
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 9,502,593 – 9,502,740 |
| Length | 147 |
| Max. P | 0.882071 |

| Location | 9,502,593 – 9,502,740 |
|---|---|
| Length | 147 |
| Sequences | 7 |
| Columns | 162 |
| Reading direction | reverse |
| Mean pairwise identity | 66.58 |
| Shannon entropy | 0.64771 |
| G+C content | 0.45567 |
| Mean single sequence MFE | -36.87 |
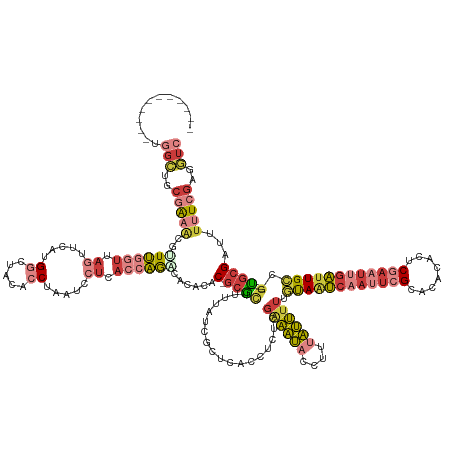
| Consensus MFE | -14.65 |
| Energy contribution | -16.72 |
| Covariance contribution | 2.07 |
| Combinations/Pair | 1.45 |
| Mean z-score | -1.58 |
| Structure conservation index | 0.40 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.05 |
| SVM RNA-class probability | 0.882071 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
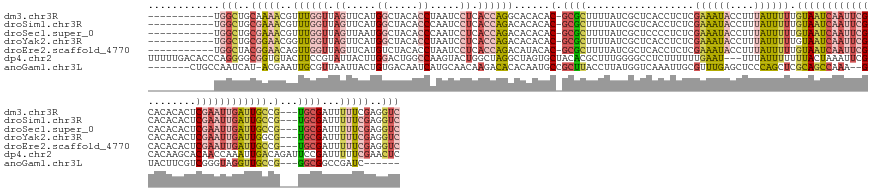
>dm3.chr3R 9502593 147 - 27905053 -----------UGGCUGCAAAACGUUUGGUUAGUUCAUGGCUACACCUAAUCCUCACCAGGCACACAC-GCGCUUUUAUCGCUCACCUCUCGAAAUACCUUUAUUUUUGUAAUCAAUUCGCACACACUCGAAUUGAUUGCCG---UGCGAUUUUUCGAGGUC -----------.((((.(.....(((((((.((.....((.....)).....)).))))))).....(-((((..................((((((....)))))).((((((((((((........)))))))))))).)---)))).......).)))) ( -34.60, z-score = -1.65, R) >droSim1.chr3R 15584386 147 + 27517382 -----------UGGCUGCGAAACGUUUGGUUAGUUCAUGGCUACACCCAAUCCUCACCAGACACACAC-GCGCUUUUAUCGCUCACCUCUCGAAAUACCUUUAUUUUUGUAAUCAAUUCGCACACACUCGAAUUGAUUGCCG---UGCGAUUUUUCGAGGUC -----------.((((.(((((.(((((((.((....(((......)))...)).))))))).....(-((((..................((((((....)))))).((((((((((((........)))))))))))).)---))))...))))).)))) ( -40.20, z-score = -3.33, R) >droSec1.super_0 8616172 147 + 21120651 -----------UGGCUGCGAAACGUUUGGUUAGUUAAUGGCUACACCCAAUCCUCACCAGACACACAC-GCGCUUUUAUCGCUCCCCUCUCGAAAUACCUUUAUUUUUGUAAUCAAUUCGCACACACUCGAAUUGAUUGCCG---UGCGAUUUUUCGAGGUC -----------.((((.(((((.(((((((.((....(((......)))...)).))))))).....(-((((..................((((((....)))))).((((((((((((........)))))))))))).)---))))...))))).)))) ( -40.20, z-score = -3.41, R) >droYak2.chr3R 13811002 147 - 28832112 -----------UGGCUGCGGAACGGUUGGUUAGUUCAUGGCUACACCUAAUCCUCACCAGACACACAC-GCGCUUUUAUCGCUCACCUCUCGAAAUACCUUUAUUUUUGUAAUCAAUUCGCACACACUCGAAUUGAUUGGCG---UGCGAUUUUUCGAGGUC -----------(((.((.(((..(((((((((.....)))))).)))...))).)))))(((...(((-((((.......))...........................(((((((((((........))))))))))))))---))(((....)))..))) ( -36.70, z-score = -1.37, R) >droEre2.scaffold_4770 5613756 147 + 17746568 -----------UGGCUACGGAACAGUUGGUUAGUUCAUGUCUACACCUAAUCCUCACCAGACAUACAC-GCGCUUUUAUCGCUCACCUCUCGAAAUACCUUUAUUUUUGUAAUCAAUUCGCACACACUCGAAUUGAUUGCCG---UGCGAUUUUUCGAGGUC -----------.((((.(((((..............((((((................))))))...(-((((..................((((((....)))))).((((((((((((........)))))))))))).)---))))...))))).)))) ( -31.99, z-score = -1.30, R) >dp4.chr2 16943934 159 - 30794189 UUUUUGACACCCAGGGGCGGUGUACUUCCGUAUUACUUGGACUGGCCAAGUACUGGCUAGGCUAGUGCUACACGCUUUGGGGCCUCUUUUUUGAAU---UUUAUUUUUUUACUAAAUUCGCACAAGCACAACCAAAUUGACAGAUUCCGAUUUUUCGAACUC ..((((....(((((.((((.......))))....))))).(((((((.....)))))))(((.((((.....(((....))).........((((---((............)))))))))).))).............))))...(((....)))..... ( -39.10, z-score = -1.15, R) >anoGam1.chr3L 22843219 143 + 41284009 -------CUGCCAAUCAU-ACGAAUUGCGUUAAUUACUGUGACAAUCAUGCAACAAGACACACAAUGCCGCUUACCUUAUGGUCAAAUUGCGUUUGAGCUCCCAGCUCGCAGCCAAA--GUACUUCGUCGGGUAGGUUGCCG---GGCGGCCGAUC------ -------(((((......-.....((((((.....)).(((.....))))))).............((.((((((((.((((.....(((.((((((((.....))))).)))))).--.....)))).)))))))).))..---)))))......------ ( -35.30, z-score = 1.12, R) >consensus ___________UGGCUGCGAAACGUUUGGUUAGUUCAUGGCUACACCUAAUCCUCACCAGACACACAC_GCGCUUUUAUCGCUCACCUCUCGAAAUACCUUUAUUUUUGUAAUCAAUUCGCACACACUCGAAUUGAUUGCCG___UGCGAUUUUUCGAGGUC ........................((((((.((.....((.....)).....)).))))))...........................(((((((.............((((((((((((........))))))))))))............)))))))... (-14.65 = -16.72 + 2.07)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:12:48 2011