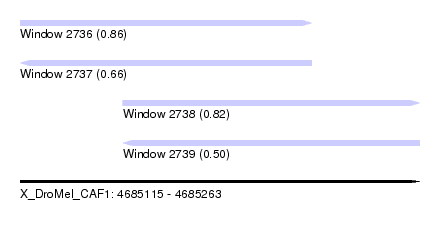

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 4,685,115 – 4,685,263 |
| Length | 148 |
| Max. P | 0.860761 |

| Location | 4,685,115 – 4,685,223 |
|---|---|
| Length | 108 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 89.47 |
| Mean single sequence MFE | -34.58 |
| Consensus MFE | -29.78 |
| Energy contribution | -29.72 |
| Covariance contribution | -0.06 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.80 |
| Structure conservation index | 0.86 |
| SVM decision value | 0.83 |
| SVM RNA-class probability | 0.860761 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
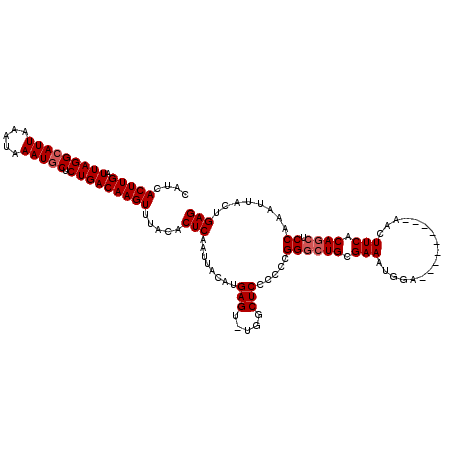
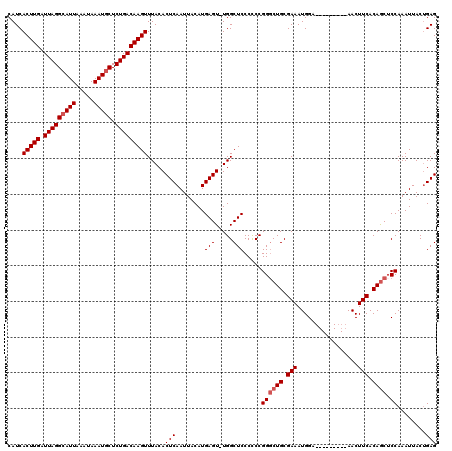
>X_DroMel_CAF1 4685115 108 + 22224390 CUGCGAA--AUGUCGCUCACAUGCAAUUAAAUUCGCUCAGCUCAGUAAUUUGGAGCUGUGAAGUU---------UCCAUUUCGCAGCCCGGGGGGAGCCA-ACUCAUGUAAUUGAGUGUA ..(((((--..((.((......)).))....)))))...((((.....(((((.((((((((((.---------...)))))))))))))))..))))..-(((((......)))))... ( -35.10) >DroSec_CAF1 38739 108 + 1 CUGCGAA--AUGUUGCUCACAUCCAAUUAAAUUCGCUCAGCUCCGUAAUUUGGAGCUGUGAAGUU---------UCCAUUUCGCAGCCCGGGGGGAGCCA-ACUCAUGUAAUUGAGUGUA ..(((((--..((((........))))....)))))...(((((....(((((.((((((((((.---------...))))))))))))))).)))))..-(((((......)))))... ( -36.90) >DroEre_CAF1 39135 108 + 1 --ACGAACGAUGUCGCUCACAUGCAAUUAAAUUUGCUCAGCUCAGUAAUUUGGCGUUGUGAAGUU---------UCCAUUUCGCAGCCCGGGGGGAGCCA-ACUCAUGUAAUUGAGUGUA --..((.((....)).))....((((......))))...((((.....(((((.((((((((((.---------...)))))))))))))))..))))..-(((((......)))))... ( -31.30) >DroYak_CAF1 35946 118 + 1 CUGCGAA--AUGUCGCUCACAUGCAAUUAAAUUCGCUCAGCUCAGUAAUUUGGAGCUGUGAAGUUGCCAAAGUUGCCAUUUCGCAGGCCGGAGGGAGCCAAACUCAUGUAAUUGAGUGUA (((((((--(((..((......(((((....(((((..(((((.........))))))))))))))).......))))))))))))...((......))..(((((......)))))... ( -35.02) >consensus CUGCGAA__AUGUCGCUCACAUGCAAUUAAAUUCGCUCAGCUCAGUAAUUUGGAGCUGUGAAGUU_________UCCAUUUCGCAGCCCGGGGGGAGCCA_ACUCAUGUAAUUGAGUGUA ..((((......))))..((((.((((((..........((((.....(((((.((((((((((.............)))))))))))))))..))))..........)))))).)))). (-29.78 = -29.72 + -0.06)
