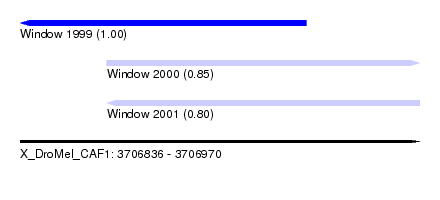
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 3,706,836 – 3,706,970 |
| Length | 134 |
| Max. P | 0.999986 |

| Location | 3,706,836 – 3,706,932 |
|---|---|
| Length | 96 |
| Sequences | 5 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 95.62 |
| Mean single sequence MFE | -40.52 |
| Consensus MFE | -39.22 |
| Energy contribution | -39.38 |
| Covariance contribution | 0.16 |
| Combinations/Pair | 1.09 |
| Mean z-score | -3.14 |
| Structure conservation index | 0.97 |
| SVM decision value | 5.40 |
| SVM RNA-class probability | 0.999986 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 3706836 96 - 22224390 GUCUUCGAGGCAUCGAAGACGCGGCUAUCCGAAGUAGCGAAAGUGGUGCGAGUGGUGCAACUUGUGACACUGCCAGCGGAGUGGAUGAGCCCCAUA ((((((((....))))))))(.(((((((((.....((...((((..((((((......))))))..))))....))....))))).)))).)... ( -40.30) >DroSec_CAF1 46506 96 - 1 GUCUUCGAGGCAUCGAAGACGCGGCUAUCUGAAAGAGCGAAAGUGGUGCGAGUGGUGCAACUUGUGCCACUGCCAGCGGAGUGGAUGAGCCCCAUA ((((((((....))))))))(.(((((((((.....((...(((((..(((((......)))))..)))))....))....))))).)))).)... ( -43.50) >DroSim_CAF1 43760 96 - 1 GUCUUCGAGGCAUCGAAGACGCGGCUAUCUGAAAGAGCGAAAGUGGUGCGAGUGGUGCAACUUGUGCCACUGCCAGCGGAGUGGAUGAGCCCCAUA ((((((((....))))))))(.(((((((((.....((...(((((..(((((......)))))..)))))....))....))))).)))).)... ( -43.50) >DroEre_CAF1 46048 96 - 1 GUCUUCGAGGCAUCGAAGACGCGGCUGUCUGAAAUAGCGAAAGUGGUGCGAGUGGUGCAACUUGUGCCACUGCCAGCGGAGUGGAUGAGUCCCAUA ((((((((....))))))))(.(((((((((.....((...(((((..(((((......)))))..)))))....))....))))).)))).)... ( -40.80) >DroYak_CAF1 43443 96 - 1 GUCUUCGAGGCAUCGAAGAAGCGGCUGUCUGAAAGAGCGAAAGUGGUGCGAGUGGUGCAACUUGAGCCACUGCCAGCGGAGUGGAUGAUUCCCAUA .(((((((....))))))).(((((((..(....)..))..((((((.(((((......))))).))))))))).))(((((.....))))).... ( -34.50) >consensus GUCUUCGAGGCAUCGAAGACGCGGCUAUCUGAAAGAGCGAAAGUGGUGCGAGUGGUGCAACUUGUGCCACUGCCAGCGGAGUGGAUGAGCCCCAUA ((((((((....))))))))(.(((((((((.....((...((((((((((((......))))))))))))....))....))))).)))).)... (-39.22 = -39.38 + 0.16)

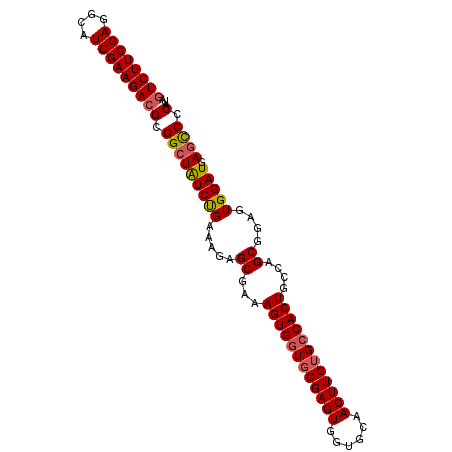
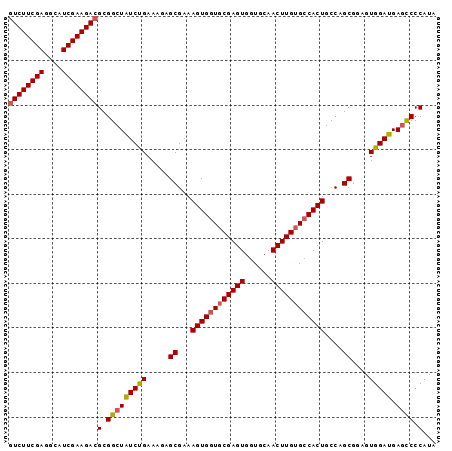
