
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 8,832,749 – 8,832,929 |
| Length | 180 |
| Max. P | 0.691070 |

| Location | 8,832,749 – 8,832,861 |
|---|---|
| Length | 112 |
| Sequences | 12 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 72.23 |
| Shannon entropy | 0.60632 |
| G+C content | 0.58468 |
| Mean single sequence MFE | -43.08 |
| Consensus MFE | -17.57 |
| Energy contribution | -17.49 |
| Covariance contribution | -0.08 |
| Combinations/Pair | 1.62 |
| Mean z-score | -1.32 |
| Structure conservation index | 0.41 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.27 |
| SVM RNA-class probability | 0.621041 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 8832749 112 - 27905053 CGGAAUGCCCUCCAGAUAGGAAUUGAAACCGGUGAUGGUGGGCGGCGGAGAGUCGUCGCCACAGUUGUGAUGCAUGUGGCAAUCGGUCGAUACGCCGCCGUUUCCAGGACCG------ .((((.((((.(((....((........)).....))).))))(((((.(.((((.(((((((((......)).)))))).....).)))).).)))))..)))).......------ ( -44.70, z-score = -1.47, R) >droSim1.chr3R 14927589 112 + 27517382 CGGAAUGCCCUCCAGAUAGGAAUUGAAACCGGUGAUGGUGGGCGGCGGCGAGUCGUCGCCACAGUUGUGAUGCAUGUGGCAAUCGGUCGUUCCGCCGCCGUUUCCAGGACCG------ .((((.((((.(((....((........)).....))).))))(((((((.((((..((((((((......)).))))))...))).)....)))))))..)))).......------ ( -47.40, z-score = -1.56, R) >droSec1.super_0 7956995 112 + 21120651 CGGAAUGCCCUCCAGAUAGGAAUUGAAACCGGUGAUGGUGGGCGGCGGAGAGUCGUCGCCACAGUUGUGAUGCAUGUGGCAAUCGGUCGUUCCGCCGCCGUUUCCAGGACCG------ .(((......))).....((........))(((..(((..((((((((((.((((..((((((((......)).))))))...))).).))))))))))....)))..))).------ ( -46.90, z-score = -1.83, R) >droYak2.chr3R 13127000 112 - 28832112 CGGAAUGCCCUCCAGGUAGGAAUUGAAACCGGUGAUGGUGGGCGGCGGAGAGUCGUCGCCACAGCUGUGAUGCAUGUGGCAAUCGCUCGAUCCACCGCCGCUUCCAGAACCC------ .(((......))).(((..........)))(((..(((.(((((((((.((((.((.((((((((......)).)))))).)).))))......))))))))))))..))).------ ( -47.60, z-score = -2.04, R) >droEre2.scaffold_4770 4930157 112 + 17746568 CGGAAUGCCCUCCAGGUAGGAAUUGAAACCGGUGAUGGUGGGCGGCGGAGAGUCGUCGCCACAGCUGUGAUGCAUGUGGCAAUCGCUUGAUCCACCGCCGCUUCCAGAACCC------ .(((......))).(((..........)))(((..(((.(((((((((.((((.((.((((((((......)).)))))).)).))))......))))))))))))..))).------ ( -45.80, z-score = -1.73, R) >droAna3.scaffold_13340 14352455 109 + 23697760 GGGUAUGCCCUCCAGGUAGGAAUUGAAACCAGUGAUCUGUGGUGGCGGAGAAUCGUCGCCGCAGCUGUGAUGCAUGUGACAGUCGCUGGA---ACCACCGCCGCUAGAGCCA------ (((....)))....(((.((...((...(((((((.(((((((((((......)))))))))))((((.((....)).))))))))))).---..)))))))((....))..------ ( -41.60, z-score = -1.27, R) >dp4.chr2 19457301 109 + 30794189 CGGUAUGCCCUCAAGGUAGGAAUUGAAACCAGUGAUGGUGGGCGGCGGAGAGUCGUCGCCACAGCUGUGAUGCAUGUGGCAUUCGCCUUGUCCGCUGCUCUGACCGCCA--------- .(((.((((.....))))((.......((((....))))(((((((((((((.((..((((((((......)).))))))...)).))).))))))))))...))))).--------- ( -49.00, z-score = -2.64, R) >droWil1.scaffold_181089 11570844 112 - 12369635 GGGUAUACCCUCAAGAUAGGAAUUGAAACCGGUGAUUGUUGGUGGUGGUGAAUCAUCGCCGCAAUUAUGAUGCAUAUGACAGUCUCCAGG---GCCGCCGUUGCCACCGCCAACG--- (((....)))((((........))))...........(((((((((((..(......((((((.......)))...((........)).)