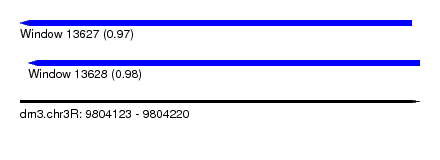
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 9,804,123 – 9,804,220 |
| Length | 97 |
| Max. P | 0.977694 |

| Location | 9,804,123 – 9,804,218 |
|---|---|
| Length | 95 |
| Sequences | 4 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 83.07 |
| Shannon entropy | 0.25107 |
| G+C content | 0.45721 |
| Mean single sequence MFE | -18.30 |
| Consensus MFE | -14.18 |
| Energy contribution | -13.99 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.47 |
| Structure conservation index | 0.77 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.83 |
| SVM RNA-class probability | 0.970119 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
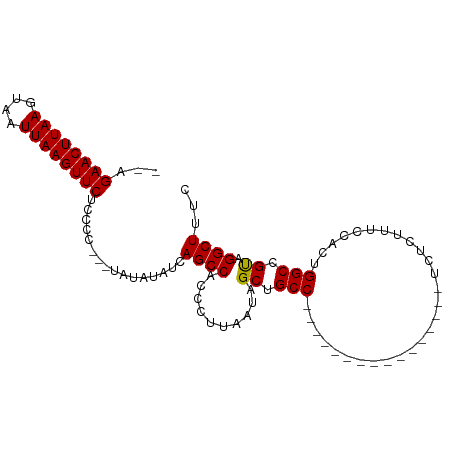
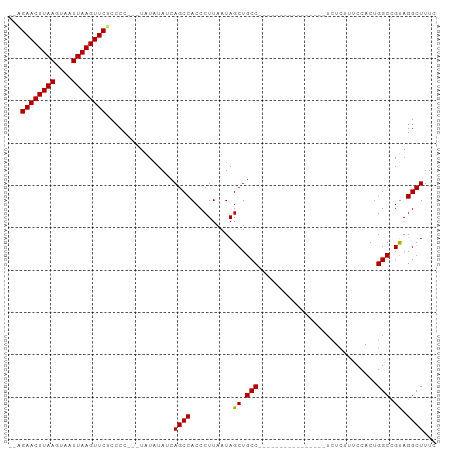
>dm3.chr3R 9804123 95 - 27905053 --GGAACUUAAGUAAUUAAGUUCUCCCC----AUAUAUCAGCCACCCUUAAUAGCUGCCUCCCAACAUCGCACCGUCUCUUUCCACUGGCCGUAGGCUUUC --(((((((((....)))))))))....----.......((((..........((..............))((.(((..........))).)).))))... ( -17.04, z-score = -1.56, R) >droEre2.scaffold_4770 9349782 94 - 17746568 ---GAACUUAAGUAAUUAAGUUCUCCCUAUAUAUGUAUCAGCCACCCUUAAUAGCUGCCUC----CAUCGCCCCGUCUCUUUCCACUGGCAGCAGGCUUUC ---((((((((....))))))))................((((....(....)((((((..----......................)))))).))))... ( -21.26, z-score = -3.19, R) >droSec1.super_0 8914191 80 + 21120651 --AGAACUUAAGUAAUUAAGUUCUCCCC---UAUAUAUCAGCCACCCUUAAUAGCUGCC----------------UCUCUUUCCACUGGCCGUAGGCUUUC --(((((((((....)))))))))..((---(((.....(((...........)))(((----------------............))).)))))..... ( -17.30, z-score = -2.53, R) >droSim1.chr3R 15894048 82 + 27517382 ACAGAACUUAAGUAAUUAAGUUCUCCCC---UAUAUAUCAGCCACCCUUAAUAGCUGCC----------------UCUCUUUCCACUGGCCGUAGGCUUUC ..(((((((((....)))))))))..((---(((.....(((...........)))(((----------------............))).)))))..... ( -17.60, z-score = -2.60, R) >consensus __AGAACUUAAGUAAUUAAGUUCUCCCC___UAUAUAUCAGCCACCCUUAAUAGCUGCC________________UCUCUUUCCACUGGCCGUAGGCUUUC ...((((((((....))))))))................((((..........((.(((............................))).)).))))... (-14.18 = -13.99 + -0.19)

