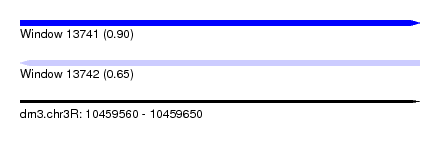

| Sequence ID | dm3.chr3R |
|---|---|
| Location | 10,459,560 – 10,459,650 |
| Length | 90 |
| Max. P | 0.902482 |

| Location | 10,459,560 – 10,459,650 |
|---|---|
| Length | 90 |
| Sequences | 8 |
| Columns | 112 |
| Reading direction | forward |
| Mean pairwise identity | 66.12 |
| Shannon entropy | 0.61324 |
| G+C content | 0.47234 |
| Mean single sequence MFE | -27.26 |
| Consensus MFE | -9.58 |
| Energy contribution | -9.66 |
| Covariance contribution | 0.08 |
| Combinations/Pair | 1.22 |
| Mean z-score | -2.07 |
| Structure conservation index | 0.35 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.16 |
| SVM RNA-class probability | 0.902482 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
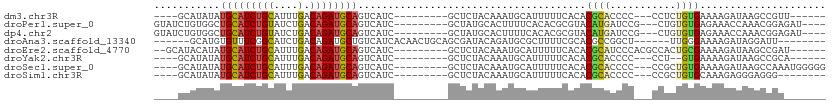
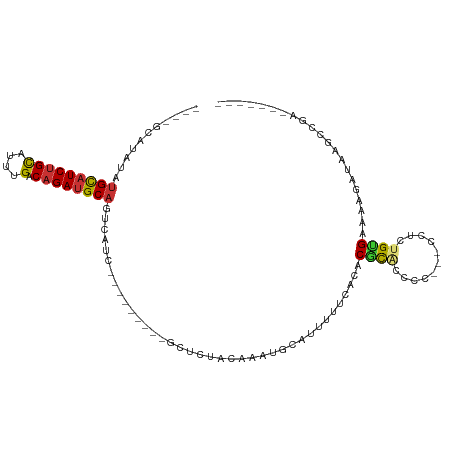
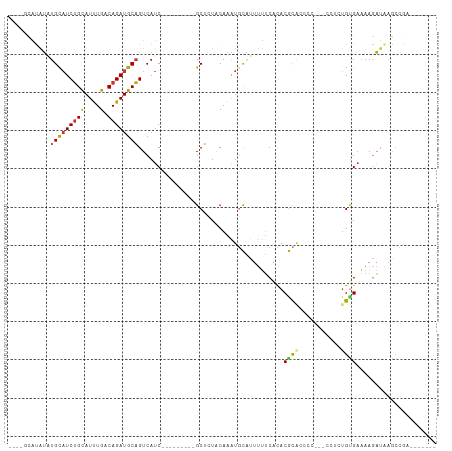
>dm3.chr3R 10459560 90 + 27905053 ----GCAUAUAUGCAUCUGCAUUUGACAGAUGCAGUCAUC---------GCUCUACAAAUGCAUUUUUCACACGCACCCC---CCUCUGUGAAAAGAUAAGCCGUU------ ----((.....(((((((((....).))))))))(((...---------((.........))..((((((((........---....)))))))))))..))....------ ( -21.30, z-score = -2.23, R) >droPer1.super_0 9622958 96 + 11822988 GUAUCUGUGGCUGCAUCUGUAUCUGACAGAUGCAGUCAUC---------GCUAUGCACUUUUCACACGCGUACAUGAUCCG---CUGUGUGAGAAACCAAACGGAGAU---- ...(((((((((((((((((.....))))))))))))...---------.........((((((((((((.........))---).))))))))).....)))))...---- ( -38.10, z-score = -3.87, R) >dp4.chr2 19942952 96 - 30794189 GUAUCUGUGGCUGCAUCUGUAUCUGACAGAUGCAGUCAUC---------GCUAUGCACUUUUCACACGCGUACAUGAUCCG---CUGUGUGAGAAACCAAACGGAGAU---- ...(((((((((((((((((.....))))))))))))...---------.........((((((((((((.........))---).))))))))).....)))))...---- ( -38.10, z-score = -3.87, R) >droAna3.scaffold_13340 22551446 92 - 23697760 ------GCAUGUGUUUCGGCAUCUGACAGAUGCUGUCAUCACAACUGCAGCGAUACAGAUGCGCUUUUCGCACGCCGGCU------UUGGGAAAAGAUAGGAUU-------- ------(((((((...(((((((.....)))))))....))))..))).((((...((.....))..))))...((..((------((....))))...))...-------- ( -26.20, z-score = -0.20, R) >droEre2.scaffold_4770 9992976 95 + 17746568 --GCAUACAUAUGCAUCUGCAUUUGACAGAUGCAGUCAUC---------GCUCUACAAAUGCAUUUUUCACACGCAUCCCACGCCACUGCGAAAAGAUAAGCCGAU------ --((.......(((((((((....).))))))))(((...---------((.........))..(((((.((.((.......))