---)).....)..))))))))))).--- ( -35.90, z-score = -0.59, R) >droVir3.scaffold_12855 6178095 112 + 10161210 UGGUAUGCCCUCAAGAUAGGAAUUGAAACCGGUGAUCGUUGGCGGCGGCGAAUCAUCGCCACAAUUAUGAUGCAUGUGACAAUCACCAGG---AGCACCAGGCCCGGCGCCACCA--- .(((.((((((.......((........))((((((.((((.(((((((((....))))).((....))..)).)))))).)))))))))---.)))...(((.....)))))).--- ( -34.40, z-score = 0.22, R) >droMoj3.scaffold_6540 25858043 118 + 34148556 GGGUAUGCCCUCGAGAUAGGAAUUGAAACCGGUGAUGGUUGGAGGCGGCGAGUCGUCGCCACAAUUAUGAUGCAUGUGACAGUCGCCUGGGCCAGCGCCGGGUCCGUUGCCACCACCA (((....)))........(((((.((..((((((.(((((.(.((((((..((((..((..((....))..))...)))).))))))).))))).)))))).)).))).))....... ( -41.50, z-score = 0.59, R) >droGri2.scaffold_14906 7569689 106 + 14172833 GGGUAUUCCUUCCAGGUAGGAAUUGAAACCGGUGAUUGUUGGUGGCGGCGAAUCGUCCCCACAAUUAUGAUGCAUGUGACAAUCACCGGG---ACCGCCGGGUCCACCA--------- .(((((((((.......))))))......(((((((((((.((((.((((...)))).))))...((((...)))).)))))))))))((---(((....)))))))).--------- ( -41.80, z-score = -1.55, R) >anoGam1.chr2R 6288341 102 + 62725911 -GGCAAGCUUCCAAGG------CUGUUGCCGUUGAAGAAUGCGCUCGGUGCUUCAUCGUCGCAAC---AGUGUGUGGCCAAGCUGGUGGAUGCAUCUGCUCCACCAAGAUAC------ -....(((((((((.(------(((((((...(((.(((.((((...))))))).)))..)))))---))).).)))..)))))((((((.((....)))))))).......------ ( -40.40, z-score = -2.00, R) >consensus CGGUAUGCCCUCCAGAUAGGAAUUGAAACCGGUGAUGGUGGGCGGCGGAGAGUCGUCGCCACAGUUGUGAUGCAUGUGGCAAUCGCUCGAUCCACCGCCGUUUCCAGCACCA______ .(((....(((......)))..........((((......((((((((....))))))))........(((..........)))...........))))...)))............. (-17.57 = -17.49 + -0.08)



| Location | 8,832,821 – 8,832,929 |
|---|---|
| Length | 108 |
| Sequences | 12 |
| Columns | 115 |
| Reading direction | forward |
| Mean pairwise identity | 82.37 |
| Shannon entropy | 0.38988 |
| G+C content | 0.51654 |
| Mean single sequence MFE | -30.92 |
| Consensus MFE | -14.20 |
| Energy contribution | -14.77 |
| Covariance contribution | 0.57 |
| Combinations/Pair | 1.43 |
| Mean z-score | -2.20 |
| Structure conservation index | 0.46 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.43 |
| SVM RNA-class probability | 0.691070 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 8832821 108 + 27905053 -------CACCAUCACCGGUUUCAAUUCCUAUCUGGAGGGCAUUCCGAACACCGGAGUCAUUCGCUACGACGACGCCGCCUUUCUCAAGAACUUGAUACCGGGCCAGCAUCUCAA -------........(((((.(((((((......(((((((..((((.....))))(((..(((...))).)))...)))))))....))).)))).)))))............. ( -31.40, z-score = -2.17, R) >droSim1.chr3R 14927661 108 - 27517382 -------CACCAUCACCGGUUUCAAUUCCUAUCUGGAGGGCAUUCCGAACACCGGAGUCAUUCGCUACGACGACGCCGCCUUCCUCAAGAACUUGAUACCGGGCCAGCAUCUCAA -------........(((((.(((((((......(((((((..((((.....))))(((..(((...))).)))...)))))))....))).)))).)))))............. ( -34.30, z-score = -2.99, R) >droSec1.super_0 7957067 108 - 21120651 -------CACCAUCACCGGUUUCAAUUCCUAUCUGGAGGGCAUUCCGAACACCGGAGUCAUUCGCUACGACGACGCCGCCUUCCUCAAGAACUUGAUACCGGGCCAGCAUCUCAA -------........(((((.(((((((......(((((((..((((.....))))(((..(((...))).)))...)))))))....))).)))).)))))