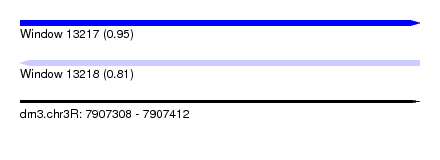

| Sequence ID | dm3.chr3R |
|---|---|
| Location | 7,907,308 – 7,907,412 |
| Length | 104 |
| Max. P | 0.949337 |

| Location | 7,907,308 – 7,907,412 |
|---|---|
| Length | 104 |
| Sequences | 5 |
| Columns | 104 |
| Reading direction | forward |
| Mean pairwise identity | 89.71 |
| Shannon entropy | 0.18672 |
| G+C content | 0.38265 |
| Mean single sequence MFE | -19.22 |
| Consensus MFE | -16.33 |
| Energy contribution | -17.58 |
| Covariance contribution | 1.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.21 |
| Structure conservation index | 0.85 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.55 |
| SVM RNA-class probability | 0.949337 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
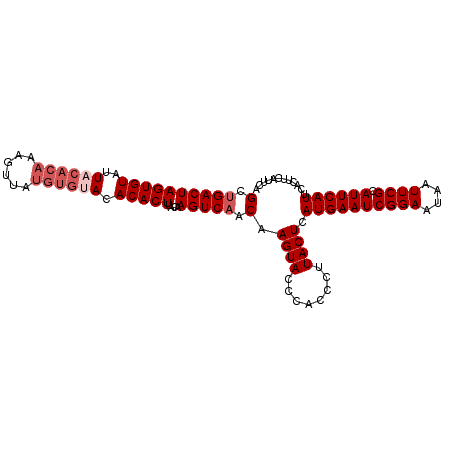
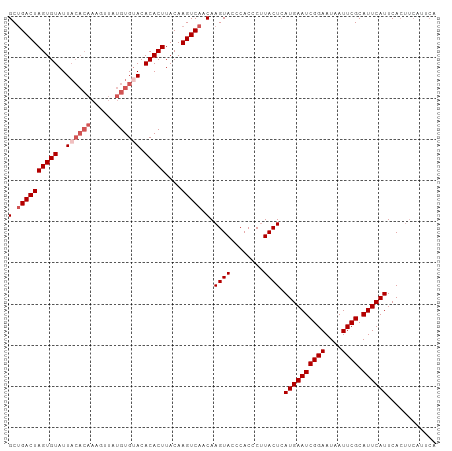
>dm3.chr3R 7907308 104 + 27905053 GCUGACUAGUGUAUUACACAAAGUUAUGUGCACACACUUACAAGUCAACAAGUACCCACCCCUACUCAUGAAUCGGAAUAAUUCGCAUUCAUUCACUUCAUUCA (.((((((((((....((((......))))...)))))....))))).).((((........)))).((((((((((....)))).))))))............ ( -17.40, z-score = -1.62, R) >droEre2.scaffold_4770 4030170 95 - 17746568 GCUGACUAGUGUAUUACACAAAGUUAUGUGUACACACUUACAAGUCAACAAGUACCCACCCUUACUGAUGAAUCGGAAUAAUUCGCAUUCAUUCA--------- (.((((((((((..((((((......)))))).)))))....))))).).((((........))))(((((((((((....)))).)))))))..--------- ( -22.40, z-score = -2.73, R) >droYak2.chr3R 12217646 92 + 28832112 GCAGACUAGUGUAUU------------GUGUACACACUUACAAGUCAACAAGUACUCACCCUUACUCAUGAAUCGGAAUAAUUCGCAUUCAUUCACUUCAUUCA ...((((..((((..------------(((....))).))))))))....((((........)))).((((((((((....)))).))))))............ ( -15.10, z-score = -1.33, R) >droSec1.super_0 7051925 104 - 21120651 GCUGACUAGUGUAUUACACAAAGUUAUGUGUACACACUUACAAGUCAACAAGUACCCACCCUUACUCAUGAAUCGGAAUAAUUCGCAUUCAUUCACUUCAUUCA (.((((((((((..((((((......)))))).)))))....))))).).((((........)))).((((((((((....)))).))))))............ ( -20.60, z-score = -2.60, R) >droSim1.chr3R 14032019 104 - 27517382 GCUGACUAGUGUAUUACACAAAGCUAUGUGUACACACUUACAAGUCAACAAGUACCCACCCUUACUCAUGAAUCGGAAUAAUUCGCAUUCAUUCACUUCAUUCA (.((((((((((..((((((......)))))).)))))....))))).).((((........)))).((((((((((....)))).))))))............ ( -20.60, z-score = -2.76, R) >consensus GCUGACUAGUGUAUUACACAAAGUUAUGUGUACACACUUACAAGUCAACAAGUACCCACCCUUACUCAUGAAUCGGAAUAAUUCGCAUUCAUUCACUUCAUUCA (.((((((((((..((((((......)))))).)))))....))))).).((((........)))).((((((((((....)))).))))))............ (-16.33 = -17.58 + 1.25)