...)).))))))))..))....------ ( -19.50, z-score = -0.74, R) >droYak2.chr3R 12858072 88 - 28832112 ----GCAUAUAUGCAUCUGCAUUUGACAGAUGCAGUCAUC---------GCUCUACAAAUGCAUUUUUCACACGCACCCC---CCU--GUGAAAAGAUAAGCCGCA------ ----((.....(((((((((....).))))))))(((...---------((.........))..((((((((.(......---).)--)))))))))).....)).------ ( -23.00, z-score = -2.70, R) >droSec1.super_0 9542933 96 - 21120651 ----GCAUAUAUGCAUCUGCAUUUGACAGAUGCAGUCAUC---------GCUCUACAAAUGCAUUUUUCACACGCACCCC---CCGCUGUGAAAAGAUAAGCCAAAUGGGGG ----((((...(((((((((....).))))))))((....---------)).......)))).(((((((((.((.....---..))))))))))).....((....))... ( -25.30, z-score = -1.21, R) >droSim1.chr3R 16513455 88 - 27517382 ----GCAUAUAUGCAUCUGCAUUUGACAGAUGCAGUCAUC---------GCUCUACAAAUGCAUUUUUCACACGCACCCC---CCGCUGUGCAAAGAGGGAGGG-------- ----((.....(((((((((....).))))))))......---------((.........))...........)).((((---((.((......)).))).)))-------- ( -26.60, z-score = -1.78, R) >consensus ____GCAUAUAUGCAUCUGCAUUUGACAGAUGCAGUCAUC_________GCUCUACAAAUGCAUUUUUCACACGCACCCC___CCUCUGUGAAAAGAUAAGCCGA_______ ...........(((((((((....).)))))))).............................................................................. ( -9.58 = -9.66 + 0.08)
| Location | 10,459,560 – 10,459,650 |
|---|---|
| Length | 90 |
| Sequences | 8 |
| Columns | 112 |
| Reading direction | reverse |
| Mean pairwise identity | 66.12 |
| Shannon entropy | 0.61324 |
| G+C content | 0.47234 |
| Mean single sequence MFE | -29.30 |
| Consensus MFE | -11.28 |
| Energy contribution | -11.81 |
| Covariance contribution | 0.53 |
| Combinations/Pair | 1.31 |
| Mean z-score | -1.49 |
| Structure conservation index | 0.38 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.34 |
| SVM RNA-class probability | 0.652129 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
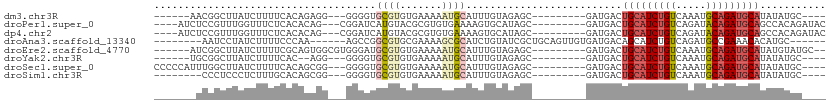
>dm3.chr3R 10459560 90 - 27905053 ------AACGGCUUAUCUUUUCACAGAGG---GGGGUGCGUGUGAAAAAUGCAUUUGUAGAGC---------GAUGACUGCAUCUGUCAAAUGCAGAUGCAUAUAUGC---- ------....(((..((((......))))---..)))(((((((.....((((((((((((((---------(.....))).)))).))))))))......)))))))---- ( -25.80, z-score = -1.22, R) >droPer1.super_0 9622958 96 - 11822988 ----AUCUCCGUUUGGUUUCUCACACAG---CGGAUCAUGUACGCGUGUGAAAAGUGCAUAGC---------GAUGACUGCAUCUGUCAGAUACAGAUGCAGCCACAGAUAC ----((((.((((..((...