Publications - Supplemental material
Please find below supplemental material corresponding to publications of our group. Currently, we list 132 supplements.
If you have problems accessing electronic information, please let us know:
You may use this URL to cite or link to us.
©NOTICE: All documents are copyrighted by the authors; If you would like to use all or a portion of any paper, please contact the author.
This supplement is also available at http://www.bioinf.uni-leipzig.de/publications/supplements/09-002You may use this URL to cite or link to us.
BIOINF 09-002: Complete HOX cluster characterization of the coelacanth provides further evidence for slow evolution of its genome
Chris T. Amemiya, Thomas P. Powers, Sonja J. Prohaska, Jane Grimwood, Jeremy Schmutz, Mark Dickson, Tsutomu Miyake, Michael A. Schoenborn, Richard M. Myers, Francis H. Ruddle, Peter F. Stadler
Figure 1 [PDF] (6K)
Figure 2 [PDF] (9K)
Figure 3 [PDF] (6K)
Figure 4 [PDF] (6K)
Figure 5 [PDF] (15K)
PNAS supplemental text
[PDF]
PNAS supplemental data
[PDF]
List of primer oligos
[PDF] List of clones
[PDF] Assembly: map of contigs
[PDF] Assembled Hox cluster sequences and annotations
Comparison of IGR length between human and coelacanth
[PDF]
VISTA plots
mVISTA overview
individual HOX clusters [PDF] List of tracker/dialign-2 footprints
[HoxA] [HoxB] [HoxC] [HoxD] PERL script mAAna_tropy.pl
source code as an archive [tar]
README [txt]
Consensus sequences of tRNA-like repetitive sequences
[fasta]
Relative Rate Tests on protein coding sequences
Extensive data with alignments: [PDF]
Short summary: [PDF] CNCN relative rate test
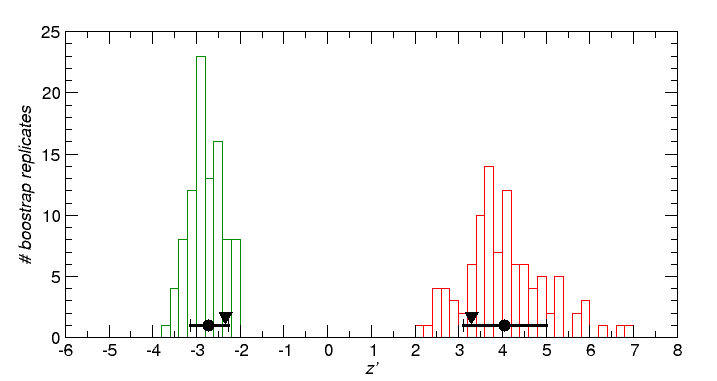
Outgroups are Heterodontus francisci HOXA and Polypterus senegalusHOXA. Latimeria is tested as variable "foreground" against the assumption of contant rates of both outgroups and the 2nd ingroup (negative z' values mean that Latimeria is slower), and as part of the "background" (positive z' values mean that the 2nd outgroup is faster than outgroups and Latimeria.
Figures of Main Text
Sequences
[PDF]
[PDF]
[PDF]
| Cluster | Genbank | fasta | Annotation | Peptides |
|---|---|---|---|---|
| HoxA | FJ497005 | [LmA.fa] | [text] | [fasta] |
| HoxB | FJ497006 | [LmB.fa] | [text] | [fasta] |
| HoxC | FJ497007 | [LmC.fa] | [text] | [fasta] |
| HoxD | FJ497008 | [LmD.fa] | [text] | [fasta] |
Noncoding Sequence Conservation
mVISTA overview
individual HOX clusters [PDF]
VISTA Evx region
HoxA/HoxD comparison
[PDF]
Comparison of A and D clusters
Reference: Human HoxA
[PDF]
Reference: Human HoxD
[PDF]
[HoxA] [HoxB] [HoxC] [HoxD]
source code as an archive [tar]
README [txt]
Repetitive Elements
[fasta]
Relative Rate Tests
Extensive data with alignments: [PDF]
Short summary: [PDF]
Outgroups are Heterodontus francisci HOXA and Polypterus senegalusHOXA. Latimeria is tested as variable "foreground" against the assumption of contant rates of both outgroups and the 2nd ingroup (negative z' values mean that Latimeria is slower), and as part of the "background" (positive z' values mean that the 2nd outgroup is faster than outgroups and Latimeria.
| ingroup | foreground | background |
|---|---|---|
| XtA | -3.112 *** | 4.939 *** |
| GgA | -3.687 *** | 6.119 *** |
| MdA | -2.325 ** | 3.301 *** |
| CfA | -1.735 * | 2.275 ** |
| MmA | -1.864 * | 2.494 ** |
| RnA | -1.840 * | 2.449 ** |
| HsA | -1.599 ns. | 2.051 ** |