| Sequence ID | dm3.chr3R |
|---|---|
| Location | 10,210,926 – 10,211,018 |
| Length | 92 |
| Max. P | 0.876617 |

| Location | 10,210,926 – 10,211,018 |
|---|---|
| Length | 92 |
| Sequences | 11 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 80.99 |
| Shannon entropy | 0.38386 |
| G+C content | 0.45869 |
| Mean single sequence MFE | -22.73 |
| Consensus MFE | -15.13 |
| Energy contribution | -14.98 |
| Covariance contribution | -0.15 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.73 |
| Structure conservation index | 0.67 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.03 |
| SVM RNA-class probability | 0.876617 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
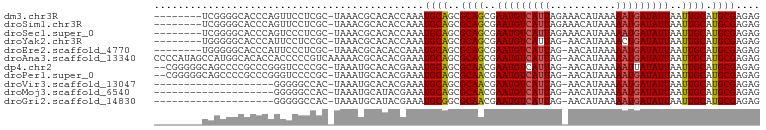
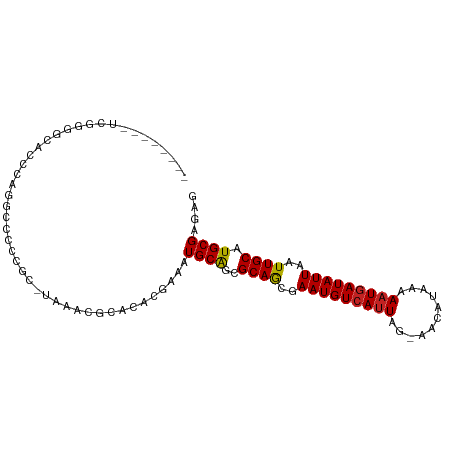
>dm3.chr3R 10210926 92 - 27905053 --------UCGGGGCACCCAGUUCCUCGC-UAAACGCACACCAAAUGCAGCGCAGCGAAUGUCAUUAGAAACAUAAAAAUGAUAUUAAUUGCAUGCGAGAG --------..(((...))).....(((((-.....(((.......)))...((((..(((((((((...........)))))))))..))))..))))).. ( -24.30, z-score = -2.05, R) >droSim1.chr3R 16272089 92 + 27517382 --------UCGGGGCACCCAGUUCCUCGC-UAAACGCACACCAAAUGCAGCGCAGCGAAUGUCAUUAGAAACAUAAAAAUGAUAUUAAUUGCAUGCGAGAG --------..(((...))).....(((((-.....(((.......)))...((((..(((((((((...........)))))))))..))))..))))).. ( -24.30, z-score = -2.05, R) >droSec1.super_0 9290819 92 + 21120651 --------UCGGGGCACCCAGUCCCUCGC-UAAACGCACACCAAAUGCAGCGCAGCGAAUGUCAUUAGAAACAUAAAAAUGAUAUUAAUUGCAUGCGAGAG --------..(((((.....)))))((((-.....(((.......)))...((((..(((((((((...........)))))))))..))))..))))... ( -26.30, z-score = -2.78, R) >droYak2.chr3R 12601665 91 + 28832112 --------UGGGGGCACCCAUUCCUCCGC-UAAACGCACACCAAAUGCAGCGCAGCGAAUGUCAUUAG-AACAUAAAACUGAUAUUAAUUGCAUGCGAGAG --------((((....))))...((((((-.....(((.......)))...((((..((((((((((.-....)))...)))))))..))))..))).))) ( -23.00, z-score = -1.44, R) >droEre2.scaffold_4770 9745280 91 - 17746568 --------UGGGGGCACCCAUUCCCUCGC-UAAACGCACACCAAAUGCAGCGCAGCGAAUGUCAUUAG-AACAUAAAAAUGAUAUUAAUUGCAUGCGAGAG --------((((....))))....(((((-.....(((.......)))...((((..(((((((((..-........)))))))))..))))..))))).. ( -27.10, z-score = -2.87, R) >droAna3.scaffold_13340 390364 100 - 23697760 CCCCAUAGCCAUGGCACACCACCCCCGUCAAAAACGCACACGAAAUGCAGCGCAGCGAAUGUCAUUAG-AACAUAAAAAUGAUAUUAAUUGCAUGCGAGAG .......((....))..............................((((..((((..(((((((((..-........)))))))))..)))).)))).... ( -18.20, z-score = -1.07, R) >dp4.chr2 16261788 97 - 30794189 --CGGGGGCAGCCCCGCCCGGGUCCCCGC-UAAAUGCACACGAAAUGCAGCGCAACGAAUGUCAUUAG-AACAUAAAAAUUAUAUUAAUUGCAUGCGAGAG --((((((..((((.....))))))))((-.....))...))...((((..((((..(((((.(((..-........))).)))))..)))).)))).... ( -24.90, z-score = -0.28, R) >droPer1.super_0 7683931 97 + 11822988 --CGGGGGCAGCCCCGCCCGGGUCCCCGC-UAAAUGCACACGAAAUGCAGCGCAACGAAUGUCAUUAG-AACAUAAAAAUGAUAUUAAUUGCAUGCGAGAG --((((((..((((.....))))))))((-.....))...))...((((..((((..(((((((((..-........)))))))))..)))).)))).... ( -31.10, z-score = -2.14, R) >droVir3.scaffold_13047 4495687 79 + 19223366 --------------------GGGGGCCAC-UAAAUGCACACGAAAUGCAGCGCAACGAAUGUCAUUAG-AACAUAAAAAUGAUAUUAAUUGCAUGCGAGAG --------------------(...((...-.....))...)....((((..((((..(((((((((..-........)))))))))..)))).)))).... ( -16.40, z-score = -1.32, R) >droMoj3.scaffold_6540 15980288 79 + 34148556 --------------------GGGGGCCAC-UAAAUGCAUACGAAAUGCAGCGCAACGAAUGUCAUUAG-AACAUAAAAAUGAUAUUAAUUGCAUGCGAGAG --------------------....((...-....(((((.....)))))..((((..(((((((((..-........)))))))))..))))..))..... ( -17.30, z-score = -1.70, R) >droGri2.scaffold_14830 5256837 79 - 6267026 --------------------GGGGGCCAC-UAAAUGCAUACGAAAUGCGGCGCAACGAAUGUCAUUAG-AACAUAAAAAUGAUAUUAAUUGCAUGCGAGAG --------------------....(((..-.....((((.....)))))))((((..(((((((((..-........)))))))))..))))......... ( -17.11, z-score = -1.38, R) >consensus ________UCGGGGCACCCAGGCCCCCGC_UAAACGCACACGAAAUGCAGCGCAGCGAAUGUCAUUAG_AACAUAAAAAUGAUAUUAAUUGCAUGCGAGAG .............................................((((..((((..(((((((((...........)))))))))..)))).)))).... (-15.13 = -14.98 + -0.15)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:14:48 2011