((((((.(---((.........))))))))).....))..)))---------).((.((((((((((.....)))))))))).)).)))).. ( -31.90, z-score = -1.97, R) >dp4.chr2 19942952 96 + 30794189 ----AUCUCCGUUUGGUUUCUCACACAG---CGGAUCAUGUACGCGUGUGAAAAGUGCAUAGC---------GAUGACUGCAUCUGUCAGAUACAGAUGCAGCCACAGAUAC ----((((.((((..((...((((((.(---((.........))))))))).....))..)))---------).((.((((((((((.....)))))))))).)).)))).. ( -31.90, z-score = -1.97, R) >droAna3.scaffold_13340 22551446 92 + 23697760 --------AAUCCUAUCUUUUCCCAA------AGCCGGCGUGCGAAAAGCGCAUCUGUAUCGCUGCAGUUGUGAUGACAGCAUCUGUCAGAUGCCGAAACACAUGC------ --------..................------...((((((((.....))))((((..(((((.......)))))(((((...)))))))))))))..........------ ( -24.20, z-score = -0.52, R) >droEre2.scaffold_4770 9992976 95 - 17746568 ------AUCGGCUUAUCUUUUCGCAGUGGCGUGGGAUGCGUGUGAAAAAUGCAUUUGUAGAGC---------GAUGACUGCAUCUGUCAAAUGCAGAUGCAUAUGUAUGC-- ------........(((((..(((....))).)))))(((((((.....((((((((((((((---------(.....))).)))).))))))))....)))))))....-- ( -29.70, z-score = -0.93, R) >droYak2.chr3R 12858072 88 + 28832112 ------UGCGGCUUAUCUUUUCAC--AGG---GGGGUGCGUGUGAAAAAUGCAUUUGUAGAGC---------GAUGACUGCAUCUGUCAAAUGCAGAUGCAUAUAUGC---- ------((((.......(((((((--(.(---......).)))))))).((((((((((((((---------(.....))).)))).))))))))..)))).......---- ( -27.30, z-score = -1.43, R) >droSec1.super_0 9542933 96 + 21120651 CCCCCAUUUGGCUUAUCUUUUCACAGCGG---GGGGUGCGUGUGAAAAAUGCAUUUGUAGAGC---------GAUGACUGCAUCUGUCAAAUGCAGAUGCAUAUAUGC---- (((((..(((.............)))..)---)))).(((((((.....((((((((((((((---------(.....))).)))).))))))))......)))))))---- ( -31.42, z-score = -1.55, R) >droSim1.chr3R 16513455 88 + 27517382 --------CCCUCCCUCUUUGCACAGCGG---GGGGUGCGUGUGAAAAAUGCAUUUGUAGAGC---------GAUGACUGCAUCUGUCAAAUGCAGAUGCAUAUAUGC---- --------.((((((.((......)).))---)))).(((((((.....((((((((((((((---------(.....))).)))).))))))))......)))))))---- ( -32.20, z-score = -2.35, R) >consensus _______UCGGCUUAUCUUUUCACAGAGG___GGGGUGCGUGUGAAAAAUGCAUUUGUAGAGC_________GAUGACUGCAUCUGUCAAAUGCAGAUGCAUAUAUGC____ .....................................((((.......))))..........................(((((((((.....)))))))))........... (-11.28 = -11.81 + 0.53)
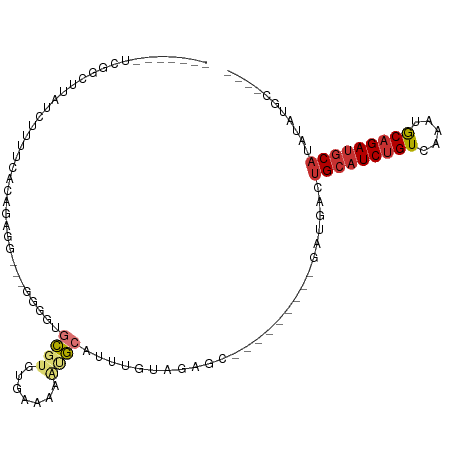
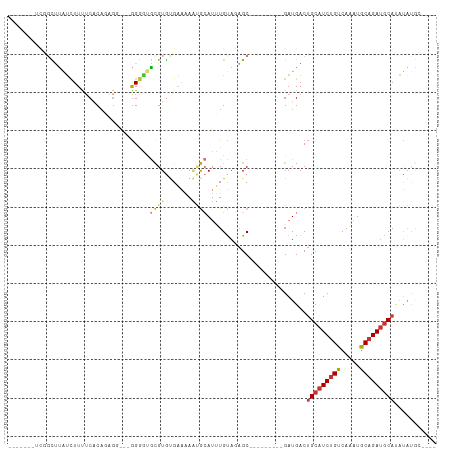
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:15:27 2011