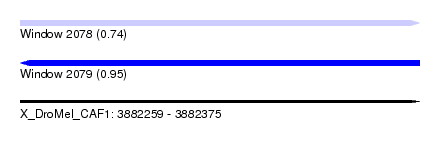

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 3,882,259 – 3,882,375 |
| Length | 116 |
| Max. P | 0.953923 |

| Location | 3,882,259 – 3,882,375 |
|---|---|
| Length | 116 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 76.28 |
| Mean single sequence MFE | -21.95 |
| Consensus MFE | -11.32 |
| Energy contribution | -11.57 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.88 |
| Structure conservation index | 0.52 |
| SVM decision value | 0.46 |
| SVM RNA-class probability | 0.744804 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
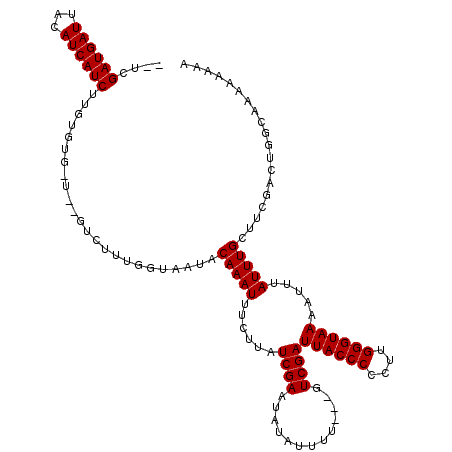
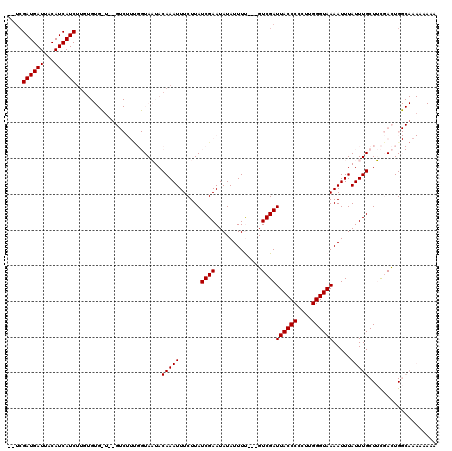
>X_DroMel_CAF1 3882259 116 + 22224390 -CUCGAUGAUUACAUCAUCUUGUGUGCUGUGUCUUUGGUAAUACAAAUUUCUUAUCGAAUAUAUUUU---GUCGAUUACCCCCUUGGGUAAAAUUUAUUUGCUUCGACUGGCAAAAAAAA -...((((((...)))))).....(((((.(((...(((((...((((((...(((((.((.....)---))))))(((((....)))))))))))..)))))..))))))))....... ( -24.30) >DroVir_CAF1 62865 91 + 1 AAUCGAUGAUUACAUCAUCUUG----------------UAUUACAAAUUUCUUAUCGAAUAUAUUUC---GUCGAUUACCCCCUUGGGUAAAAUUUAUUUGCUGUCAUUG---------- ...(((((((((((......))----------------))...(((((.......((((.....)))---)....((((((....)))))).....)))))..)))))))---------- ( -16.40) >DroGri_CAF1 52441 91 + 1 AAUCGAUGAUUACAUCAUCUUG----------------UAUUACAAAUUUCUUAUCGAAUAUAUUUC---GUCGAUUACCCCCUUGGGUAAAAUUUAUUUGCUGUCAUUG---------- ...(((((((((((......))----------------))...(((((.......((((.....)))---)....((((((....)))))).....)))))..)))))))---------- ( -16.40) >DroSec_CAF1 4 93 + 1 ----------------------UGUGCU--GUCUUUGGUAAUACAAAUUUCUUAUCGAAUAUAUUUU---GUCGAUUACCCCCUUGGGUAAAAUUUAUUUGCUUCGACUGGCAAAAAAAA ----------------------..((((--(((...(((((...((((((...(((((.((.....)---))))))(((((....)))))))))))..)))))..))).))))....... ( -19.50) >DroEre_CAF1 422 118 + 1 --UCGAUGAUUACAUCAUCUUGUGUGUGUUGUCUUCGGUAAUACAAAUUUCUUAUCGAAUAUAUUUUGUUGUCGAUUACCCCCUUGGGUAAAAUUUAUUUGCUUCGACUGGCACACAAAA --..((((((...))))))((((((((..(((.((((((((..........)))))))).))).......(((((((((((....))))))............)))))..)))))))).. ( -31.20) >DroYak_CAF1 2531 116 + 1 -CUCGAUGAUUACAUCAUCUUGUGUGUUGUGUCUUUGGUAAUACAAAUUUCUUAUCGAAUAUAUUUU---GUCGAUUACCCCCUUGGGUAAAAUUUAUUUGCUUCGACUGGCAAAAAAAA -.((((..(.(((((......)))))..((((.((((((((..........)))))))).)))).).---.))))((((((....))))))..(((.((((((......)))))).))). ( -23.90) >consensus __UCGAUGAUUACAUCAUCUUGUGUG_U__GUCUUUGGUAAUACAAAUUUCUUAUCGAAUAUAUUUU___GUCGAUUACCCCCUUGGGUAAAAUUUAUUUGCUUCGACUGGCAAAAAAAA ....((((((...))))))........................(((((......((((.............))))((((((....)))))).....)))))................... (-11.32 = -11.57 + 0.25)

