
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 9,496,123 – 9,496,214 |
| Length | 91 |
| Max. P | 0.500000 |

| Location | 9,496,123 – 9,496,214 |
|---|---|
| Length | 91 |
| Sequences | 3 |
| Columns | 93 |
| Reading direction | reverse |
| Mean pairwise identity | 80.61 |
| Shannon entropy | 0.25402 |
| G+C content | 0.50117 |
| Mean single sequence MFE | -14.90 |
| Consensus MFE | -10.57 |
| Energy contribution | -10.57 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.67 |
| Structure conservation index | 0.71 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.02 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
>dm3.chr3R 9496123 91 - 27905053 CCAAGCGAGGCGGCCCAUUUCCCAACCCUCCACCGCCAAACCCAUUCGUUUUGGUAGUCUAAACGCUUUACUCUACCGCUUUCUUACCUUA-- ..((((..(((((...................)))))...............(((((..(((.....)))..)))))))))..........-- ( -18.61, z-score = -2.48, R) >droSim1.chr3R 15578026 77 + 27517382 CCAAGCGA---GGCCCAUUUCCCGCCC------------AUCCACUCGUUUUGGUAGUCUAAACGCUUUACUCUACCGCUUU-UUACCUUGUU ..(((((.---(((.........))))------------.............(((((..(((.....)))..))))))))).-.......... ( -13.10, z-score = -1.25, R) >droSec1.super_0 8609840 77 + 21120651 CCAAGCGA---GGCCCAUUUCCCGCCC------------ACUCAUUCGUUUUGGUAGUCUAACCGCUUUACUCUACCGCUUU-UUACCUUGUA ..(((((.---(((.........))))------------.............(((((..(((.....)))..))))))))).-.......... ( -13.00, z-score = -1.26, R) >consensus CCAAGCGA___GGCCCAUUUCCCGCCC____________ACCCAUUCGUUUUGGUAGUCUAAACGCUUUACUCUACCGCUUU_UUACCUUGU_ ..((((.....((........)).............................(((((..(((.....)))..)))))))))............ (-10.57 = -10.57 + 0.00)

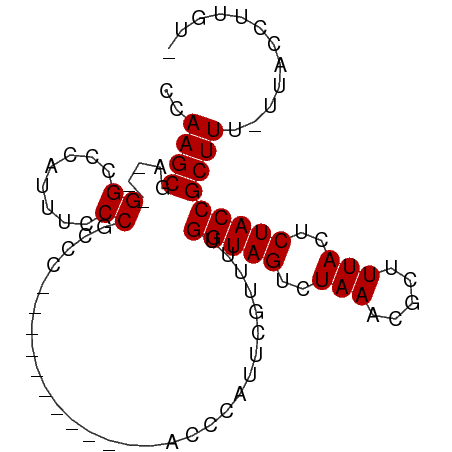

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:12:44 2011