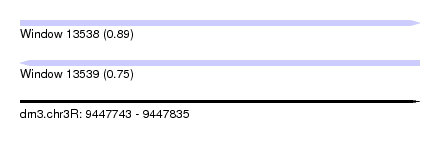

| Sequence ID | dm3.chr3R |
|---|---|
| Location | 9,447,743 – 9,447,835 |
| Length | 92 |
| Max. P | 0.893550 |

| Location | 9,447,743 – 9,447,835 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 92 |
| Reading direction | forward |
| Mean pairwise identity | 91.69 |
| Shannon entropy | 0.14021 |
| G+C content | 0.46000 |
| Mean single sequence MFE | -26.98 |
| Consensus MFE | -25.46 |
| Energy contribution | -24.98 |
| Covariance contribution | -0.48 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.59 |
| Structure conservation index | 0.94 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.11 |
| SVM RNA-class probability | 0.893550 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
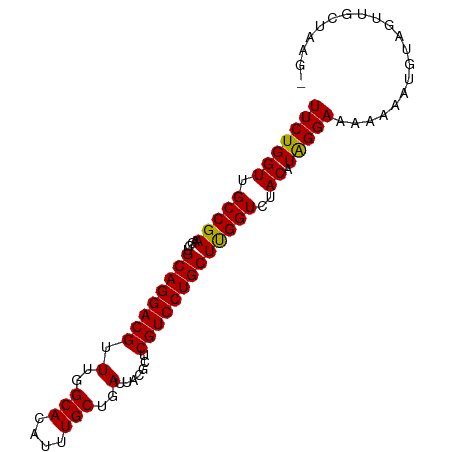
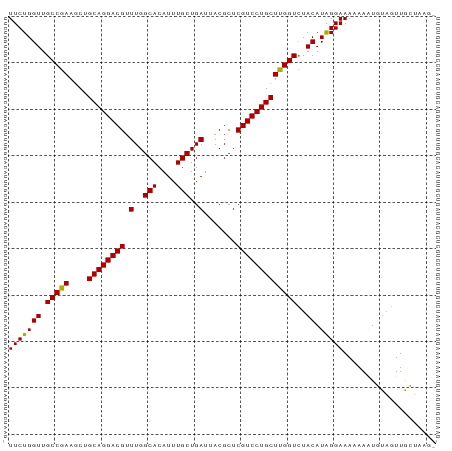
>dm3.chr3R 9447743 92 + 27905053 UUCUGGUUGCCGAAUCUGCAGGACGUUUGGCACAUUUGCUGAUUACGCUCGUCCUGCUUGGUCUACAUAGGAAAAAAAUGUAGUUACUAAGC ...((((.(((((....((((((((.(..(((....)))..).......)))))))))))))((((((.........))))))..))))... ( -25.60, z-score = -1.48, R) >droSim1.chr3R 15528582 92 - 27517382 UUCUGGUUGCCGAAGCUGCAGGACGUUUGGCACAUUUGCUGAUUACGCUCGUCCUGCUUGGUCUACAUGGGAAAAAAAUGUAGUUGCUAAGA ....((((.....))))((((((((.(..(((....)))..).......))))))))(((((((((((.........))))))..))))).. ( -26.90, z-score = -0.98, R) >droSec1.super_0 8561240 92 - 21120651 UUCUGGUUGCCGAAGCUGCAGGACGUUUGGCACAUUUGCUGAUUACGCUCGUCCUGCUUGGUCUACAUGGGAAAAAAAUGUAGUUGCUAAGA ....((((.....))))((((((((.(..(((....)))..).......))))))))(((((((((((.........))))))..))))).. ( -26.90, z-score = -0.98, R) >droYak2.chr3R 13756919 87 + 28832112 UUCUGGUUGCCGAAGCUGCAGGACGUUUGGCACAUUUGCUGAUUACGCUCGUCCUGCUCGGUCUACAUAGGAAACAA-UUUAGUUGAU---- (((((((.(((((....((((((((.(..(((....)))..).......)))))))))))))..)).))))).....-..........---- ( -27.30, z-score = -1.92, R) >droEre2.scaffold_4770 5558980 87 - 17746568 UUCUGGUUGCCGAAGCUGCAGGACGUUUGGCACAUUUGCUGAUUACGCUCGUCCUGCUCGGUAUACAUAGGAAAAAA-UUUUGUUGAU---- (((((((((((((....((((((((.(..(((....)))..).......)))))))))))))).)).))))).....-..........---- ( -28.20, z-score = -2.57, R) >consensus UUCUGGUUGCCGAAGCUGCAGGACGUUUGGCACAUUUGCUGAUUACGCUCGUCCUGCUUGGUCUACAUAGGAAAAAAAUGUAGUUGCUAAG_ (((((((.(((((....((((((((.(..(((....)))..).......)))))))))))))..)).))))).................... (-25.46 = -24.98 + -0.48)