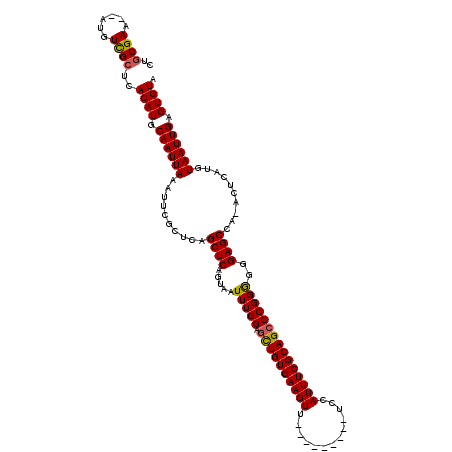


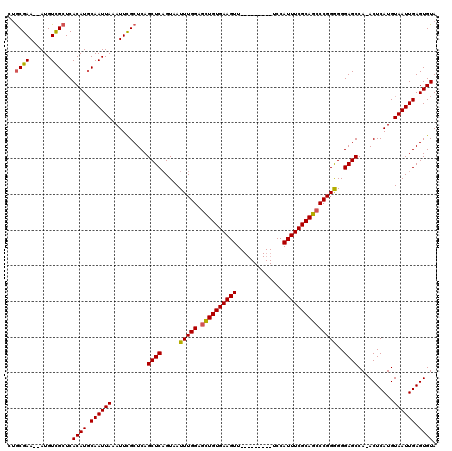
| Location | 4,685,115 – 4,685,223 |
|---|---|
| Length | 108 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 89.47 |
| Mean single sequence MFE | -32.26 |
| Consensus MFE | -23.09 |
| Energy contribution | -23.65 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.88 |
| Structure conservation index | 0.72 |
| SVM decision value | 0.26 |
| SVM RNA-class probability | 0.657590 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 4685115 108 - 22224390 UACACUCAAUUACAUGAGU-UGGCUCCCCCCGGGCUGCGAAAUGGA---------AACUUCACAGCUCCAAAUUACUGAGCUGAGCGAAUUUAAUUGCAUGUGAGCGACAU--UUCGCAG ...(((((......)))))-.(((((.....)))))((((((((..---------...((((((((((.........)))))..((((......))))..)))))...)))--))))).. ( -32.60) >DroSec_CAF1 38739 108 - 1 UACACUCAAUUACAUGAGU-UGGCUCCCCCCGGGCUGCGAAAUGGA---------AACUUCACAGCUCCAAAUUACGGAGCUGAGCGAAUUUAAUUGGAUGUGAGCAACAU--UUCGCAG ...(((((......)))))-.(((((.....)))))((((((((..---------.......(((((((.......))))))).((..((((....))))....))..)))--))))).. ( -34.70) >DroEre_CAF1 39135 108 - 1 UACACUCAAUUACAUGAGU-UGGCUCCCCCCGGGCUGCGAAAUGGA---------AACUUCACAACGCCAAAUUACUGAGCUGAGCAAAUUUAAUUGCAUGUGAGCGACAUCGUUCGU-- .(((..((((((.((..((-((((((......(((((.(((..(..---------..)))).))..)))........))))).)))..)).))))))..)))(((((....)))))..-- ( -27.04) >DroYak_CAF1 35946 118 - 1 UACACUCAAUUACAUGAGUUUGGCUCCCUCCGGCCUGCGAAAUGGCAACUUUGGCAACUUCACAGCUCCAAAUUACUGAGCUGAGCGAAUUUAAUUGCAUGUGAGCGACAU--UUCGCAG ...(((((......)))))..((((......))))(((((((((((......(....)....((((((.........)))))).))........((((......)))))))--)))))). ( -34.70) >consensus UACACUCAAUUACAUGAGU_UGGCUCCCCCCGGGCUGCGAAAUGGA_________AACUUCACAGCUCCAAAUUACUGAGCUGAGCGAAUUUAAUUGCAUGUGAGCGACAU__UUCGCAG ((((..((((((.((..((.((((((.....((((((.(((.................))).)))).))........)))))).))..)).))))))..)))).((((......)))).. (-23.09 = -23.65 + 0.56)
