
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 10,239,881 – 10,239,973 |
| Length | 92 |
| Max. P | 0.755097 |

| Location | 10,239,881 – 10,239,973 |
|---|---|
| Length | 92 |
| Sequences | 10 |
| Columns | 104 |
| Reading direction | reverse |
| Mean pairwise identity | 72.59 |
| Shannon entropy | 0.54065 |
| G+C content | 0.46421 |
| Mean single sequence MFE | -24.92 |
| Consensus MFE | -9.54 |
| Energy contribution | -9.69 |
| Covariance contribution | 0.15 |
| Combinations/Pair | 1.25 |
| Mean z-score | -1.94 |
| Structure conservation index | 0.38 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.59 |
| SVM RNA-class probability | 0.755097 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 10239881 92 - 27905053 UGAGACAGUGGAGCUGAGGGC--CAUGAUUUAUGAGCUACCAACGGCAGCAUAAAUCAAAGGCAA--------CAAAAUCGGCCAUUUGUAUCGAGGGCACA-- .((.((((..(.(((((..((--(.(((((((((.(((......)))..)))))))))..)))..--------.....))))))..)))).)).........-- ( -24.60, z-score = -1.27, R) >droSim1.chr3R 16299592 92 + 27517382 UGAGACAGUGGAGCUGAGGGC--CAUGAUUUAUGAGCUACCAACGGCAGCAUAAAUCAAAGGCAA--------CAAAAUCGGCCUUUUGUAUCGAGGGCACA-- .....(..(((.((.((((((--(.(((((((((.(((......)))..)))))))))..(....--------)......))))))).)).)))..).....-- ( -28.00, z-score = -2.19, R) >droSec1.super_0 9319656 92 + 21120651 UGAGACAGUGGAGCUGAGGGC--CAUGAUUUAUGAGCUACCAACGGCAGCAUAAAUCAAAGGCAA--------CAAAAUCGGCCUUUUGUAUCGAGGGCACA-- .....(..(((.((.((((((--(.(((((((((.(((......)))..)))))))))..(....--------)......))))))).)).)))..).....-- ( -28.00, z-score = -2.19, R) >droYak2.chr3R 12631312 92 + 28832112 UGCGACAGUGGAACUGAGGGC--CAUGAUUUAUGAGCUACCAACGGCAGCAUAAAUCAAAGCCAG--------CAAAAUCGGCCUUUUGUAUCGAGGGCACA-- (((..(((.....)))..(((--..(((((((((.(((......)))..)))))))))..))).)--------))......((((((......))))))...-- ( -29.00, z-score = -2.00, R) >droEre2.scaffold_4770 9773231 91 - 17746568 UGAGACAGUGGAACUGAGGGC--CAUGAUUUAUGAGCCACCAACGGCAGCAUAAAUCAAAGGCAG--------CCAAAUCGGCC-UUUGCAUCGAGGGCACA-- .((..(((.....))).((((--(.(((((((((.(((......)))..)))))))))..)))..--------))...)).(((-((((...)))))))...-- ( -28.80, z-score = -1.76, R) >droAna3.scaffold_13340 416987 86 - 23697760 CGAGACAGUGGGUCUGAGGGC--CGAGAUUUAUGAGCUACCAACGGCAGCAUAAAUCAGAG---A--------CAAGAUCGGCCUUUU---UCGAGGCCACA-- .......((((.(((((((((--(((((((((((.(((......)))..))))))))....---.--------.....))))))))).---...))))))).-- ( -28.70, z-score = -2.70, R) >dp4.chr2 16292065 79 - 30794189 ----UCAGU--CUCUGCGGGC--AAUGAUUUAUAAGCUACCAAAGACAGCAUAAAUCAAAG---A--------CAACAUCAACCUU----AUCGAGGGCACA-- ----...((--((((.(((..--..((((((((..(((.........)))))))))))..(---(--------.....))......----.))))))).)).-- ( -12.20, z-score = -0.50, R) >droWil1.scaffold_181089 2315110 86 + 12369635 UCAAUCAGUCAUUUUG--GGGUUAAUGAUUUAUCAGCU-CUAGCAGUGGCAUAAAUCAAAGA-----------CAACAUCAACCUU----AUCGAGGACACACA .......(((....((--(((((..(((((((((.(((-.....))).).))))))))..((-----------.....))))))))----).....)))..... ( -16.40, z-score = -1.77, R) >droVir3.scaffold_13047 4540837 98 + 19223366 UCAGUCAGUCAGAGCGUUGGCUGGAUGAUUUAUAUGCUACCAAUAGCUGUAUAAAUCAAAGACAGCACACAGCAAGCAGCAACCUU----AUCGAGGACACA-- ...(((..((.(((.((((.(((..((((((((((((((....)))).))))))))))......((.....))...))))))))))----...)).)))...-- ( -25.90, z-score = -1.93, R) >droGri2.scaffold_14830 5286148 89 - 6267026 UCAGUCAGUCAGAGAGCUGGC--AUUGAUUUAUAUGCAGUCGAUGGCUGUAUAAAUCAAAAA-------CAGCAAGCGGCAACCUU----AUCGAGGACACA-- .......(((...((((((..--.((((((..((((((((.....))))))))))))))...-------))))(((.(....))))----.))...)))...-- ( -27.60, z-score = -3.14, R) >consensus UGAGACAGUGGAACUGAGGGC__CAUGAUUUAUGAGCUACCAACGGCAGCAUAAAUCAAAGGCAA________CAAAAUCGGCCUUUU__AUCGAGGGCACA__ .........................(((((((((.(((......)))..)))))))))........................((((.......))))....... ( -9.54 = -9.69 + 0.15)

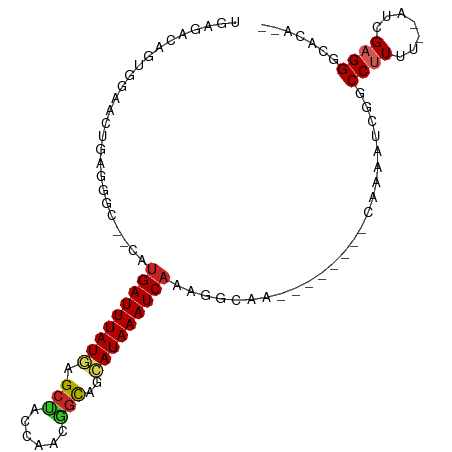

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:14:56 2011