
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 7,155,149 – 7,155,245 |
| Length | 96 |
| Max. P | 0.769458 |

| Location | 7,155,149 – 7,155,245 |
|---|---|
| Length | 96 |
| Sequences | 7 |
| Columns | 113 |
| Reading direction | reverse |
| Mean pairwise identity | 72.19 |
| Shannon entropy | 0.54834 |
| G+C content | 0.41040 |
| Mean single sequence MFE | -28.69 |
| Consensus MFE | -11.32 |
| Energy contribution | -11.76 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.45 |
| Mean z-score | -1.87 |
| Structure conservation index | 0.39 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.63 |
| SVM RNA-class probability | 0.769458 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
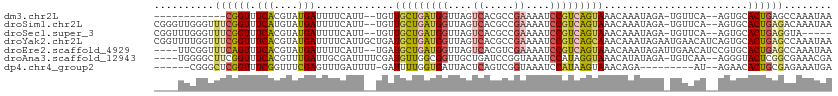
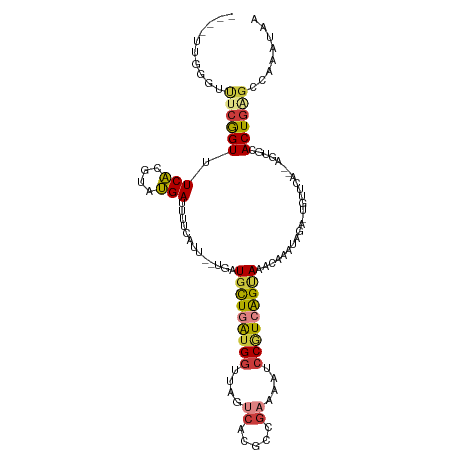
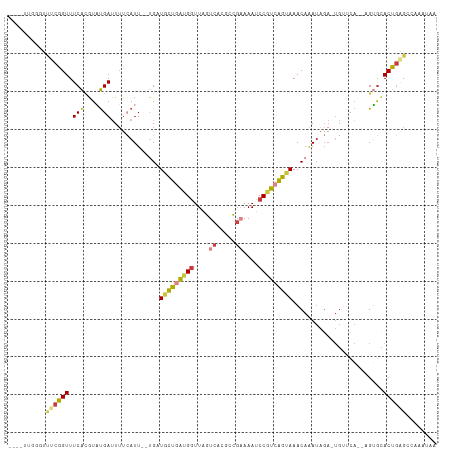
>dm3.chr2L 7155149 96 - 23011544 ------------CGGUUUCACGUAUGAUUUUCAUU--UGUUGCUGAUGGUUAGUCACGCCGAAAAUCCGUCAGUAAACAAAUAGA-UGUUCA--AGUGCACUGAGCCAAAUAA ------------.((((((((...(((...(((((--((((((((((((....((.....))....)))))))).))))))).))-...)))--.)))....)))))...... ( -28.40, z-score = -3.25, R) >droSim1.chr2L 6944550 108 - 22036055 CGGGUUGGGUUUCGGUUUCAUGUAUGAUUUUCAUU--UGUUGCUGAUGGUUAGUCACGCCGAAAAUCCGUCAGUAAACAAAUAGA-UGUUCA--AGUGCACUGAGACAAAUAA ........((((((((..(((...(((...(((((--((((((((((((....((.....))....)))))))).))))))).))-...)))--.))).))))))))...... ( -32.20, z-score = -2.77, R) >droSec1.super_3 2678711 103 - 7220098 CGGUUUGGGUUUCGGUUUCACGUAUGAUUUUCAUU--UGUUGCUGAUGGUUAGUCACGCCGAAAAUCCGUCAGUAAACAAAUAGA-UGUUCA--AGUGCACUGAGGUA----- ........(..(((((..(((...(((...(((((--((((((((((((....((.....))....)))))))).))))))).))-...)))--.))).)))))..).----- ( -29.60, z-score = -2.18, R) >droYak2.chr2L 16573983 113 + 22324452 CGGUUUUGGUUUCGGUUUCACGUAUGAUUUUCAUUGCUGAUGCUGAUGGUUAGUCACGCCGAAAAUCCGUCAGCAAACAAAUAGAAUGAACAUCAGUGCACUGAGCCAAAUAA ....(((((((.((((..(((...((((.((((((.(((.(((((((((....((.....))....)))))))))......))))))))).))))))).)))))))))))... ( -34.70, z-score = -2.67, R) >droEre2.scaffold_4929 16070497 107 - 26641161 ----UUCGGUUUCAGUUUCACGUAUGAUUUUCAUU--UGAUGCUGAUGGUUAGUCACGUCGAAAAUCCGUCAGUAAACAAAUAGAUUGAACAUCCGUGCACUGAGCCAAAUAA ----...((((.((((..((((.(((.((.(((((--((.(((((((((....((.....))....)))))))))..))))).))..)).))).)))).))))))))...... ( -32.50, z-score = -3.68, R) >droAna3.scaffold_12943 1003914 106 + 5039921 ----UGGGGCUUCGGUUUCACGUUUGAUUGCGAUUUUCGAUGUUGGCGGUUGCUGAUCCGGUAAAUCCAUAGGUAAACAUAUAGA-UGUCAA--AGGGUACUCGGCGAAACGA ----((..((....))..))(((((((((((.(..........).))))))((((((((((.....))........((((....)-)))...--.)))...))))).))))). ( -25.70, z-score = 0.53, R) >dp4.chr4_group2 793558 95 - 1235136 ------CGGGCUCGGUUUCGGUUUCGAGUUUGAUUUU-GAUUUUGGUGAUUACUCAGUCGGUAAAUCCAUAAGUAAACAGA---------AU--AGAACACUGCGAGAAAUGA ------((..(((((((((...(((..(((((....(-(..((((.(((((....))))).))))..))....))))).))---------).--.))).))..))))...)). ( -17.70, z-score = 0.93, R) >consensus ____UUGGGUUUCGGUUUCACGUAUGAUUUUCAUU__UGAUGCUGAUGGUUAGUCACGCCGAAAAUCCGUCAGUAAACAAAUAGA_UGUUCA__AGUGCACUGAGCCAAAUAA ..........((((((.(((....))).............(((((((((....((.....))....)))))))))........................))))))........ (-11.32 = -11.76 + 0.44)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:23:02 2011