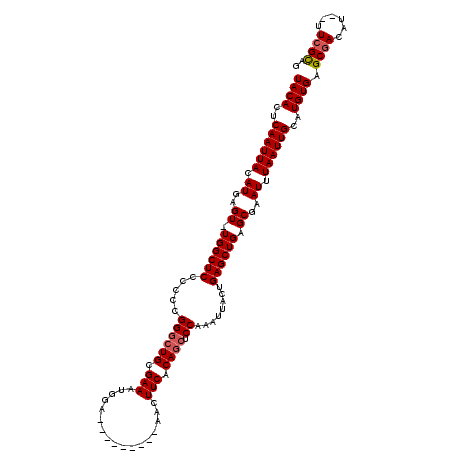


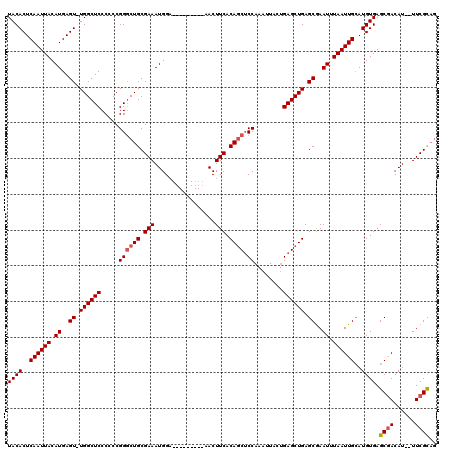
| Location | 4,685,153 – 4,685,263 |
|---|---|
| Length | 110 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 92.61 |
| Mean single sequence MFE | -32.80 |
| Consensus MFE | -28.75 |
| Energy contribution | -28.87 |
| Covariance contribution | 0.12 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.73 |
| Structure conservation index | 0.88 |
| SVM decision value | 0.67 |
| SVM RNA-class probability | 0.816841 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 4685153 110 + 22224390 CUCAGUAAUUUGGAGCUGUGAAGUU---------UCCAUUUCGCAGCCCGGGGGGAGCCA-ACUCAUGUAAUUGAGUGUAAACUUGUCAGAGAAUUUAUUUAAUGCCUAAUCAAGUGAUG (((......((((.((((((((((.---------...))))))))))))))((.(((...-(((((......))))).....))).)).)))...((((((.((.....)).)))))).. ( -28.70) >DroSec_CAF1 38777 110 + 1 CUCCGUAAUUUGGAGCUGUGAAGUU---------UCCAUUUCGCAGCCCGGGGGGAGCCA-ACUCAUGUAAUUGAGUGUAAACUUGUCAGAGCAUUUAUUUAAUGCCUAAUCAAGUGAUG ((((....(((((.((((((((((.---------...))))))))))))))).))))...-(((((......)))))....(((((..((.(((((.....)))))))...))))).... ( -36.60) >DroEre_CAF1 39173 110 + 1 CUCAGUAAUUUGGCGUUGUGAAGUU---------UCCAUUUCGCAGCCCGGGGGGAGCCA-ACUCAUGUAAUUGAGUGUAAACUUGUCAGAGCAUUUAUUUAAUGCCUAAUCAAGUGAUG ..........((((((((((((((.---------...))))))))))((....)).))))-(((((......)))))....(((((..((.(((((.....)))))))...))))).... ( -32.10) >DroYak_CAF1 35984 120 + 1 CUCAGUAAUUUGGAGCUGUGAAGUUGCCAAAGUUGCCAUUUCGCAGGCCGGAGGGAGCCAAACUCAUGUAAUUGAGUGUAAACUUGUCAGAGCAUUUAUUUAAUGCCUAAUCAAGUGAUG (((.....(((((..(((((((((.((.......)).))))))))).)))))..)))....(((((......)))))....(((((..((.(((((.....)))))))...))))).... ( -33.80) >consensus CUCAGUAAUUUGGAGCUGUGAAGUU_________UCCAUUUCGCAGCCCGGGGGGAGCCA_ACUCAUGUAAUUGAGUGUAAACUUGUCAGAGCAUUUAUUUAAUGCCUAAUCAAGUGAUG (((.....(((((.((((((((((.............)))))))))))))))..)))....(((((......)))))....(((((..((.(((((.....)))))))...))))).... (-28.75 = -28.87 + 0.12)