| Location | 3,706,865 – 3,706,970 |
|---|---|
| Length | 105 |
| Sequences | 5 |
| Columns | 107 |
| Reading direction | forward |
| Mean pairwise identity | 92.25 |
| Mean single sequence MFE | -29.86 |
| Consensus MFE | -25.22 |
| Energy contribution | -25.46 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.54 |
| Structure conservation index | 0.84 |
| SVM decision value | 0.80 |
| SVM RNA-class probability | 0.852892 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
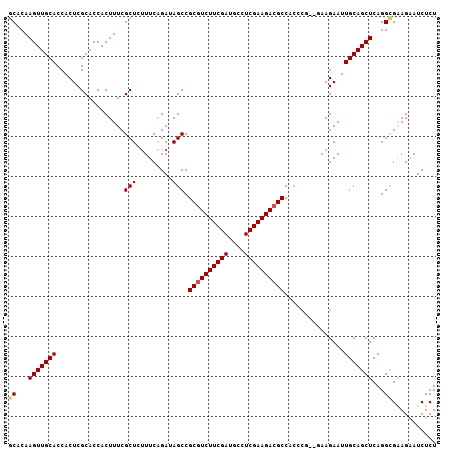
>X_DroMel_CAF1 3706865 105 + 22224390 UCACAAGUUGCACCACUCGCACCACUUUCGCUACUUCGGAUAGCCGCGUCUUCGAUGCCUCGAAGACGCCACCCU--UAAGAAUUGCAGCUCAGGCGAAGAAUCUCU ((.(.(((((((.................(((((....).)))).((((((((((....))))))))))......--.......)))))))...).))......... ( -31.10) >DroSec_CAF1 46535 107 + 1 GCACAAGUUGCACCACUCGCACCACUUUCGCUCUUUCAGAUAGCCGCGUCUUCGAUGCCUCGAAGACGCCAGAAUUGCAAGAAUUGCAGCUCAGGCGAAGAAUCUCU ((..(((((((.......)))..))))..))(((((......(((((((((((((....))))))))))..((.(((((.....))))).)).))))))))...... ( -32.80) >DroSim_CAF1 43789 105 + 1 GCACAAGUUGCACCACUCGCACCACUUUCGCUCUUUCAGAUAGCCGCGUCUUCGAUGCCUCGAAGACGCCACCCC--UAAGAAUUGCAGCUCAGGCGAAGAAUCUCU ((...(((((((.................(((.........))).((((((((((....))))))))))......--.......)))))))...))........... ( -28.50) >DroEre_CAF1 46077 105 + 1 GCACAAGUUGCACCACUCGCACCACUUUCGCUAUUUCAGACAGCCGCGUCUUCGAUGCCUCGAAGACGCCACCCG--GAAGAAUUGCAGCUCAGGUGAAGAAUCUCU .(((.(((((((....((...((......(((.........))).((((((((((....)))))))))).....)--)..))..)))))))...))).......... ( -30.50) >DroYak_CAF1 43472 105 + 1 GCUCAAGUUGCACCACUCGCACCACUUUCGCUCUUUCAGACAGCCGCUUCUUCGAUGCCUCGAAGACGCCACCCG--GAAGAAUUGCAGCUCAGGUGGAGGAUCUCU (((..(((((((......((.........))((((((.(.....)((.(((((((....))))))).)).....)--)))))..)))))))..)))((((...)))) ( -26.40) >consensus GCACAAGUUGCACCACUCGCACCACUUUCGCUCUUUCAGAUAGCCGCGUCUUCGAUGCCUCGAAGACGCCACCCG__GAAGAAUUGCAGCUCAGGCGAAGAAUCUCU ((...(((((((.................(((.........))).((((((((((....))))))))))...............)))))))...))........... (-25.22 = -25.46 + 0.24)

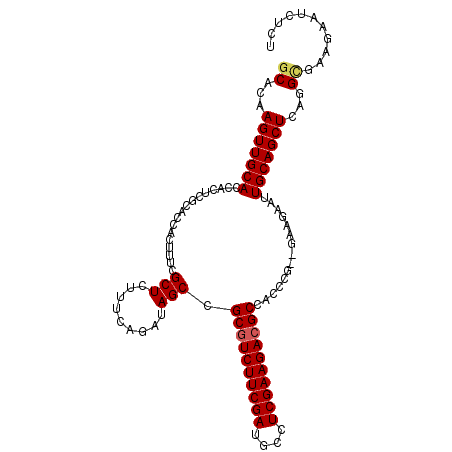
| Location | 3,706,865 – 3,706,970 |
