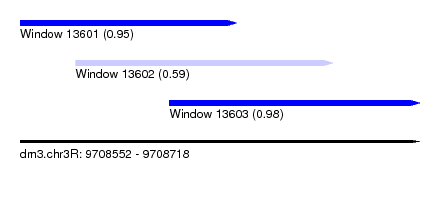
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 9,708,552 – 9,708,718 |
| Length | 166 |
| Max. P | 0.981185 |

| Location | 9,708,552 – 9,708,642 |
|---|---|
| Length | 90 |
| Sequences | 4 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 87.93 |
| Shannon entropy | 0.19453 |
| G+C content | 0.54518 |
| Mean single sequence MFE | -33.55 |
| Consensus MFE | -29.83 |
| Energy contribution | -29.70 |
| Covariance contribution | -0.13 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.97 |
| Structure conservation index | 0.89 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.51 |
| SVM RNA-class probability | 0.945829 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
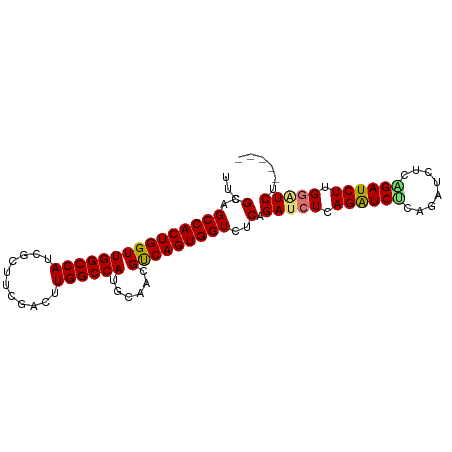
>dm3.chr3R 9708552 90 + 27905053 UUCGCAGCCACUGGUUGGCCAUCGGCCCGACUUGGCCAUGCAACGCCAGUGGUCUCAGAUCUCAGAUCGCAGAUCUCCGAUCUUGGGUCU----- ...(..(((((((((..((.((.((((......))))))))...)))))))))..)((((((.((((((........)))))).))))))----- ( -38.30, z-score = -2.77, R) >droSec1.super_0 8819935 95 - 21120651 UUCGCAGCCACUGGUUGGCCAUCGGUUCGACUUGGCCAUGCAACGUCAGUGGUCUCAGAUCUCAGGUCUCAGAUCGCAGAUCUUGGAUCCGAUCU ...(..((((((((.(((((((((...)))..)))))).......))))))))..)(((((...(((((.(((((...))))).))))).))))) ( -32.90, z-score = -1.23, R) >droYak2.chr3R 14022738 90 + 28832112 UUCGCAGCCACUGGUUGGCCAUCGCUUCGACUUGGCCAUGCAACGUCAGUGGUCUCAGAAAUCAGAUCUCUGAUCUUGGAUCUUGGAUCU----- (((...((((((((.(((((((((...)))..)))))).......))))))))....)))...((((((..((((...))))..))))))----- ( -29.80, z-score = -1.53, R) >droEre2.scaffold_4770 5823463 90 - 17746568 UUCGCAGCCACUGGUUGGCCAUCGCUUCGUCUUGGCCAUGCAACGUCAGUGGUCUCAGAUCUCAGGUCUCAGAUCUGUGAUCUUGGAUCU----- ...(..((((((((.((((((..((...))..)))))).......))))))))..)((((((.(((((.((....)).))))).))))))----- ( -33.20, z-score = -2.34, R) >consensus UUCGCAGCCACUGGUUGGCCAUCGCUUCGACUUGGCCAUGCAACGUCAGUGGUCUCAGAUCUCAGAUCUCAGAUCUCAGAUCUUGGAUCU_____ ...(..(((((((((((((((...........))))))......)))))))))..).(((((.((((((........)))))).)))))...... (-29.83 = -29.70 + -0.13)

| Location | 9,708,575 – 9,708,682 |
|---|---|
| Length | 107 |
| Sequences | 4 |
| Columns | 112 |
| Reading direction | forward |
| Mean pairwise identity | 87.67 |