| Location | 9,804,125 – 9,804,220 |
|---|---|
| Length | 95 |
| Sequences | 4 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 82.36 |
| Shannon entropy | 0.26031 |
| G+C content | 0.45578 |
| Mean single sequence MFE | -18.58 |
| Consensus MFE | -14.99 |
| Energy contribution | -14.62 |
| Covariance contribution | -0.37 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.39 |
| Structure conservation index | 0.81 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.98 |
| SVM RNA-class probability | 0.977694 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
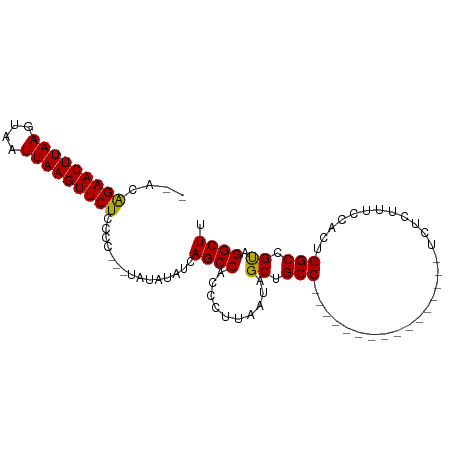
>dm3.chr3R 9804125 95 - 27905053 --ACGGAACUUAAGUAAUUAAGUUCUCCCC----AUAUAUCAGCCACCCUUAAUAGCUGCCUCCCAACAUCGCACCGUCUCUUUCCACUGGCCGUAGGCUU --..(((((((((....)))))))))....----.......((((..........((..............))((.(((..........))).)).)))). ( -17.34, z-score = -1.40, R) >droEre2.scaffold_4770 9349784 93 - 17746568 ----AGAACUUAAGUAAUUAAGUUCUCCCUAUAUAUGUAUCAGCCACCCUUAAUAGCUGCCUC----CAUCGCCCCGUCUCUUUCCACUGGCAGCAGGCUU ----(((((((((....)))))))))...............((((....(....)((((((..----......................)))))).)))). ( -21.76, z-score = -3.01, R) >droSec1.super_0 8914193 80 + 21120651 --ACAGAACUUAAGUAAUUAAGUUCUCCCC---UAUAUAUCAGCCACCCUUAAUAGCUGCC----------------UCUCUUUCCACUGGCCGUAGGCUU --..(((((((((....)))))))))..((---(((.....(((...........)))(((----------------............))).)))))... ( -17.60, z-score = -2.59, R) >droSim1.chr3R 15894050 82 + 27517382 ACACAGAACUUAAGUAAUUAAGUUCUCCCC---UAUAUAUCAGCCACCCUUAAUAGCUGCC----------------UCUCUUUCCACUGGCCGUAGGCUU ....(((((((((....)))))))))..((---(((.....(((...........)))(((----------------............))).)))))... ( -17.60, z-score = -2.54, R) >consensus __ACAGAACUUAAGUAAUUAAGUUCUCCCC___UAUAUAUCAGCCACCCUUAAUAGCUGCC________________UCUCUUUCCACUGGCCGUAGGCUU ....(((((((((....)))))))))...............((((..........((.(((............................))).)).)))). (-14.99 = -14.62 + -0.37)

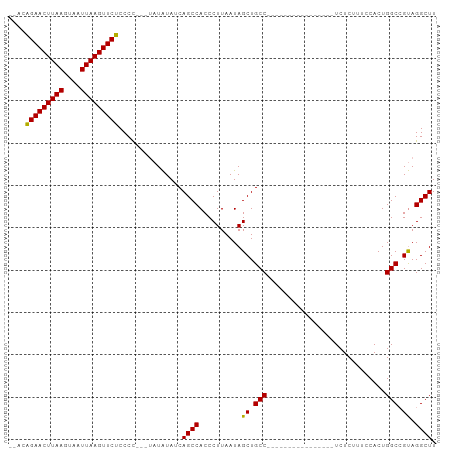
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:13:53 2011