| Location | 7,907,308 – 7,907,412 |
|---|---|
| Length | 104 |
| Sequences | 5 |
| Columns | 104 |
| Reading direction | reverse |
| Mean pairwise identity | 89.71 |
| Shannon entropy | 0.18672 |
| G+C content | 0.38265 |
| Mean single sequence MFE | -25.88 |
| Consensus MFE | -22.65 |
| Energy contribution | -23.78 |
| Covariance contribution | 1.13 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.57 |
| Structure conservation index | 0.88 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.75 |
| SVM RNA-class probability | 0.806772 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
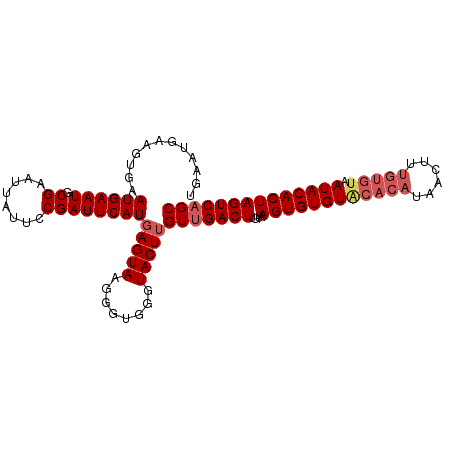
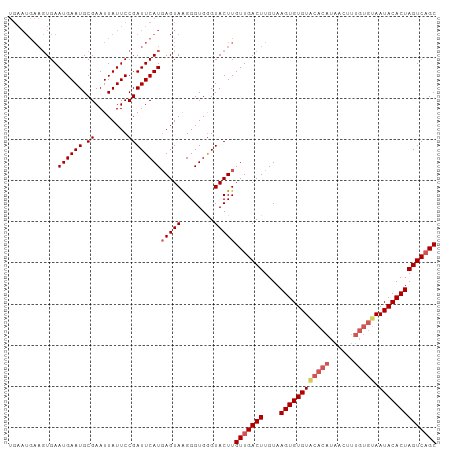
>dm3.chr3R 7907308 104 - 27905053 UGAAUGAAGUGAAUGAAUGCGAAUUAUUCCGAUUCAUGAGUAGGGGUGGGUACUUGUUGACUUGUAAGUGUGUGCACAUAACUUUGUGUAAUACACUAGUCAGC ...(((((..((((((.......))))))...)))))(((((........)))))(((((((....((((((((((((......))))).)))))))))))))) ( -28.20, z-score = -1.83, R) >droEre2.scaffold_4770 4030170 95 + 17746568 ---------UGAAUGAAUGCGAAUUAUUCCGAUUCAUCAGUAAGGGUGGGUACUUGUUGACUUGUAAGUGUGUACACAUAACUUUGUGUAAUACACUAGUCAGC ---------.((((((.......))))))..(((((((......)))))))....(((((((....((((((((((((......))))).)))))))))))))) ( -27.00, z-score = -2.11, R) >droYak2.chr3R 12217646 92 - 28832112 UGAAUGAAGUGAAUGAAUGCGAAUUAUUCCGAUUCAUGAGUAAGGGUGAGUACUUGUUGACUUGUAAGUGUGUACAC------------AAUACACUAGUCUGC ...(((((..((((((.......))))))...)))))(((((........)))))((.((((....(((((((....------------.))))))))))).)) ( -17.60, z-score = 0.22, R) >droSec1.super_0 7051925 104 + 21120651 UGAAUGAAGUGAAUGAAUGCGAAUUAUUCCGAUUCAUGAGUAAGGGUGGGUACUUGUUGACUUGUAAGUGUGUACACAUAACUUUGUGUAAUACACUAGUCAGC ...(((((..((((((.......))))))...)))))(((((........)))))(((((((....((((((((((((......))))).)))))))))))))) ( -28.30, z-score = -2.14, R) >droSim1.chr3R 14032019 104 + 27517382 UGAAUGAAGUGAAUGAAUGCGAAUUAUUCCGAUUCAUGAGUAAGGGUGGGUACUUGUUGACUUGUAAGUGUGUACACAUAGCUUUGUGUAAUACACUAGUCAGC ...(((((..((((((.......))))))...)))))(((((........)))))(((((((....((((((((((((......))))).)))))))))))))) ( -28.30, z-score = -1.99, R) >consensus UGAAUGAAGUGAAUGAAUGCGAAUUAUUCCGAUUCAUGAGUAAGGGUGGGUACUUGUUGACUUGUAAGUGUGUACACAUAACUUUGUGUAAUACACUAGUCAGC ............((((((.((........))))))))(((((........)))))(((((((....((((((((((((......))))).)))))))))))))) (-22.65 = -23.78 + 1.13)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:08:11 2011