
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 9,615,291 – 9,615,395 |
| Length | 104 |
| Max. P | 0.654252 |

| Location | 9,615,291 – 9,615,395 |
|---|---|
| Length | 104 |
| Sequences | 8 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 76.08 |
| Shannon entropy | 0.43514 |
| G+C content | 0.40217 |
| Mean single sequence MFE | -30.26 |
| Consensus MFE | -12.49 |
| Energy contribution | -13.40 |
| Covariance contribution | 0.91 |
| Combinations/Pair | 1.20 |
| Mean z-score | -2.17 |
| Structure conservation index | 0.41 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.35 |
| SVM RNA-class probability | 0.654252 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
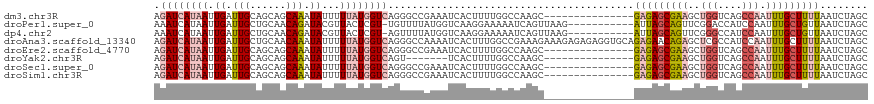
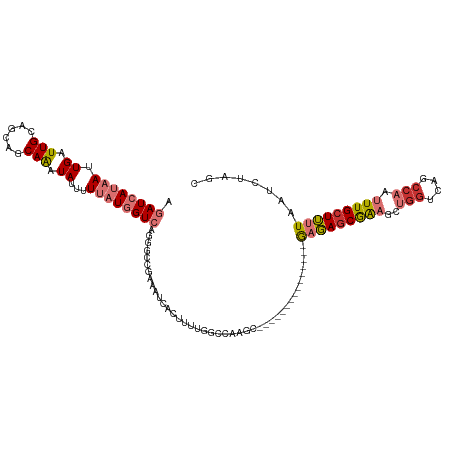
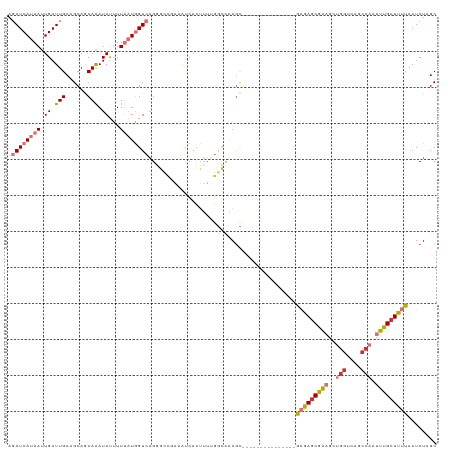
>dm3.chr3R 9615291 104 + 27905053 AGAUCAUAAUUGAUUGCAGCAGCAAAUAUUUUUAUGGUCAGGGCCGAAAUCACUUUUGGCCAAGC---------------GAGAGCGAAGCUGGUCAGCCAAUUUGCUUUUAAUCUAGC .((((((((.((.((((....)))).))...))))))))..((((((((....))))))))....---------------(((((((((..(((....))).)))))))))........ ( -35.10, z-score = -3.15, R) >droPer1.super_0 7020722 107 - 11822988 AAAUCAUAAUUGAUUGCUGCAACAGAUACGUUACUCGU-UGUUUUAUGGUCAAGGAAAAAUCAGUUAAG-----------AUUAGCAGUUCGGACCAUCCAAUUUGCUGUUAAUCUAGC ......(((((((((.(((((((((........)).))-)))...........))...)))))))))((-----------(((((((((..((.....)).....)))))))))))... ( -24.00, z-score = -1.58, R) >dp4.chr2 15610901 107 + 30794189 AAAUCAUAAUUGAUUGCUGCAACAGAUACGUUACUCGU-AGUUUUAUGGUCAAGGAAAAAUCAGUUAAG-----------AUUAGCAGUUCGGGCCAUCCAAUUUGCUGUUAAUCUAGC ......(((((((((.((....((..((((.....)))-)......)).....))...)))))))))((-----------(((((((((...((....)).....)))))))))))... ( -25.30, z-score = -1.89, R) >droAna3.scaffold_13340 17940111 119 + 23697760 AGAUCAUAAUUGAUUGCUGCAACAAAUAUUUUUAUGGUCAGGGCCAAAAUCACUUUGGCCGAAAGAAAGAGAGAGGUGCAGAGAACAGAGCUCGCCAUCCAAUUUGCUUUUAAUCUAGC .((((((((.((.(((......))).))...))))))))..(((((((.....)))))))((..((((((((..((.((.(((.......)))))...))..))).)))))..)).... ( -27.60, z-score = -0.42, R) >droEre2.scaffold_4770 5729416 104 - 17746568 AGAUCAUAAUUGAUUGCAGCAGCAAAUAUUUUUAUGGUCAGGGCCGAAAUCACUUUUGGCCAAGC---------------GAGAGCGAAGCUGGUCAGCCAAUUUGCUUUUAAUCUAGC .((((((((.((.((((....)))).))...))))))))..((((((((....))))))))....---------------(((((((((..(((....))).)))))))))........ ( -35.10, z-score = -3.15, R) >droYak2.chr3R 13924235 97 + 28832112 AGAUCAUAAUUGAUUGCAGCAGCAAAUAUUUUUAUGGUCAGU-------UCACUUUUGGCCAAGC---------------GAGAGCGAAGCUGGUCAGCCAAUUUGCUUUUAAUCUAGC .(((((....)))))...((.(((((....))).(((((((.-------......))))))).))---------------(((((((((..(((....))).)))))))))......)) ( -24.80, z-score = -0.90, R) >droSec1.super_0 8728533 104 - 21120651 AGAUCAUAAUUGAUUGCAGCAGCAAAUAUUUUUAUGGUCAGGGCCGAAAUCACUUUUGGCCAAGC---------------GAGAGCGAAGCUGGUCAGCCAAUUUGCUUUUAAUCUAGC .((((((((.((.((((....)))).))...))))))))..((((((((....))))))))....---------------(((((((((..(((....))).)))))))))........ ( -35.10, z-score = -3.15, R) >droSim1.chr3R 15703716 104 - 27517382 AGAUCAUAAUUGAUUGCAGCAGCAAAUAUUUUUAUGGUCAGGGCCGAAAUCACUUUUGGCCAAGC---------------GAGAGCGAAGCUGGUCAGCCAAUUUGCUUUUAAUCUAGC .((((((((.((.((((....)))).))...))))))))..((((((((....))))))))....---------------(((((((((..(((....))).)))))))))........ ( -35.10, z-score = -3.15, R) >consensus AGAUCAUAAUUGAUUGCAGCAGCAAAUAUUUUUAUGGUCAGGGCCGAAAUCACUUUUGGCCAAGC_______________GAGAGCGAAGCUGGUCAGCCAAUUUGCUUUUAAUCUAGC .((((((((.((.(((......))).))...)))))))).........................................(((((((((..(((....))).)))))))))........ (-12.49 = -13.40 + 0.91)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:13:10 2011