bbq
Discovering cis-regulatory modules (clusters of transcription factor binding sites)
Axel Mosig, 2005.
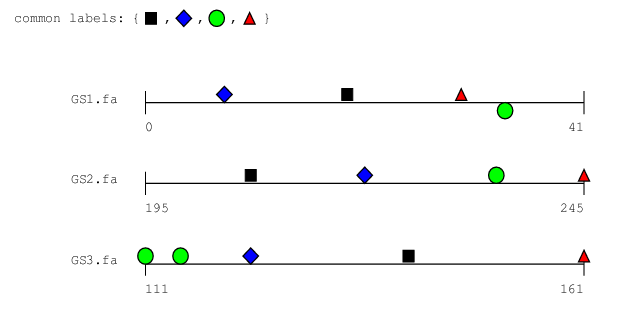
bbq is a command-line tool for discovering clusters
of transcription factor binding sites that occur simultaneously in
several genome sequences. Finding such clusters – which are
sometimes also referred to as cis-regulatory modules – is
done in a multiple-alignment-like fashion by solving a certain
combinatorial and geometric optimization problem, the so-called best
barbeque problem (explaining the name bbq). As opposed to
classical, typically dynamic programming based, alignment procedures,
the order of the binding sites' occurences can be arbitrarily
shuffled, so that bbq is the result of developing
completely new algorithms.
The implementation provided here supports numerous features such
as weighted sets, p-value based weighting schemes or computing the
best h solutions instead of only the best solution. Most
of these features are supported by two algorithms representing two
different approaches to solving the given optimization problem. Output
can be visualized in postscript.
bbq
source code release
[download]
Contains documented source code, tutorial and some example files.
Compilation requires a working installation of the GNU MP library!
(Configure gmp using ./configure --enable-cxx)
bbq
binary (built under Fedora)
[download]
For the impacient with a suitable Linux environment
Darwin (Mac OS X) binary
 bbq
bbq
[download]
For the impacient apple user; stand-alone version compiled under OS X 10.4
windows binary
 bbq.exe
bbq.exe
[download]
For the impacient windows user; stand-alone version compiled using MinGW. Although most features work fine, note that this build has not yet been tested in detail.
bbq
tutorial
[view]
A step-by-step introduction into using
bbq
...and why is it called bbq?
bbq
implementation documentation
[view]
Doxygen generated for those interested in every detail
bbq
publication preprint
[pdf] [ps]
How
bbq works