
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 7,765,469 – 7,765,566 |
| Length | 97 |
| Max. P | 0.778404 |

| Location | 7,765,469 – 7,765,566 |
|---|---|
| Length | 97 |
| Sequences | 9 |
| Columns | 115 |
| Reading direction | reverse |
| Mean pairwise identity | 53.84 |
| Shannon entropy | 0.95321 |
| G+C content | 0.36350 |
| Mean single sequence MFE | -20.43 |
| Consensus MFE | -3.84 |
| Energy contribution | -3.97 |
| Covariance contribution | 0.13 |
| Combinations/Pair | 1.78 |
| Mean z-score | -1.78 |
| Structure conservation index | 0.19 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.66 |
| SVM RNA-class probability | 0.778404 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 7765469 97 - 27905053 ---------------AGUGAUUCUCAUGAACUUAUCUGGAGUACCUC--GCUUUGCUUUAGAGUCACUUUGUU-UGCUCUAAAUUCCUCGUGGAUUUUUUGCAUACAUAUAUGUG ---------------((((((((....)).....(((((((((....--....))))))))))))))).(((.-(((...(((.(((....))).)))..))).)))........ ( -22.20, z-score = -2.16, R) >droSim1.chr3R 13889719 103 + 27517382 ----AGUAUUUCCCCUAAGAUUCUCAUGAACUUAUCUAGAGUACCUC-------GCUUUAGAGUCACUUUGUU-UGCUCUAAAUUCCUCGUGGAAUUUUUGCAUACAUAUAUGUG ----...........((((.(((....)))))))((((((((.....-------))))))))..(((..(((.-(((...(((((((....)))))))..))).))).....))) ( -24.20, z-score = -3.43, R) >droSec1.super_0 6916372 103 + 21120651 ----AGUAUUUCCCCUAAGAUUUUUAUGAACUUAUCUAGAGUACCUC-------GCUUUAGAGUCACAUGGUU-UGCUCUAAAUUCCUCGUGGAAUUUUUGCAUACAUAUAUGUG ----...........((((.(((....)))))))((((((((.....-------))))))))..((((((((.-(((...(((((((....)))))))..))).))...)))))) ( -23.30, z-score = -2.60, R) >droYak2.chr3R 12177711 104 + 28832112 ----AGUAUUUCCCCUAAGAUUCUCGAGAACUUAUCUAGAGUACCUC--GCUUUGCUUUAGAGUCACUUUGUU-UGCUCUAAAUUCCUCGUGGAAUUUUUGCAUAUAUGUG---- ----...........((((.(((....)))))))(((((((((....--....)))))))))..(((..(((.-(((...(((((((....)))))))..))).))).)))---- ( -23.90, z-score = -2.63, R) >droEre2.scaffold_4770 3891617 104 + 17746568 ----AGUAUUUCCCCUAAGAUACUCGUGAACUUAUCUAGAGUACCUC--GCUUUGCUUUAGAGUCACUUUGUU-UGCUCUAAAUUCCUCGUGGAAUUUUUGCAUAUAUGUG---- ----(((((((......))))))).((((.....(((((((((....--....))))))))).))))..(((.-(((...(((((((....)))))))..))).)))....---- ( -26.90, z-score = -4.02, R) >droAna3.scaffold_12911 2433393 102 - 5364042 ----GGAGUUUCCCCUUCGAUUUUC--GGACUUAUCCGGAGUACUCU--GC-CUGCGUUAUCUUCCCCAUAUUGUGCCUCGAAUUCCUUGCGGAAUUUGCGGAGAUAUGUG---- ----(((((((((.....(.....)--(((....))))))).)))))--..-...............((((((...((.((((((((....)))))))).)).))))))..---- ( -27.00, z-score = -1.32, R) >dp4.chr2 5449986 107 + 30794189 AGUACUUAGGCUCUUCUCUUCCUUGAUUAUCUUGAGUGAAAUCUCUCUAGCAUUGCGUCACUCUCUCACAAUAAUGAGUUACAUUCCUUGCGGAAUAUUUUUUUUCU-------- ..((((((((....((........))....))))))))..................((.((((............)))).))(((((....)))))...........-------- ( -15.80, z-score = 0.14, R) >droWil1.scaffold_181132 844289 96 + 1035393 -----------UACAAAUUGUUCUCACAAUUUUACUU----UUCUUU---UUGGGUUUUUGUUUCUUUUUGUU-UGUUUUAAAUUGCUCAUAUACUUUUCAUCUACACAUGUGUG -----------...(((((((....))))))).....----......---.(((((.((((...(........-.)...))))..)))))...............(((....))) ( -6.40, z-score = 0.59, R) >droMoj3.scaffold_6540 20826740 98 - 34148556 ---------AGUAUUGCAAAGCUGAAAGCCUUUGAGUUAAAUAUAUC--AGCAUUUCGAACAAUGUGAUCAUUAGUUCUCACAUUUUCCACCAAGCAUAUUUAAAAACG------ ---------.....(((...(((((....................))--))).....(((.(((((((..........))))))))))......)))............------ ( -14.15, z-score = -0.61, R) >consensus ____AGUAUUUCCCCUAAGAUUCUCAUGAACUUAUCUAGAGUACCUC__GCUUUGCUUUAGAGUCACUUUGUU_UGCUCUAAAUUCCUCGUGGAAUUUUUGCAUACAUAUG____ ...........................................................................((...(((((((....)))))))..))............. ( -3.84 = -3.97 + 0.13)
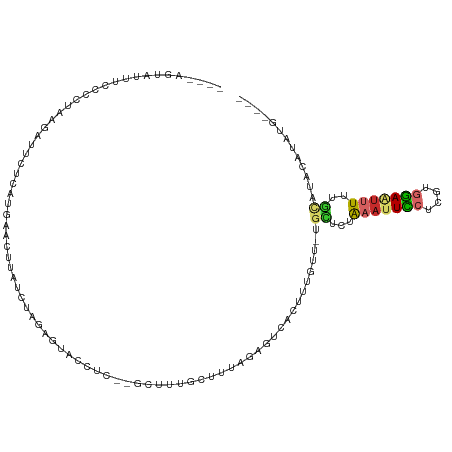


Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:08:00 2011