
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 8,713,426 – 8,713,529 |
| Length | 103 |
| Max. P | 0.631168 |

| Location | 8,713,426 – 8,713,529 |
|---|---|
| Length | 103 |
| Sequences | 9 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 68.05 |
| Shannon entropy | 0.62711 |
| G+C content | 0.46754 |
| Mean single sequence MFE | -25.60 |
| Consensus MFE | -8.58 |
| Energy contribution | -8.26 |
| Covariance contribution | -0.33 |
| Combinations/Pair | 1.59 |
| Mean z-score | -1.64 |
| Structure conservation index | 0.34 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.29 |
| SVM RNA-class probability | 0.631168 |
| Prediction | RNA |
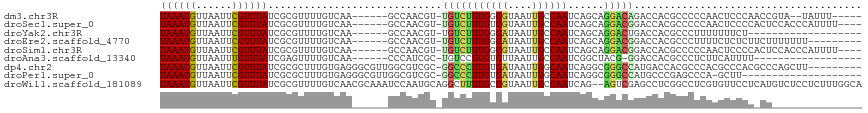
Download alignment: ClustalW | MAF
>dm3.chr3R 8713426 103 - 27905053 UAAAUGUUAAUUCGUUUAUCGCGUUUUGUCAA------GCCAACGU-UGUCUUUGGCGUAAUUGCCAAUCAGCAGGACAGACCACGCCCCCAACUCCCAACCGUA--UAUUU----- ((((((......))))))..((((((((((..------(....)((-((...((((((....)))))).))))..))))))..))))..................--.....----- ( -22.30, z-score = -1.85, R) >droSec1.super_0 7844466 106 + 21120651 UAAAUGUUAAUUCGUUUAUCGCGUUUUGUCAA------GCCAACGU-UGUCUUUGUCGUAAUUGCCAAUCAGCAGGACGGACCACGCCCCCAACUCCCCACUCCACCCAUUUU---- ((((((......))))))..((((...(((..------.((...((-((...(((.((....)).))).)))).))...))).))))..........................---- ( -15.90, z-score = -0.06, R) >droYak2.chr3R 13009164 91 - 28832112 UAAAUGUUAAUUCGUUUAUCGCGUUUUGUCAA------GCCAACGU-UGUCUUUGGCAUAAUUGCCAAUCAGCAGGACUGACCACGCCCUUUUUUUCU------------------- ((((((......))))))..((((...((((.------.((...((-((...((((((....)))))).)))).))..)))).))))...........------------------- ( -25.00, z-score = -3.51, R) >droEre2.scaffold_4770 4812963 101 + 17746568 UAAAUGUUAAUUCGUUUAUCGCGUUUUGUCAA------GCCAACGU-UGUCUUUGGCAUAAUUGCCAAUCAGCAGGACGGACCACGCCCUUUUCUCUCUUCUUUUUUU--------- ((((((......))))))..((((...(((..------.((...((-((...((((((....)))))).)))).))...))).)))).....................--------- ( -23.70, z-score = -2.66, R) >droSim1.chr3R 14814028 106 + 27517382 UAAAUGUUAAUUCGUUUAUCGCGUUUUGUCAA------GCCAACGU-UGUCUUUGGCGUAAUUGCCAAUCAGCAGGACGGACCACGCCCCCAACUCCCCACUCCACCCAUUUU---- ((((((......))))))..((((...(((..------.((...((-((...((((((....)))))).)))).))...))).))))..........................---- ( -23.00, z-score = -1.70, R) >droAna3.scaffold_13340 14246645 91 + 23697760 UAAAUGUUAAUUUGUUUAUCGAGUUUUGUCAA------CCCAUCGC-UGUCCUUGUCUUAAUUGCCAAUCGGCUACG-GGACCACGCCCUCUUCAUUUU------------------ ....................(((...(((...------(((...((-((...(((.(......).))).))))...)-))...)))..)))........------------------ ( -11.90, z-score = 0.66, R) >dp4.chr2 19323262 107 + 30794189 UAAAUGUUAAUUCGUUUAUCGCGCUUUGUGAGGGCGUUGGCGUCGC-GGCCCUUGUCAUAAUUGGCAAUCAGGCGGGCCAUGACCACGCCCACGCCCACGCCCAGCUU--------- ((((((......))))))....(((..(((.((((((.(((((((.-(((((((((((....))))))......))))).))))...))).)))))).)))..)))..--------- ( -45.40, z-score = -2.75, R) >droPer1.super_0 9047557 95 - 11822988 UAAAUGUUAAUUCGUUUAUCGCGCUUUGUGAGGGCGUUGGCGUCGC-GGCCCUUGUCAUAAUUGGCAAUCAGGCGGGCCAUGCCCGAGCCCA-GCUU-------------------- ((((((......))))))((((.....))))((((...(((((...-(((((((((((....))))))......))))))))))...)))).-....-------------------- ( -34.30, z-score = -0.69, R) >droWil1.scaffold_181089 11439001 115 - 12369635 UAAAUGUUAAUUCGUUUAUCGCGUUUUGUCAACGCAAAUCCAAUGCAGGCUUUUGCCGUAAUUGCCAAUCAG--AGUCGAGCCUCGGCCUCGUGUUCCUCAUGUCUCCUCUUUGGCA ((((((......))))))..(((((.....)))))........(((.(((....))))))..((((((..((--((..((((....))..((((.....))))))..)))))))))) ( -28.90, z-score = -2.17, R) >consensus UAAAUGUUAAUUCGUUUAUCGCGUUUUGUCAA______GCCAACGU_UGUCUUUGGCAUAAUUGCCAAUCAGCAGGACGGACCACGCCCCCAUCUCCCC_CU_______________ ((((((......)))))).............................(((((((((((....))))))......)))))...................................... ( -8.58 = -8.26 + -0.33)


Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:10:27 2011