
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 9,144,209 – 9,144,302 |
| Length | 93 |
| Max. P | 0.606708 |

| Location | 9,144,209 – 9,144,302 |
|---|---|
| Length | 93 |
| Sequences | 3 |
| Columns | 105 |
| Reading direction | reverse |
| Mean pairwise identity | 51.46 |
| Shannon entropy | 0.67964 |
| G+C content | 0.50689 |
| Mean single sequence MFE | -23.27 |
| Consensus MFE | -6.01 |
| Energy contribution | -6.90 |
| Covariance contribution | 0.89 |
| Combinations/Pair | 1.22 |
| Mean z-score | -1.91 |
| Structure conservation index | 0.26 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.24 |
| SVM RNA-class probability | 0.606708 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 9144209 93 - 27905053 -GUCUCACCGCAAAUCG--------GUGAUCGAGCACCCUCACCUUCAACUAACUCGA---AACUUUUGCCCUCAGAGAACGACAGACCGCGUUCUUGCUCUCCC -...((((((.....))--------))))..((((.........(((.........))---).............(((((((........))))))))))).... ( -20.30, z-score = -1.71, R) >droYak2.chr2R 4794119 96 - 21139217 -UUCUCAACUAGAGAUAU--AGAUCAAAAUCAGA---AGUCGACAAAGAGCAACUCGA---CUCUACUGACCGUCGACAGCAAGAGAGCCAGCUCUUGCUCACCC -.((((.....))))...--..............---.((((((...(((...)))..---.((....))..))))))((((((((......))))))))..... ( -24.50, z-score = -2.01, R) >droAna3.scaffold_13137 462583 105 + 1373759 AAAGCAACCGAAAACCAUGCAAGCUCUGAUCAAGCGCGGUCAAUUACCCCCAAAUUGACCGAAUUCUUAAACGUAGAGAGCGAGAGAGCGCGUUCCUGCUCCCCC ..((((...((.....((((..(((((..((..((.(((((((((.......)))))))))...((((.......)))))))).))))))))))).))))..... ( -25.00, z-score = -2.02, R) >consensus _AUCUCACCGAAAAACAU__A____AUGAUCAAGC_CAGUCAACUACAACCAACUCGA___AACUAUUGACCGUAGAGAGCGAGAGAGCGCGUUCUUGCUCACCC .............................................................................(((((((((......))))))))).... ( -6.01 = -6.90 + 0.89)

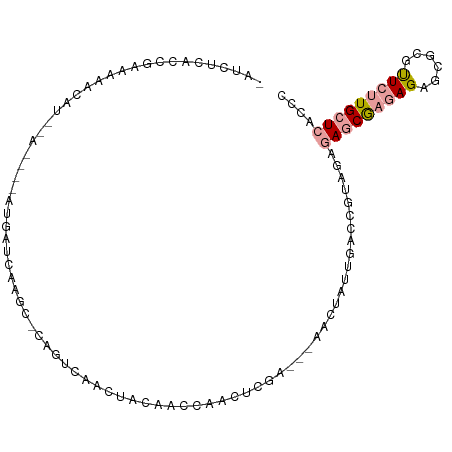
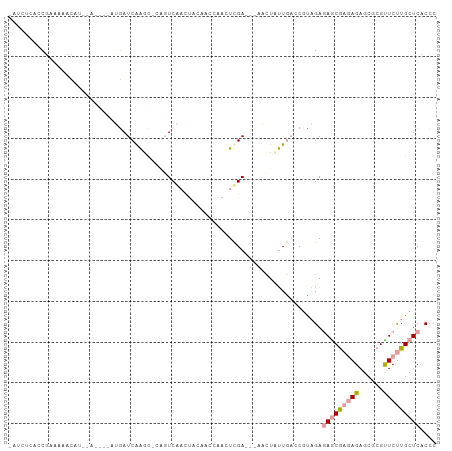
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:11:26 2011