| Shannon entropy | 0.19735 |
| G+C content | 0.54026 |
| Mean single sequence MFE | -38.90 |
| Consensus MFE | -26.44 |
| Energy contribution | -27.50 |
| Covariance contribution | 1.06 |
| Combinations/Pair | 1.19 |
| Mean z-score | -2.13 |
| Structure conservation index | 0.68 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.19 |
| SVM RNA-class probability | 0.587917 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
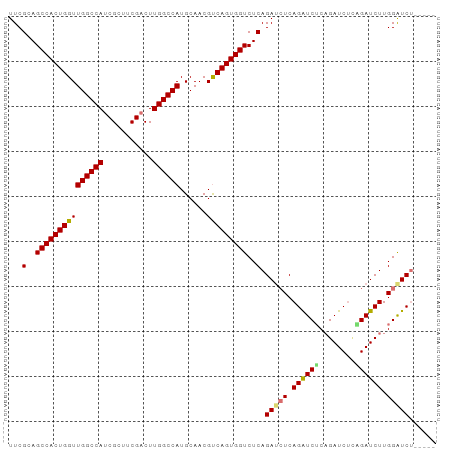
>dm3.chr3R 9708575 107 + 27905053 GGCCCGACUUGGCCAUGCAACGCCAGUGGUCUCAGAUCUCAGAUCGCAGAUCUCCGAUCUUGGGUC-----UCCCAUUCCGGGUCCAUCUCAUCUGGUCCAAAUUGGAGCGG ((((((...((((........))))((((....((((((.((((((........)))))).)))))-----).))))..))))))........(((.(((.....))).))) ( -37.70, z-score = -1.06, R) >droSec1.super_0 8819958 112 - 21120651 GGUUCGACUUGGCCAUGCAACGUCAGUGGUCUCAGAUCUCAGGUCUCAGAUCGCAGAUCUUGGAUCCGAUCUCCCAUCCCGGGUCCAUCUCAUCCUGUCCAAAUUGGAGCGG (.(((((.((((.((..........(((((((.((((.....)))).))))))).((...((((((((...........))))))))..))....)).)))).))))).).. ( -36.20, z-score = -0.69, R) >droYak2.chr3R 14022761 107 + 28832112 GCUUCGACUUGGCCAUGCAACGUCAGUGGUCUCAGAAAUCAGAUCUCUGAUCUUGGAUCUUGGAUC-----UCCCAUCCCAGAUCCAUCUCAUCUGGUCCAAAUUGGAGCGG (((((((.(((((((......(((((.(((((.(....).))))).)))))..(((((((.((((.-----....)))).))))))).......))).)))).))))))).. ( -43.20, z-score = -4.08, R) >droEre2.scaffold_4770 5823486 107 - 17746568 GCUUCGUCUUGGCCAUGCAACGUCAGUGGUCUCAGAUCUCAGGUCUCAGAUCUGUGAUCUUGGAUC-----UCCCAUCCCAGAUCCAUCUCAUCUGGUCCAAAUUGGAGCGG ((((((..(((((((............((((.(((((((........))))))).)))).((((((-----(........))))))).......))).))))..)))))).. ( -38.50, z-score = -2.70, R) >consensus GCUUCGACUUGGCCAUGCAACGUCAGUGGUCUCAGAUCUCAGAUCUCAGAUCUCAGAUCUUGGAUC_____UCCCAUCCCAGAUCCAUCUCAUCUGGUCCAAAUUGGAGCGG (((((((.(((((((............(((((.((((.....)))).)))))...(((((.((((..........)))).))))).........))).)))).))))))).. (-26.44 = -27.50 + 1.06)

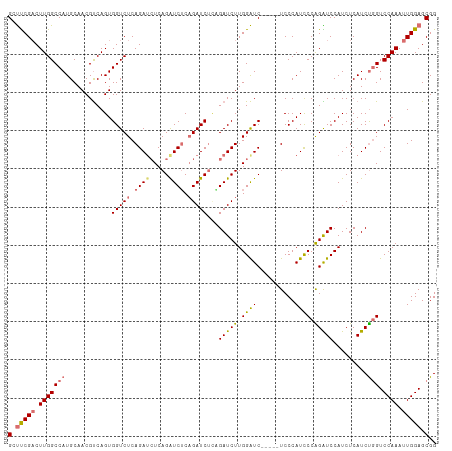
| Location | 9,708,614 – 9,708,718 |
|---|---|
| Length | 104 |
| Sequences | 5 |
| Columns | 109 |
| Reading direction | forward |
| Mean pairwise identity | 86.60 |
| Shannon entropy | 0.23371 |
| G+C content | 0.47146 |
| Mean single sequence MFE | -36.36 |