............. ( -34.30, z-score = -2.99, R) >droYak2.chr3R 13127072 108 + 28832112 -------CACCAUCACCGGUUUCAAUUCCUACCUGGAGGGCAUUCCGAACACCGGAGUGAUUCGCUACGACGACGCCGCCUUCCUCAAGAACUUGAUACCGGGCCAGCAUCUCAA -------........(((((.(((((((......(((((((((((((.....))))))...(((......)))....)))))))....))).)))).)))))............. ( -34.40, z-score = -2.64, R) >droEre2.scaffold_4770 4930229 108 - 17746568 -------CACCAUCACCGGUUUCAAUUCCUACCUGGAGGGCAUUCCGAACACCGGAGUGAUUCGCUACGACGACGCCGCCUUCCUCAAGAACUUGAUACCGGGCCAGCAUCUCAA -------........(((((.(((((((......(((((((((((((.....))))))...(((......)))....)))))))....))).)))).)))))............. ( -34.40, z-score = -2.64, R) >droAna3.scaffold_13340 14352524 108 - 23697760 ------ACAG-AUCACUGGUUUCAAUUCCUACCUGGAGGGCAUACCCAACACCGGCGUUAUACGCUACGACGACGCCGCCUUCCUAAAGAACUUGAUACCGGGACAGCAUCUCAA ------....-.((.(((((.(((((((......(((((((............(((((..(.((...)))..))))))))))))....))).)))).)))))))........... ( -29.80, z-score = -2.23, R) >dp4.chr2 19457370 108 - 30794189 -------CACCAUCACUGGUUUCAAUUCCUACCUUGAGGGCAUACCGAACACCGGGGUCAUACGCUACGACGACGCCGCCUUCCUCAAGAACUUGAUACCGGGACAGCAUCUCAA -------.....((.(((((.(((((((.......((((((...(((.....)))(((....((......))..))))))))).....))).)))).)))))))........... ( -26.50, z-score = -0.85, R) >droWil1.scaffold_181089 11570916 108 + 12369635 -------AACAAUCACCGGUUUCAAUUCCUAUCUUGAGGGUAUACCCAAUACGGGCGUUAUACGUUACGAUAAUGCGGCAUUCCUUAAGAAUUUGAUACCAGGUCAACAUCUUAA -------.......((((((.((((......(((((((((.....((.....))(((((((.((...))))))))).....)))))))))..)))).))).)))........... ( -26.00, z-score = -1.82, R) >droVir3.scaffold_12855 6178167 108 - 10161210 -------AACGAUCACCGGUUUCAAUUCCUAUCUUGAGGGCAUACCAAACACUGGCGUCAUACGCUAUGACGAUGCGGCGUUCCUCAAGAAUUUGAUACCGGGCCAGCAUCUCAA -------........(((((.((((......((((((((((((.(((.....)))(((((((...)))))))))))......))))))))..)))).)))))............. ( -33.20, z-score = -2.57, R) >droMoj3.scaffold_6540 25858121 108 - 34148556 -------AACCAUCACCGGUUUCAAUUCCUAUCUCGAGGGCAUACCCAACACCGGCGUCAUUCGCUAUGACGAUGCGGCCUUCCUCAAGAAUUUGAUACCGGGCCAGCAUCUCAA -------........(((((......((.......))(((....)))...))))).(((((.....)))))((((((((((...((((....))))....))))).))))).... ( -31.00, z-score = -2.12, R) >droGri2.scaffold_14906 7569755 108 - 14172833 -------AACAAUCACCGGUUUCAAUUCCUACCUGGAAGGAAUACCCAACACUGGCGUCAUACGCUAUGAUGACGCCGCAUUCCUCAAAAACUUGAUACCGGGUCAGCAUCUCAA -------........(((((.((((((((.....))))(((((.........(((((((((........))))))))).)))))........)))).)))))............. ( -31.60, z-score = -3.67, R) >anoGam1.chr2R 6288410 111 - 62725911 AUUCUUCAACGGCAACAGCCUUGGAAGCUUGCCCACCGGG-ACCAUUGGCACUCGA---ACACGAUAUGACGAUCUUGCCUUCCUGCAAAACUUACAACCGGGCCAACGGCUUAA .....((((.(((....)))))))(((((.((((..((((-(.....((((.(((.---...........)))...)))).)))))..............))))....))))).. ( -24.09, z-score = 0.28, R) >consensus _______CACCAUCACCGGUUUCAAUUCCUACCUGGAGGGCAUACCGAACACCGGAGUCAUACGCUACGACGACGCCGCCUUCCUCAAGAACUUGAUACCGGGCCAGCAUCUCAA ...............(((((.((((.......((.(((((....(((.....)))........((..((....))..))..))))).))...)))).)))))............. (-14.20 = -14.77 + 0.57)

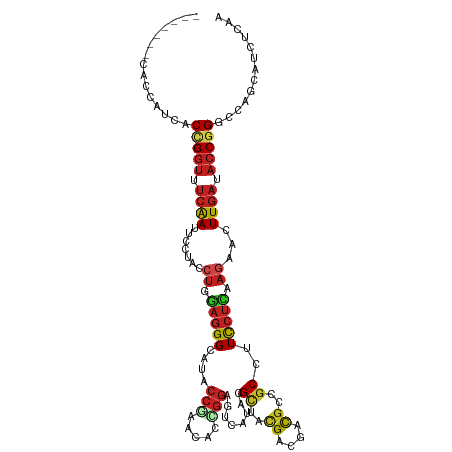

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:10:46 2011