
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 7,432,396 – 7,432,501 |
| Length | 105 |
| Max. P | 0.847892 |

| Location | 7,432,396 – 7,432,501 |
|---|---|
| Length | 105 |
| Sequences | 8 |
| Columns | 111 |
| Reading direction | forward |
| Mean pairwise identity | 70.75 |
| Shannon entropy | 0.57154 |
| G+C content | 0.41225 |
| Mean single sequence MFE | -26.33 |
| Consensus MFE | -9.97 |
| Energy contribution | -10.54 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.31 |
| Mean z-score | -1.95 |
| Structure conservation index | 0.38 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.90 |
| SVM RNA-class probability | 0.847892 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
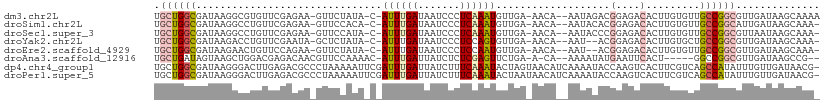
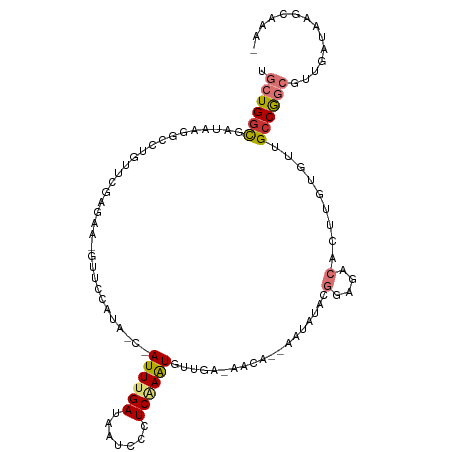
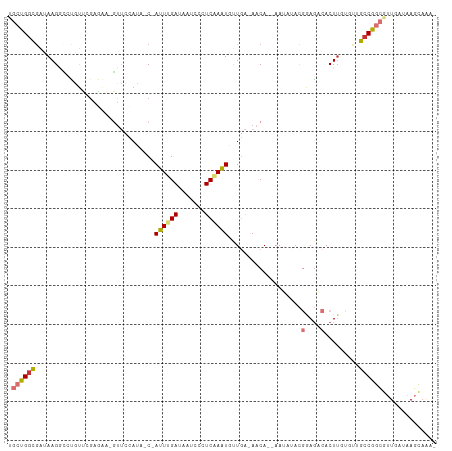
>dm3.chr2L 7432396 105 + 23011544 UGCUGGCGAUAAGGCGUGUUCGAGAA-GUUCUAUA-C-AUUUGAUAAUCCCUCAAAUGUUGA-AACA--AAUAGACGGAGACACUUGUGUUGCCGGCGUUGAUAAGCAAAA .(((((((((((((.((.((((....-.(((...(-(-((((((.......)))))))).))-)...--......)))).)).))).))))))))))(((....))).... ( -32.24, z-score = -3.01, R) >droSim1.chr2L 7224090 104 + 22036055 UGCUGGCGAUAAGGCCUGUUCGAGAA-GUUCCACA-C-AUUUGAUAAUCCCUCAAAUGUUGA-AACA--AAUACACGGAGACACUUGUGUUGCCGGCAUUGAUAAGCAAA- (((((((((((.........((((..-(((.((.(-(-((((((.......)))))))))).-))).--.......(....).)))))))))))))))............- ( -28.40, z-score = -1.86, R) >droSec1.super_3 2948930 104 + 7220098 UGCUGGCGAUAAGGCCUGUUCGAGAA-GUUCCAUA-C-AUUUGAUAAUCCCUCAAAUGUUGA-AACA--AAUACCCGGAGACACUUGUGUUGCCGGCGUUAAUAAGCAAA- .((((((((((.........((((..-(((.((.(-(-((((((.......)))))))))).-))).--.......(....).)))))))))))))).............- ( -27.90, z-score = -1.95, R) >droYak2.chr2L 16857278 102 - 22324452 UGCUGGCGAUAAGACCUGUUCGAAUA-GCUCUAUA-C-AUUUGAUAAUCCCUCCAGUGUUGA-AACA--AAU--ACGGAGACACUUGUGCUGCCGGCGUUGAUAAGCAAA- .((((((((((((......((((((.-........-.-))))))......((((.(((((..-....--)))--))))))...)))))..)))))))(((....)))...- ( -26.70, z-score = -1.70, R) >droEre2.scaffold_4929 16351337 102 + 26641161 UGCUGGCGAUAAGAACUGUUCCAGAA-GUUCUAUA-C-AUUUGAUAAUCCCUCCAAUGUUGA-AACA--AAU--ACGGAGACACUUGUGUUGCCGGCGUUGAUAAGCAAA- .((((((((((((((((........)-)))))...-.-............((((..((((..-....--)))--).)))).......))))))))))(((....)))...- ( -28.60, z-score = -2.77, R) >droAna3.scaffold_12916 15340524 99 + 16180835 UGCUGAUAGUAAGCUGGACGAGACAACGUUCCAAAAC-AUUUGAUUAUCUCUCGAGUUCUGA-A-CA--AAAAUAUGAAUUCACU-----GGCCGGCGUUGAUAAGCCG-- .(((((((((.....(((((......)))))(((...-..)))))))))....((((((...-.-..--.......))))))...-----)))((((........))))-- ( -20.02, z-score = -0.16, R) >dp4.chr4_group1 4183054 110 + 5278887 UGCUGGCGAUAAGGGACUUGAGACGCCCUAAAAAUUCGAUUUGAUUAUCUUUCAAAUACUAGUAACAUCAAAAUACCAAGUCACUUCGUCAGCCAUAUUUGUUGAUAACG- .((((((((..((.((((((((.....)).........((((((.......))))))...................)))))).)))))))))).................- ( -23.40, z-score = -1.85, R) >droPer1.super_5 5734457 110 - 6813705 UGCUGGCGAUAAGGGACUUGAGACGCCCUAAAAAUUCGAUUUGAUUAUCUUUCAAAUACUAAUAACAUCAAAAUACCAAGUCACUUCGUCAGCCAUAUUUGUUGAUAACG- .((((((((..((.((((((((.....)).........((((((.......))))))...................)))))).)))))))))).................- ( -23.40, z-score = -2.27, R) >consensus UGCUGGCGAUAAGGCCUGUUCGAGAA_GUUCCAUA_C_AUUUGAUAAUCCCUCAAAUGUUGA_AACA__AAUAUACGGAGACACUUGUGUUGCCGGCGUUGAUAAGCAAA_ .((((((...............................((((((.......))))))...................(....).........)))))).............. ( -9.97 = -10.54 + 0.56)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:23:42 2011