
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 9,438,045 – 9,438,120 |
| Length | 75 |
| Max. P | 0.754738 |

| Location | 9,438,045 – 9,438,120 |
|---|---|
| Length | 75 |
| Sequences | 9 |
| Columns | 81 |
| Reading direction | reverse |
| Mean pairwise identity | 74.17 |
| Shannon entropy | 0.52437 |
| G+C content | 0.56126 |
| Mean single sequence MFE | -22.46 |
| Consensus MFE | -12.74 |
| Energy contribution | -12.74 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.50 |
| Mean z-score | -1.14 |
| Structure conservation index | 0.57 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.59 |
| SVM RNA-class probability | 0.754738 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 9438045 75 - 27905053 GUAGCAGAAGUUUCCUCGAAAAUGCGCACAUUUGUUUGCCUUGCCCUGGCGACC---AUUCU---CCUGGUCGCCGGCUCA ...(((....((((...)))).)))(((........)))...(((..(((((((---(....---..)))))))))))... ( -23.70, z-score = -1.83, R) >droSim1.chr3R 15517718 75 + 27517382 GUAGCAGAAGUUUCCUCGAAAAUGCGCACAUUUGUUUGCCUUGCCCUGGCGACC---AUUCU---CCUGGUCGCCGGCUCA ...(((....((((...)))).)))(((........)))...(((..(((((((---(....---..)))))))))))... ( -23.70, z-score = -1.83, R) >droSec1.super_0 8551512 75 + 21120651 GUAGCAGAAGUUUCCUCGAAAAUGCGCACAUUUGUUUGCCUUGCCCUGGCGACC---AUUCU---CCUAGUCGCCGGCUCA ...(((....((((...)))).)))(((........)))...(((..((((((.---.....---....)))))))))... ( -17.90, z-score = -0.39, R) >droYak2.chr3R 13747070 75 - 28832112 GUAGCAGGAGUUUUCUCGAAUAUGCGCACAUUUGUUUGCCUCGCCCUGGCGACC---AUCCU---CCUGGUCGCCGGCUCC ......((((......(((....(((.((....)).))).)))..(((((((((---(....---..)))))))))))))) ( -25.20, z-score = -1.90, R) >droEre2.scaffold_4770 5549602 75 + 17746568 GUAGCAGAAGUUUUCACGAAUAUGCGCACAUUUGUUUGCCUUGUCUUGGCGACC---AUCCU---CCUGGUCGCCGGCUCC ..(((............((....(((.((....)).)))....)).((((((((---(....---..)))))))))))).. ( -21.80, z-score = -1.49, R) >droAna3.scaffold_13340 1987389 74 - 23697760 GUGGCAGAAGUUUUCCU-GAAAUGCGCACAUUCGUUUGUCUUGCCCUGGCGGCC---GUCCU---CCUGGUCGCCAGCUCA ((((((..((.....))-....))).)))............((..(((((((((---(....---..))))))))))..)) ( -24.00, z-score = -1.14, R) >dp4.chr2 16874306 80 - 30794189 GUGAAAGAAGUUUGUCC-AAAAUGCGCACAUUUACUUGCCUCGUCCUGGCGGCCCUUAUCCUUGCCCUGGUCGCCAGCUCC ......((.((..((..-.(((((....)))))))..)).))(..(((((((((..............)))))))))..). ( -19.34, z-score = -0.73, R) >droPer1.super_0 8305525 80 + 11822988 GUGAAAGAAGUUUGUCC-AAAAUGCGCACAUUUAUUUGCCUCGUCCUGGCGGCCCUCGUCCUUGCCCUGGUCGCCAGCACC (..((..((((.(((.(-.....).))).)))).))..)...((.(((((((((...((....))...))))))))).)). ( -21.10, z-score = -0.90, R) >droMoj3.scaffold_6500 13360965 72 + 32352404 GUGGCUCAAGUGCCAGCGCCAACGCAAAUGCCAAUGCGCAAUCAUUCGGCGGUA---AU------CGCGGUGGCGGCUUCG .((((......)))).(((((.(((...((((((((......)))).))))...---..------.))).)))))...... ( -25.40, z-score = -0.03, R) >consensus GUAGCAGAAGUUUCCUCGAAAAUGCGCACAUUUGUUUGCCUUGCCCUGGCGACC___AUCCU___CCUGGUCGCCGGCUCA .......................(((.((....)).)))......(((((((((..............))))))))).... (-12.74 = -12.74 + 0.00)

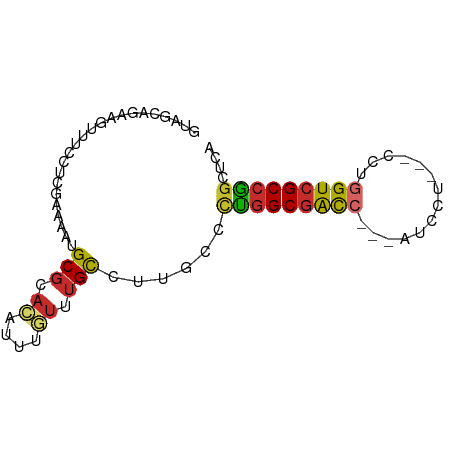

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:12:37 2011