
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 9,920,226 – 9,920,319 |
| Length | 93 |
| Max. P | 0.541980 |

| Location | 9,920,226 – 9,920,319 |
|---|---|
| Length | 93 |
| Sequences | 9 |
| Columns | 96 |
| Reading direction | forward |
| Mean pairwise identity | 78.84 |
| Shannon entropy | 0.45820 |
| G+C content | 0.53578 |
| Mean single sequence MFE | -33.27 |
| Consensus MFE | -12.43 |
| Energy contribution | -13.31 |
| Covariance contribution | 0.88 |
| Combinations/Pair | 1.23 |
| Mean z-score | -2.14 |
| Structure conservation index | 0.37 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.10 |
| SVM RNA-class probability | 0.541980 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
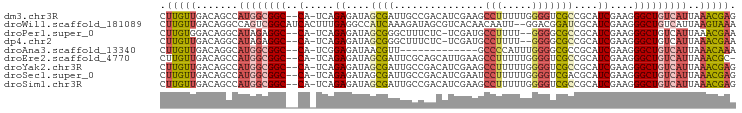
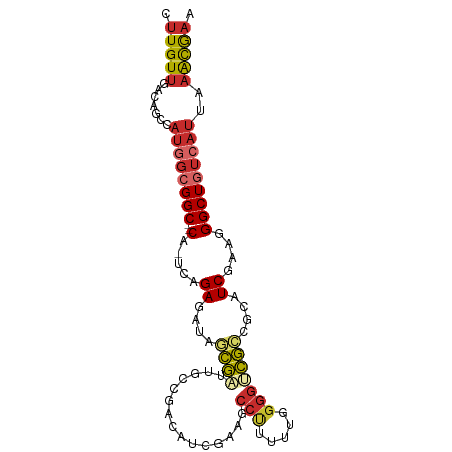
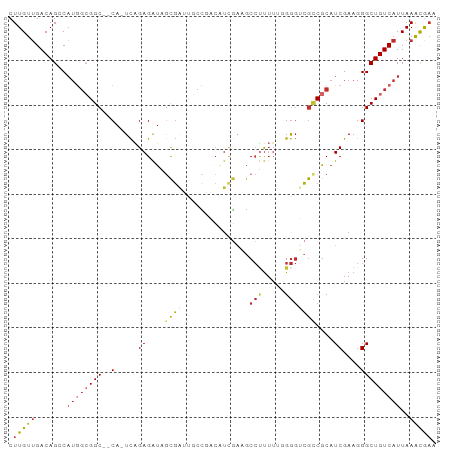
>dm3.chr3R 9920226 93 + 27905053 CUUGUUGACAGCCAUGGCGGC--CA-UCAGAGAUAGCGAUUGCCGACAUCGAAGCCUUUUUGGGGUCGCCGCAUCGAAGGGCUGUCAUUAAACGAG (((((((((((((.(((((((--(.-(((((((...((((.......)))).....)))))))))))))))........))))))))....))))) ( -35.70, z-score = -2.51, R) >droWil1.scaffold_181089 10797359 94 + 12369635 CUUGUUGACAGGCCAGUCGGCAUCACUUUGAGGCCAUCAAAGAUAGCGUCACAACAAUU--GGACGGAUCGCAUCGAAGGGCUGUCAUUAAGUAAA ((((.((((((.((..((((((((.((((((.....))))))....((((.((.....)--)))))))).))..)))..)))))))).)))).... ( -31.50, z-score = -3.18, R) >droPer1.super_0 6982932 90 + 11822988 CUUGUGGACAGGCAUAGAGGC--CA-UCAGAGAUAGCGGGCUUUCUC-UCGAUGCCUUUU--GGGGCGCCGCAUCGAAGGGCUGUCAUUAAACGAA .((((.(((((((......))--)(-(((((((.((....)).))))-).)))(((((((--(..((...))..)))))))))))).....)))). ( -30.70, z-score = -0.90, R) >dp4.chr2 17327658 90 - 30794189 CUUGUUGACAGGCAUAGAGGC--CA-UCAGAGAUAGCGGGCUUUCUC-UCGAUGCCUUUU--GGGGCGCCGCAUCGAAGGGCUGUCAUUAAACGAA .((((((((((((......))--)(-(((((((.((....)).))))-).)))(((((((--(..((...))..)))))))))))))....)))). ( -30.70, z-score = -1.08, R) >droAna3.scaffold_13340 3146441 80 - 23697760 CUUGUUGACAGGCAUGGCGGC--CA-UCGGAGAUAACGUU-------------GCCCCAUUUGGGGCGCCGCAUCGAAGGGCUGUCAUUAAACAAA .((((((((((.(...(((((--..-.((.......))..-------------(((((....)))))))))).......).))))))....)))). ( -31.50, z-score = -2.19, R) >droEre2.scaffold_4770 9464974 92 + 17746568 CUUGUUGACAGCCAUGGCGGC--CA-UCAGAGAUAGCGAUUCGCAGCAUUGAAGCCUUUUUGGGGUCGCCGCAUCGAAGGGCUGUCAUUAAACGC- .(((.((((((((.(((((((--(.-(((((((..(((...))).((......)).)))))))))))))))........)))))))).)))....- ( -34.80, z-score = -2.15, R) >droYak2.chr3R 12321202 93 - 28832112 CUUGUUGACAGCCAUGGCGGC--CA-UCAGAGAUAGCGAUUGCCGACAUCGAAGCCUUUUUGGGGUCGCCGCAUCGAAGGGCUGUCAUUAAACGAG (((((((((((((.(((((((--(.-(((((((...((((.......)))).....)))))))))))))))........))))))))....))))) ( -35.70, z-score = -2.51, R) >droSec1.super_0 9024387 93 - 21120651 CUUGUUGACAGCCAUGGCGGC--CA-UCAGAGAUAGCGAUUGCCGACAUCGAAUCCUUUUUGGGGUCGACGCAUCGAAGGGCUGUCAUUAAACGAG (((((((((((((.((.((((--.(-((.........))).)))).))((((..((.....))..))))..........))))))))....))))) ( -33.10, z-score = -2.27, R) >droSim1.chr3R 16007952 93 - 27517382 CUUGUUGACAGCCAUGGCGGC--CA-UCAGAGAUAGCGAUUGCCGACAUCGAAGCCUUUUUGGGGUCGCCGCAUCGAAGGGCUGUCAUUAAACGAG (((((((((((((.(((((((--(.-(((((((...((((.......)))).....)))))))))))))))........))))))))....))))) ( -35.70, z-score = -2.51, R) >consensus CUUGUUGACAGCCAUGGCGGC__CA_UCAGAGAUAGCGAUUGCCGACAUCGAAGCCUUUUUGGGGUCGCCGCAUCGAAGGGCUGUCAUUAAACGAA .(((.((((((.....(((((...............(((.........)))...(((.....)))..))))).((....)))))))).)))..... (-12.43 = -13.31 + 0.88)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:14:10 2011