
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 7,671,766 – 7,671,864 |
| Length | 98 |
| Max. P | 0.927521 |

| Location | 7,671,766 – 7,671,864 |
|---|---|
| Length | 98 |
| Sequences | 3 |
| Columns | 107 |
| Reading direction | reverse |
| Mean pairwise identity | 60.95 |
| Shannon entropy | 0.54914 |
| G+C content | 0.51024 |
| Mean single sequence MFE | -33.14 |
| Consensus MFE | -16.29 |
| Energy contribution | -15.64 |
| Covariance contribution | -0.65 |
| Combinations/Pair | 1.45 |
| Mean z-score | -1.72 |
| Structure conservation index | 0.49 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.33 |
| SVM RNA-class probability | 0.927521 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
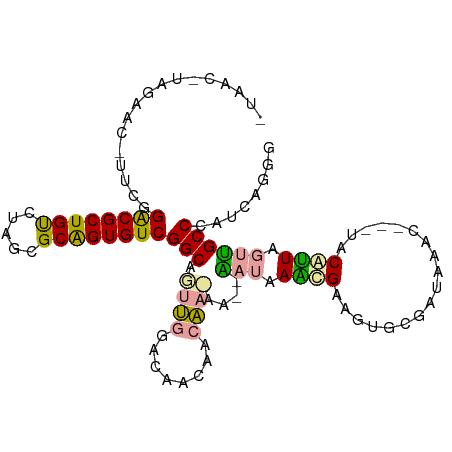
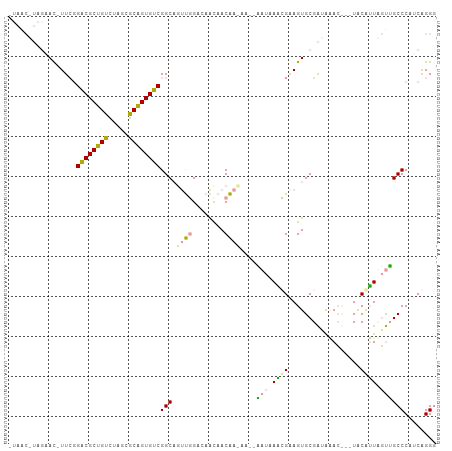
>dm3.chr3R 7671766 98 - 27905053 -------AGUACAUUCGGGCGCUGCUUCGUGCGGUGUCGGCAUUUGGCCAACAACAAAAAGCGUUAAUCGAAGUGCGGUGUAGC--UACGAUACCUGCCCUUCGGGC -------.((((.((((((((((((.....))))))))(((.....)))...................))))))))(((((...--....))))).((((...)))) ( -34.20, z-score = -0.82, R) >droYak2.chrU 26016142 99 + 28119190 AUAACAUAAGAC-UU--GACGCUGUCUAGCGCAGUGUCGGCAGUUGGACAAUAACAACAA--CAUAAGUGUAGUGCGAUAGAC---UGCAUUUGUGGCCCGUCAGGG .........(((-..--((((((((.....))))))))(((.((((........))))..--((((((((((((.(....)))---))))))))))))).))).... ( -37.90, z-score = -3.73, R) >droMoj3.scaffold_6498 1458851 101 + 3408170 -UAACGUAGAAU-UUCGGACGCUGUCCAGCGCAGUGUCGGCUGCCCGA-AGCAACGA-AA--AAUAAAGGGAGUGCGACAAACUAAUACAUUCGUUGCCCAUCAGGG -.......((.(-((((((((((((.....)))))))).(((......-)))..)))-))--......(((...((((.............))))..))).)).... ( -27.32, z-score = -0.60, R) >consensus _UAAC_UAGAAC_UUCGGACGCUGUCUAGCGCAGUGUCGGCAGUUGGACAACAACAA_AA__AAUAAACGAAGUGCGAUAAAC___UACAUUAGUUGCCCAUCAGGG .................((((((((.....))))))))(((.((((........))))....(((.((((..................)))).))))))........ (-16.29 = -15.64 + -0.65)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:07:42 2011