|---|---|
| Length | 105 |
| Sequences | 5 |
| Columns | 107 |
| Reading direction | reverse |
| Mean pairwise identity | 92.25 |
| Mean single sequence MFE | -38.19 |
| Consensus MFE | -31.26 |
| Energy contribution | -31.82 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.58 |
| Structure conservation index | 0.82 |
| SVM decision value | 0.63 |
| SVM RNA-class probability | 0.803725 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
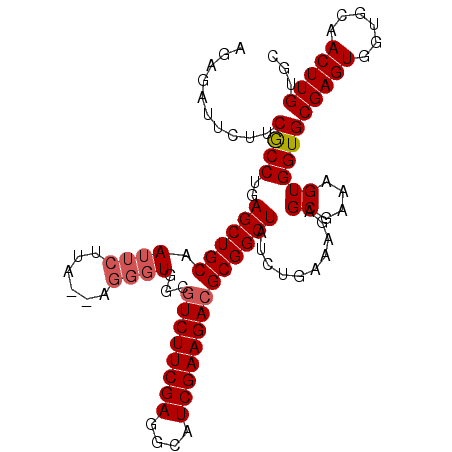
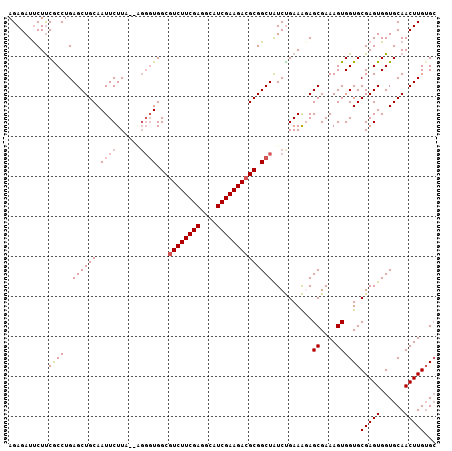
>X_DroMel_CAF1 3706865 105 - 22224390 AGAGAUUCUUCGCCUGAGCUGCAAUUCUUA--AGGGUGGCGUCUUCGAGGCAUCGAAGACGCGGCUAUCCGAAGUAGCGAAAGUGGUGCGAGUGGUGCAACUUGUGA .........((((.(..(((((..(((...--..(((.((((((((((....)))))))))).)))....))))))))..).)))).((((((......)))))).. ( -38.70) >DroSec_CAF1 46535 107 - 1 AGAGAUUCUUCGCCUGAGCUGCAAUUCUUGCAAUUCUGGCGUCUUCGAGGCAUCGAAGACGCGGCUAUCUGAAAGAGCGAAAGUGGUGCGAGUGGUGCAACUUGUGC ..((((.....(((.(((.((((.....)))).)))..((((((((((....))))))))))))).))))......((....)).(..(((((......)))))..) ( -38.90) >DroSim_CAF1 43789 105 - 1 AGAGAUUCUUCGCCUGAGCUGCAAUUCUUA--GGGGUGGCGUCUUCGAGGCAUCGAAGACGCGGCUAUCUGAAAGAGCGAAAGUGGUGCGAGUGGUGCAACUUGUGC ......(((((.((((((........))))--))(((.((((((((((....)))))))))).)))....).))))((....)).(..(((((......)))))..) ( -40.60) >DroEre_CAF1 46077 105 - 1 AGAGAUUCUUCACCUGAGCUGCAAUUCUUC--CGGGUGGCGUCUUCGAGGCAUCGAAGACGCGGCUGUCUGAAAUAGCGAAAGUGGUGCGAGUGGUGCAACUUGUGC .....(((..((((((..............--))))))((((((((((....)))))))))).(((((.....))))))))....(..(((((......)))))..) ( -37.44) >DroYak_CAF1 43472 105 - 1 AGAGAUCCUCCACCUGAGCUGCAAUUCUUC--CGGGUGGCGUCUUCGAGGCAUCGAAGAAGCGGCUGUCUGAAAGAGCGAAAGUGGUGCGAGUGGUGCAACUUGAGC ....((((((((((...........((((.--((((..((.(((((((....)))))))....))..)))).))))((....)))))).))).)))........... ( -35.30) >consensus AGAGAUUCUUCGCCUGAGCUGCAAUUCUUA__AGGGUGGCGUCUUCGAGGCAUCGAAGACGCGGCUAUCUGAAAGAGCGAAAGUGGUGCGAGUGGUGCAACUUGUGC ..........((((..((((((.((((......))))...((((((((....))))))))))))))..........((....))))))(((((......)))))... (-31.26 = -31.82 + 0.56)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:04:51 2006