| Consensus MFE | -26.66 |
| Energy contribution | -26.66 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.19 |
| Mean z-score | -2.85 |
| Structure conservation index | 0.73 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.07 |
| SVM RNA-class probability | 0.981185 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
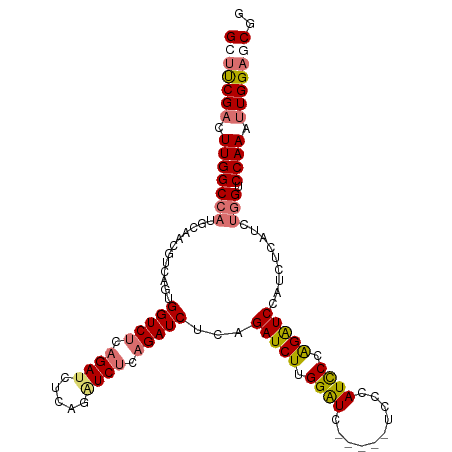
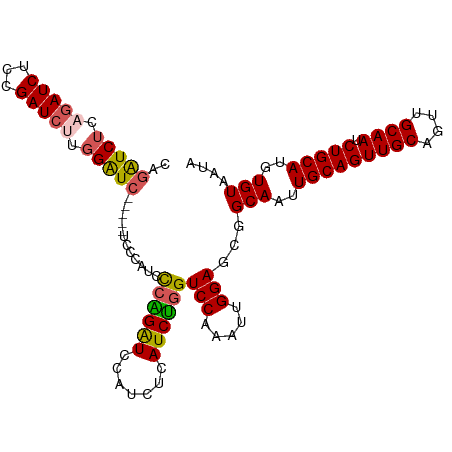
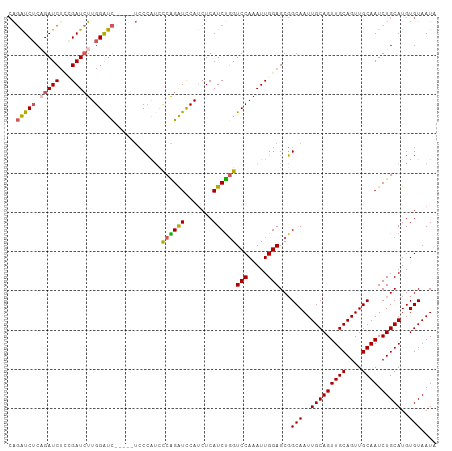
>dm3.chr3R 9708614 104 + 27905053 CAGAUCGCAGAUCUCCGAUCUUGGGUC-----UCCCAUUCCGGGUCCAUCUCAUCUGGUCCAAAUUGGAGCGGCAAUUGCAGUUGCAGUUGCAAUCUGCAUGUGUAAUA ((.((.(((((((((((((.((((..(-----.(((.....)))............)..)))))))))))..((((((((....)))))))).)))))))).))..... ( -38.42, z-score = -3.24, R) >droSec1.super_0 8819997 109 - 21120651 CAGGUCUCAGAUCGCAGAUCUUGGAUCCGAUCUCCCAUCCCGGGUCCAUCUCAUCCUGUCCAAAUUGGAGCGGCAAUUGCAGUUGCAGUUGCAAUCUGCAUGUGUAAUA ((.((..((((((((.((...((((((((...........))))))))..))......(((.....))))))((((((((....)))))))).))))).)).))..... ( -35.80, z-score = -2.07, R) >droYak2.chr3R 14022800 104 + 28832112 CAGAUCUCUGAUCUUGGAUCUUGGAUC-----UCCCAUCCCAGAUCCAUCUCAUCUGGUCCAAAUUGGAGCGGCAAUUGCAGUUGCAGUUGCAAUCUGCAUGUGUAAUA ((((((((..((.(((((((.((((((-----(........)))))))........)))))))))..)))..((((((((....)))))))).)))))........... ( -36.70, z-score = -2.92, R) >droEre2.scaffold_4770 5823525 104 - 17746568 CAGGUCUCAGAUCUGUGAUCUUGGAUC-----UCCCAUCCCAGAUCCAUCUCAUCUGGUCCAAAUUGGAGCGGCAAUUGCAGUUGCAGUUGCAAUCUGCAUGUGUAAUA ..((.(.(((((.((.(((((.((((.-----....)))).)))))))....)))))).))....((.((..((((((((....))))))))...)).))......... ( -32.60, z-score = -1.69, R) >droAna3.scaffold_13340 1063755 100 - 23697760 -GCAUCUCUGAUCUCCGAU-UUGGAUU-------CAGAUUCAGAUUCAACUCAUCUGGUCCAAAUUGGAGCUGCAAUUGCAGUUGCAGUUGCAAUCUGCAUGUGUAAUA -(((........(((((((-((((((.-------(((((..((......)).)))))))))))))))))).(((((((((....)))))))))...))).......... ( -38.30, z-score = -4.33, R) >consensus CAGAUCUCAGAUCUCCGAUCUUGGAUC_____UCCCAUCCCAGAUCCAUCUCAUCUGGUCCAAAUUGGAGCGGCAAUUGCAGUUGCAGUUGCAAUCUGCAUGUGUAAUA ..(((((.(((((...))))).)))))............((((((.......))))))(((.....)))...(((..(((((((((....)))).)))))..))).... (-26.66 = -26.66 + 0.00)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:13:32 2011