| Location | 3,882,259 – 3,882,375 |
|---|---|
| Length | 116 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 76.28 |
| Mean single sequence MFE | -20.20 |
| Consensus MFE | -15.15 |
| Energy contribution | -15.40 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.68 |
| Structure conservation index | 0.75 |
| SVM decision value | 1.44 |
| SVM RNA-class probability | 0.953923 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
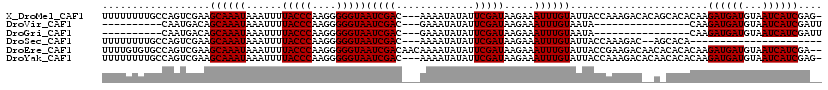
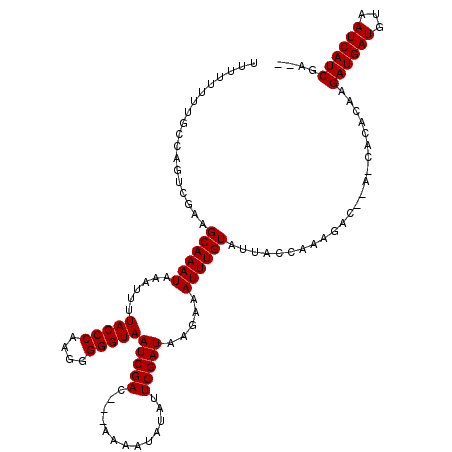
>X_DroMel_CAF1 3882259 116 - 22224390 UUUUUUUUGCCAGUCGAAGCAAAUAAAUUUUACCCAAGGGGGUAAUCGAC---AAAAUAUAUUCGAUAAGAAAUUUGUAUUACCAAAGACACAGCACACAAGAUGAUGUAAUCAUCGAG- .......(((..(((.......(((((((((((((....)))).(((((.---.........)))))..))))))))).........)))...))).....((((((...))))))...- ( -20.99) >DroVir_CAF1 62865 91 - 1 ----------CAAUGACAGCAAAUAAAUUUUACCCAAGGGGGUAAUCGAC---GAAAUAUAUUCGAUAAGAAAUUUGUAAUA----------------CAAGAUGAUGUAAUCAUCGAUU ----------........((((((.....((((((....))))))....(---(((.....)))).......))))))....----------------...((((((...)))))).... ( -17.80) >DroGri_CAF1 52441 91 - 1 ----------CAAUGACAGCAAAUAAAUUUUACCCAAGGGGGUAAUCGAC---GAAAUAUAUUCGAUAAGAAAUUUGUAAUA----------------CAAGAUGAUGUAAUCAUCGAUU ----------........((((((.....((((((....))))))....(---(((.....)))).......))))))....----------------...((((((...)))))).... ( -17.80) >DroSec_CAF1 4 93 - 1 UUUUUUUUGCCAGUCGAAGCAAAUAAAUUUUACCCAAGGGGGUAAUCGAC---AAAAUAUAUUCGAUAAGAAAUUUGUAUUACCAAAGAC--AGCACA---------------------- .......(((..(((.......(((((((((((((....)))).(((((.---.........)))))..))))))))).........)))--.)))..---------------------- ( -16.39) >DroEre_CAF1 422 118 - 1 UUUUGUGUGCCAGUCGAAGCAAAUAAAUUUUACCCAAGGGGGUAAUCGACAACAAAAUAUAUUCGAUAAGAAAUUUGUAUUACCGAAGACAACACACACAAGAUGAUGUAAUCAUCGA-- (((((((((...(((((............((((((....)))))))))))...........((((.(((..........))).)))).......)))))))))(((((....))))).-- ( -27.40) >DroYak_CAF1 2531 116 - 1 UUUUUUUUGCCAGUCGAAGCAAAUAAAUUUUACCCAAGGGGGUAAUCGAC---AAAAUAUAUUCGAUAAGAAAUUUGUAUUACCAAAGACACAACACACAAGAUGAUGUAAUCAUCGAG- ...((((((...(((((............((((((....)))))))))))---..(((((((((.....)))...))))))..))))))............((((((...))))))...- ( -20.80) >consensus UUUUUUUUGCCAGUCGAAGCAAAUAAAUUUUACCCAAGGGGGUAAUCGAC___AAAAUAUAUUCGAUAAGAAAUUUGUAUUACCAAAGAC__A_CACACAAGAUGAUGUAAUCAUCGA__ ..................((((((......(((((....)))))(((((.............))))).....)))))).......................((((((...)))))).... (-15.15 = -15.40 + 0.25)
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:06:03 2006