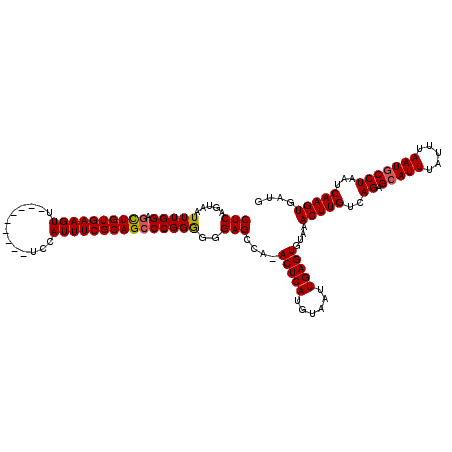
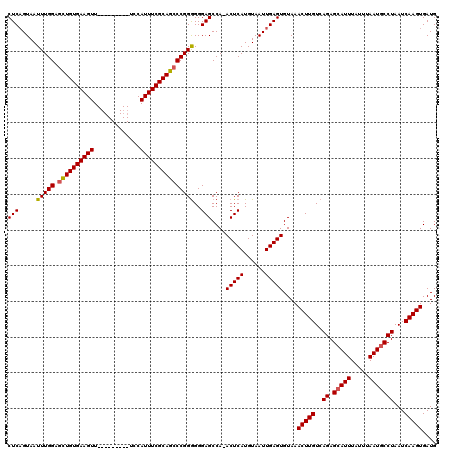
| Location | 4,685,153 – 4,685,263 |
|---|---|
| Length | 110 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 92.61 |
| Mean single sequence MFE | -26.96 |
| Consensus MFE | -20.25 |
| Energy contribution | -21.01 |
| Covariance contribution | 0.75 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.79 |
| Structure conservation index | 0.75 |
| SVM decision value | -0.06 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 4685153 110 - 22224390 CAUCACUUGAUUAGGCAUUAAAUAAAUUCUCUGACAAGUUUACACUCAAUUACAUGAGU-UGGCUCCCCCCGGGCUGCGAAAUGGA---------AACUUCACAGCUCCAAAUUACUGAG ....(((((.(((((.(((.....))).).))))))))).....((((.......(((.-...))).....((((((.(((..(..---------..)))).)))).)).......)))) ( -22.70) >DroSec_CAF1 38777 110 - 1 CAUCACUUGAUUAGGCAUUAAAUAAAUGCUCUGACAAGUUUACACUCAAUUACAUGAGU-UGGCUCCCCCCGGGCUGCGAAAUGGA---------AACUUCACAGCUCCAAAUUACGGAG ....(((((.(((((((((.....))))).)))))))))...((((((......)))).-)).((((....((((((.(((..(..---------..)))).))))))........)))) ( -30.40) >DroEre_CAF1 39173 110 - 1 CAUCACUUGAUUAGGCAUUAAAUAAAUGCUCUGACAAGUUUACACUCAAUUACAUGAGU-UGGCUCCCCCCGGGCUGCGAAAUGGA---------AACUUCACAACGCCAAAUUACUGAG ....(((((.(((((((((.....))))).))))))))).....((((......((.((-((((..((....))..))(((..(..---------..)))).))))..))......)))) ( -26.00) >DroYak_CAF1 35984 120 - 1 CAUCACUUGAUUAGGCAUUAAAUAAAUGCUCUGACAAGUUUACACUCAAUUACAUGAGUUUGGCUCCCUCCGGCCUGCGAAAUGGCAACUUUGGCAACUUCACAGCUCCAAAUUACUGAG ....(((((.(((((((((.....))))).))))))))).....((((.......(((((.((((......))))(((.(((.(....)))).))).......)))))........)))) ( -28.76) >consensus CAUCACUUGAUUAGGCAUUAAAUAAAUGCUCUGACAAGUUUACACUCAAUUACAUGAGU_UGGCUCCCCCCGGGCUGCGAAAUGGA_________AACUUCACAGCUCCAAAUUACUGAG ....(((((.(((((((((.....))))).))))))))).....(((........(((.....))).....((((((.(((.................))).)))).))........))) (-20.25 = -21.01 + 0.75)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:16:10 2006