| Location | 9,447,743 – 9,447,835 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 92 |
| Reading direction | reverse |
| Mean pairwise identity | 91.69 |
| Shannon entropy | 0.14021 |
| G+C content | 0.46000 |
| Mean single sequence MFE | -23.12 |
| Consensus MFE | -18.76 |
| Energy contribution | -18.96 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.98 |
| Structure conservation index | 0.81 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.57 |
| SVM RNA-class probability | 0.745245 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
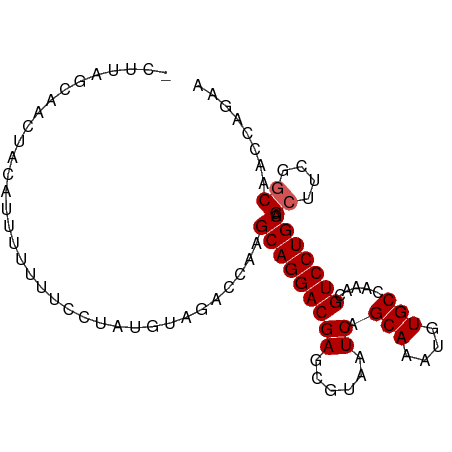
>dm3.chr3R 9447743 92 - 27905053 GCUUAGUAACUACAUUUUUUUCCUAUGUAGACCAAGCAGGACGAGCGUAAUCAGCAAAUGUGCCAAACGUCCUGCAGAUUCGGCAACCAGAA (((.(((..((((((.........)))))).....(((((((((......)).(((....))).....)))))))..))).)))........ ( -24.40, z-score = -2.35, R) >droSim1.chr3R 15528582 92 + 27517382 UCUUAGCAACUACAUUUUUUUCCCAUGUAGACCAAGCAGGACGAGCGUAAUCAGCAAAUGUGCCAAACGUCCUGCAGCUUCGGCAACCAGAA .........((((((.........)))))).....(((((((((......)).(((....))).....))))))).((....))........ ( -24.70, z-score = -2.46, R) >droSec1.super_0 8561240 92 + 21120651 UCUUAGCAACUACAUUUUUUUCCCAUGUAGACCAAGCAGGACGAGCGUAAUCAGCAAAUGUGCCAAACGUCCUGCAGCUUCGGCAACCAGAA .........((((((.........)))))).....(((((((((......)).(((....))).....))))))).((....))........ ( -24.70, z-score = -2.46, R) >droYak2.chr3R 13756919 87 - 28832112 ----AUCAACUAAA-UUGUUUCCUAUGUAGACCGAGCAGGACGAGCGUAAUCAGCAAAUGUGCCAAACGUCCUGCAGCUUCGGCAACCAGAA ----.((.......-....................(((((((((......)).(((....))).....))))))).((....)).....)). ( -20.90, z-score = -0.95, R) >droEre2.scaffold_4770 5558980 87 + 17746568 ----AUCAACAAAA-UUUUUUCCUAUGUAUACCGAGCAGGACGAGCGUAAUCAGCAAAUGUGCCAAACGUCCUGCAGCUUCGGCAACCAGAA ----.((.......-....................(((((((((......)).(((....))).....))))))).((....)).....)). ( -20.90, z-score = -1.69, R) >consensus _CUUAGCAACUACAUUUUUUUCCUAUGUAGACCAAGCAGGACGAGCGUAAUCAGCAAAUGUGCCAAACGUCCUGCAGCUUCGGCAACCAGAA ...................................(((((((((......)).(((....))).....))))))).((....))........ (-18.76 = -18.96 + 0.20)
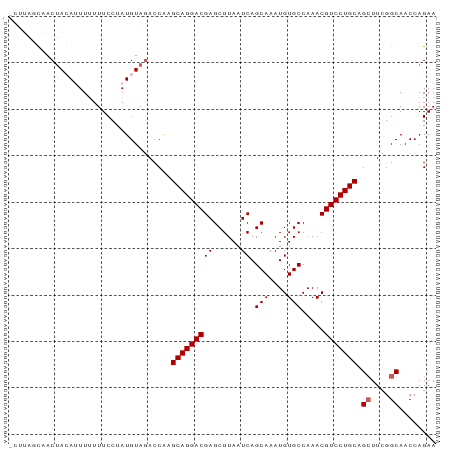
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:12:39 2011