| Sequence ID | dm3.chr3R |
|---|---|
| Location | 9,362,838 – 9,362,933 |
| Length | 95 |
| Max. P | 0.972892 |

| Location | 9,362,838 – 9,362,933 |
|---|---|
| Length | 95 |
| Sequences | 9 |
| Columns | 115 |
| Reading direction | reverse |
| Mean pairwise identity | 71.67 |
| Shannon entropy | 0.54827 |
| G+C content | 0.48481 |
| Mean single sequence MFE | -32.94 |
| Consensus MFE | -12.10 |
| Energy contribution | -13.02 |
| Covariance contribution | 0.91 |
| Combinations/Pair | 1.13 |
| Mean z-score | -2.63 |
| Structure conservation index | 0.37 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.88 |
| SVM RNA-class probability | 0.972892 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 9362838 95 - 27905053 -----------------CGUGG-CCCAUGUUGCAUUUACUUUGCAUGCAACAGUUCGCUGCUUUUUCGCAUACAAGUGACCCAAUUACGGGAAAGGCAGUGGGUUGAUGGCCU-- -----------------...((-((..(((((((.......)))).))).(((..((((((((((((((......))))(((......)))))))))))))..)))..)))).-- ( -37.90, z-score = -3.22, R) >droSim1.chr3R 15454350 95 + 27517382 -----------------CGUGC-CCCAUGUUGCAUGUACUUUGCAUGCAACAGUUCGCUGCUUUUUCGCAUACAAGUGACCCAAUUACGGGAAAGGCAGUGGGUUGAUGGGCU-- -----------------...((-((...(((((((((.....)))))))))((..((((((((((((((......))))(((......)))))))))))))..))...)))).-- ( -42.90, z-score = -4.36, R) >droSec1.super_0 8476361 95 + 21120651 -----------------CGUGC-CCCAUGUUGCAUGUACUUUGCAUGCAACAGUUCGCUGCUUUUUCGCAUACAAGUGACCCAAUUACGGGAAAGGCAGUGGGUUGAUGGGCU-- -----------------...((-((...(((((((((.....)))))))))((..((((((((((((((......))))(((......)))))))))))))..))...)))).-- ( -42.90, z-score = -4.36, R) >droYak2.chr3R 13669421 93 - 28832112 -----------------CAUGC-CCCAUGUUGCAUGUACUUUGCAUGCAACAGUUCGCUGCUUUUUCGCAUACAAGUGACCCAAUUACGGCAAAGGCAGUGGG--AGUGGGCA-- -----------------..(((-(((.((((((((((.....)))))))))).((((((((((((((((......)))).((......)).))))))))))))--.).)))))-- ( -39.20, z-score = -3.30, R) >droEre2.scaffold_4770 5466697 93 + 17746568 -----------------GAUGC-CCCAUGUUGCAUGUACUUUGCAUGCAACAGUUCGCUGCUUUUUCGCAUACAAGUGACCCAAUUACGGGAAAGGCAGUGGA--AGUGGGCA-- -----------------..(((-(((.((((((((((.....)))))))))).((((((((((((((((......))))(((......)))))))))))))))--.).)))))-- ( -44.30, z-score = -5.33, R) >droAna3.scaffold_13340 1915404 88 - 23697760 ------------CCGUUUAUCC-UUGAUGUUGCAUGUACUUCGCAUGCAACAGUUCGCUGCUUUUUCGCAUACAAGUGACCCAAUUACCCAAGAG--GGGACA------------ ------------..(((..(((-(((.((((((((((.....))))))))))......(((......)))..)))).))........(((....)--))))).------------ ( -25.00, z-score = -2.30, R) >droPer1.super_0 8234499 97 + 11822988 -----------------UCUCC-UGGAUGUCGCAUGUACUUUGCAUGCAACAGUUCGCUGCUUUUUCGCAUAGAAGUGACCCAAUUACAGAGGGGUAGGGGCAGGGGAAGGGGCA -----------------.((((-(...(((.((((((.....)))))).))).(((.((((((((((((......))))(((.........)))..)))))))).)))))))).. ( -32.90, z-score = -1.34, R) >droWil1.scaffold_181130 6433371 112 - 16660200 UAAUAUGUGCAUACGAACAUCCAUACAUACUACCUCCACUUGGCAUGCAACAGUUCGCUGCUUUUCC-CAUACAAGUGACCCAAUUACAGAAAGGGUGGCGCAACAACACUCG-- .....(((((..........................(((((((((.((........)))))......-....))))))((((...........)))).)))))..........-- ( -20.40, z-score = 0.02, R) >droGri2.scaffold_14906 10079343 73 + 14172833 -----------------UCGGC-UAUAUUUC-CAUGUAGUUUACAUGCAACAGUUCGCUGCUUUUUCGCAUACAAGUGACCUAAUUACUGGG----------------------- -----------------.....-.......(-((.((((((.........(((....))).....((((......))))...))))))))).----------------------- ( -11.00, z-score = 0.51, R) >consensus _________________CAUGC_CCCAUGUUGCAUGUACUUUGCAUGCAACAGUUCGCUGCUUUUUCGCAUACAAGUGACCCAAUUACGGGAAAGGCAGUGGA__GAUGGGCU__ ...........................((((((((((.....))))))))))..(((((...............))))).................................... (-12.10 = -13.02 + 0.91)

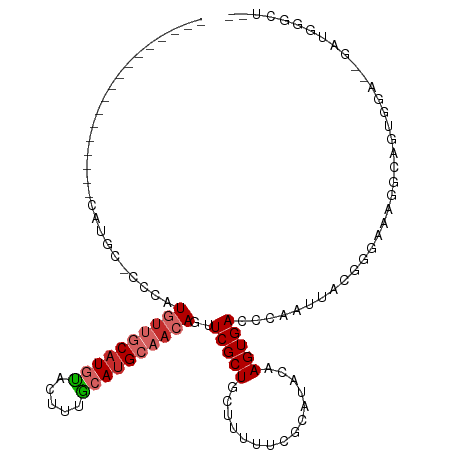
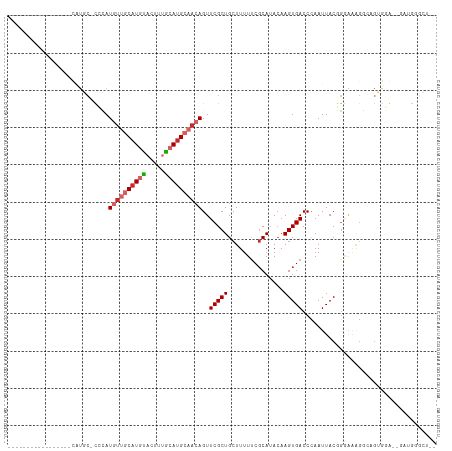
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:12:13 2011