCCAUUGCCCCCAAUCAUAGCCUGGGGCUUUGGAUAAAAAGUUGUGGUUAUAGCGAACU---UUCAUUUGCGAUUAAACUGCUUGCGGAUUCAAUAUCCGU ..((((..(((((............)))))..)))).........(..(((...(((((..---....))))).....)))..)..((((((.....)))))) ( -29.50) >consensus UUUCCACUGCCCUCCAAGUUAACCAGGGGCUCUGGAUAAAAAGUUGUGGUUAUAGUGAAUUCACUCCAUUUGGCGAUAAACUGCCAGCGGACUU__UUCCCGU ..((((..(((((............)))))..)))).........(..(((...........................)))..)..((((.........)))) (-18.70 = -18.48 + -0.22)

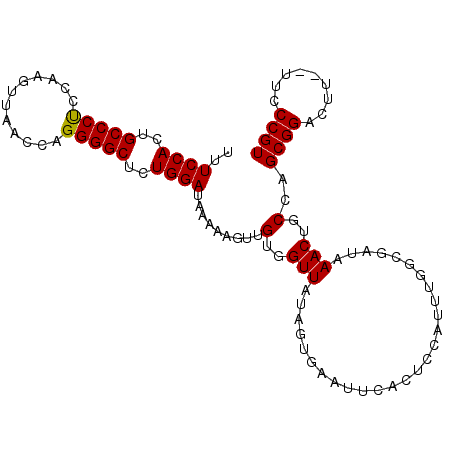
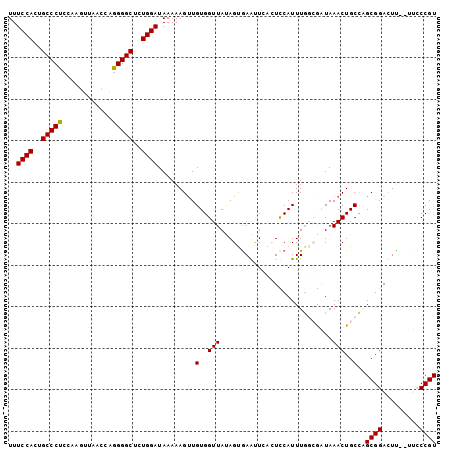
| Location | 620,819 – 620,925 |
|---|---|
| Length | 106 |
| Sequences | 6 |
| Columns | 112 |
| Reading direction | reverse |
| Mean pairwise identity | 76.50 |
| Mean single sequence MFE | -27.51 |
| Consensus MFE | -20.71 |
| Energy contribution | -20.49 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.14 |
| Mean z-score | -0.96 |
| Structure conservation index | 0.75 |
| SVM decision value | 0.33 |
| SVM RNA-class probability | 0.691690 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 620819 106 - 22224390 ---UCGAAAUCUGUAUA-G-UCUUACCGCACCAAGCUUUCCACUGCCCUCCAAGUUAA-CCAGGGGCUCUGGAUAAAAAGUUGUGGUUAUAGUGAGUUCACUCCAUUUUGCG ---.((((((..((...-.-.(((((...(((((((((((((..(((((.........-...)))))..))))....))))).))))....)))))...))...)))))).. ( -24.60) >DroPse_CAF1 10015 100 - 1 CAGCUGAACUCUG--------GCUACCGCACCAAGCUUUCCAUUGCCCCCAAUCAUAG-CCUGGGGCUUUGGAUAAAAAGUUGUGGUUAUAGCGAACU---UUCAUUUGCGA .((((.......)--------)))((((((....((((((((..(((((.........-...)))))..))))....))))))))))....(((((..---....))))).. ( -29.20) >DroSec_CAF1 5062 108 - 1 CGAUCGAAAUCAGUAUA-G-UCUUACCGCACCAAGCUUUCCACUGCCCUGCAAGUUAA-CCAGGGGCUCUGGAUAAAAAG-UGUGGUUAUAGUGAAUUGACUCCAUUUGGCG .((((((..(((.((((-.-....(((((((.......((((..(((((.........-...)))))..))))......)-)))))))))).))).)))).))......... ( -29.12) >DroEre_CAF1 5432 109 - 1 CCAACCAAAUCAGUAUA-G-UCUUACCGCACCAAGCUUUCCAUUGCCCUCCAAGUUAA-CUAGGGGCUCUGGAUAAAAAGUUGUGGUUACAGUGAGGUCACUCCAUUUGGCG ....((((((.(((...-.-((((((...(((((((((((((..(((((...((....-)).)))))..))))....))))).))))....))))))..)))..)))))).. ( -29.30) >DroAna_CAF1 5253 108 - 1 ---UCAGAAUCAGAAUCAGUUGCUACCGCACCAAGCUUUUCAUUGCCCGCCAACUCAUCCCAGGGGCUUUGGAUAAAAAGUUGUGGUUAUAGCGAAGCCUCUCCAUUU-GCG ---.((((...(((.....(((((((((((....((((((((..((((...............))))..))))....))))))))))...)))))....)))...)))-).. ( -23.66) >DroPer_CAF1 17985 100 - 1 CAGCUGAACUCUG--------GCUACCGCACCAAGCUUUCCAUUGCCCCCAAUCAUAG-CCUGGGGCUUUGGAUAAAAAGUUGUGGUUAUAGCGAACU---UUCAUUUGCGA .((((.......)--------)))((((((....((((((((..(((((.........-...)))))..))))....))))))))))....(((((..---....))))).. ( -29.20) >consensus C__UCGAAAUCAGUAUA_G_UCCUACCGCACCAAGCUUUCCAUUGCCCUCCAACUUAA_CCAGGGGCUCUGGAUAAAAAGUUGUGGUUAUAGCGAACUCACUCCAUUUGGCG ..........................((((((((((((((((..(((((.............)))))..))))....))))).))))....))).................. (-20.71 = -20.49 + -0.22)
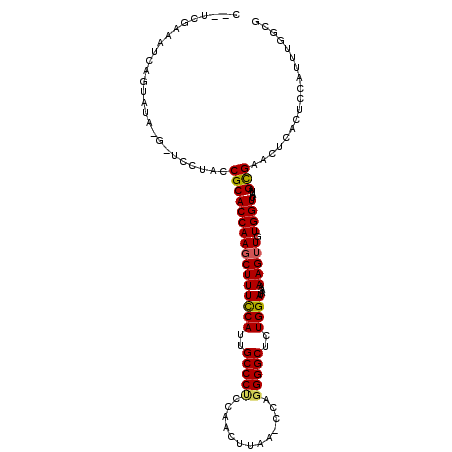


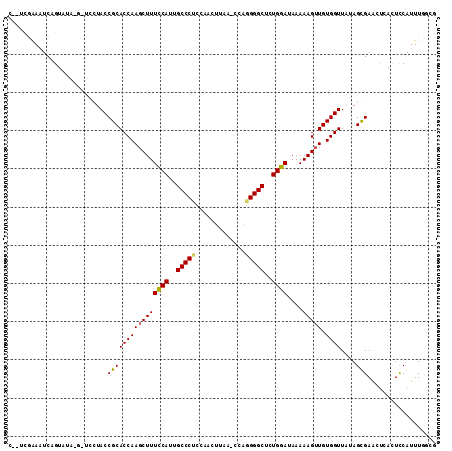
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:37:55 2006