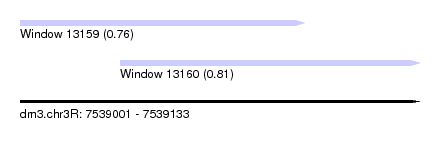

| Sequence ID | dm3.chr3R |
|---|---|
| Location | 7,539,001 – 7,539,133 |
| Length | 132 |
| Max. P | 0.809714 |

| Location | 7,539,001 – 7,539,095 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 83.62 |
| Shannon entropy | 0.25435 |
| G+C content | 0.41770 |
| Mean single sequence MFE | -25.82 |
| Consensus MFE | -20.06 |
| Energy contribution | -21.06 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.69 |
| Structure conservation index | 0.78 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.59 |
| SVM RNA-class probability | 0.755124 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
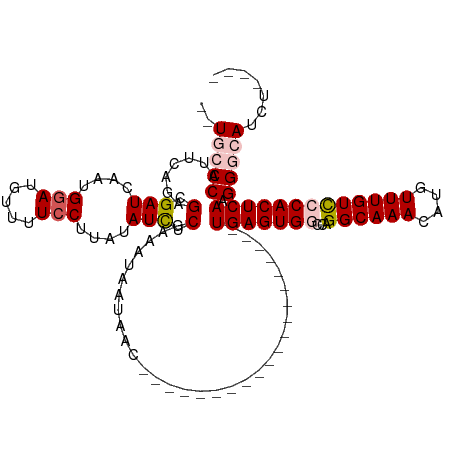
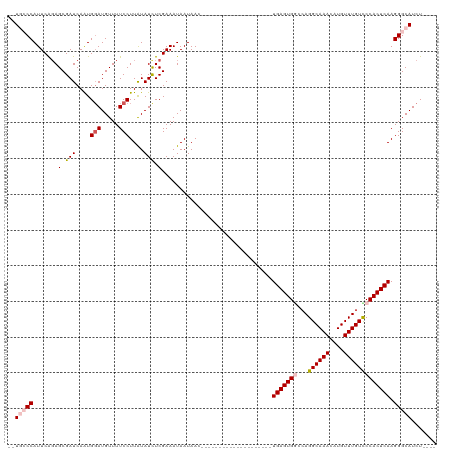
>dm3.chr3R 7539001 94 + 27905053 --UGCCCAUUCAGAGCGAUCAAUGGAUGUUUUCCUUAUAUCUCGAAAUAAUAAC--------------------UGAGUGGCCAGGCAAACAUGUUUGUUCCACUCAAGGGCAUCU---- --(((((.(((.(((........(((.....)))......))))))........--------------------(((((((.((((((....))))))..))))))).)))))...---- ( -29.44, z-score = -2.96, R) >droEre2.scaffold_4770 3668508 98 - 17746568 --UUCCCAUUCAGAGCGAUCAAUGGAUGUUUUCCUUGUAUUUCGAAAUAAUAAC--------------------UGAGUGCCCAGGCAAACGUGUUUGUCCCACUCAAGGGCAUCUAUCU --..(((........(((.(((.(((.....))))))....)))..........--------------------((((((....((((((....)))))).)))))).)))......... ( -22.20, z-score = -0.84, R) >droYak2.chr3R 11949124 116 - 28832112 UUUUGCCAUUCGUAGCGAUCAAUGGAUGUUUUCCUUAUAUCUCGAAAUAAUAACGUAUAUCUCGAAAUAAUAACUGAGUGAAGAGGCAAACAUGUUUGUUCCACUCAAGGGCAUCU---- ...((((((((((........)))))))....((((.(((.((((.(((........))).)))).)))......(((((((.(((((....))))).)).))))))))))))...---- ( -24.90, z-score = -1.13, R) >droSec1.super_0 6693844 94 - 21120651 --UGCCCAUUCAGAGCGAUCAGUGGAUGUUUUCCUUAUAUCUCGAAAUAAUAAG--------------------UGAGUGGCCAGGCAAACAUGUUUGUCCCACUCAAGGGCAUCU---- --(((((.(((.(((........(((.....)))......))))))........--------------------(((((((.((((((....))))))..))))))).)))))...---- ( -29.84, z-score = -2.70, R) >droSim1.chr3R 13660317 94 - 27517382 --UGGCCAUUCAGAGCGAUCAAUGUAUGUUUUCCUUAUAUCUCGAAAUAAUAAG--------------------UGAGUGGCCAGGCAAACAUGUUUGUCCCACUCAAGGGCAUCU---- --((((((((((((....)).............(((((...........)))))--------------------))))))))))((((((....))))))((......))......---- ( -22.70, z-score = -0.83, R) >consensus __UGCCCAUUCAGAGCGAUCAAUGGAUGUUUUCCUUAUAUCUCGAAAUAAUAAC____________________UGAGUGGCCAGGCAAACAUGUUUGUCCCACUCAAGGGCAUCU____ ..(((((.......(.(((....(((.....)))....))).)...............................(((((((...((((((....))))))))))))).)))))....... (-20.06 = -21.06 + 1.00)

| Location | 7,539,034 – 7,539,133 |
|---|---|
| Length | 99 |
| Sequences | 5 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 87.76 |
| Shannon entropy | 0.19770 |
| G+C content | 0.39310 |
| Mean single sequence MFE | -25.44 |
| Consensus MFE | -21.86 |
| Energy contribution | -22.22 |
| Covariance contribution | 0.36 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.64 |
| Structure conservation index | 0.86 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.76 |
| SVM RNA-class probability | 0.809714 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
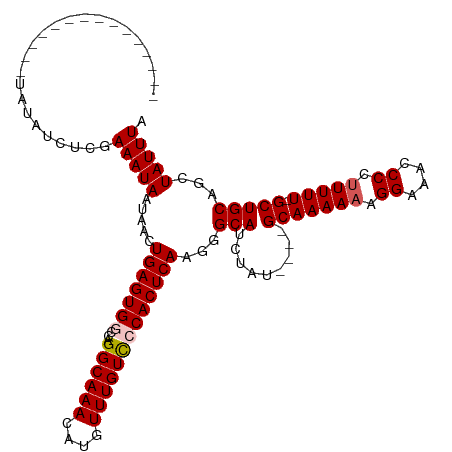
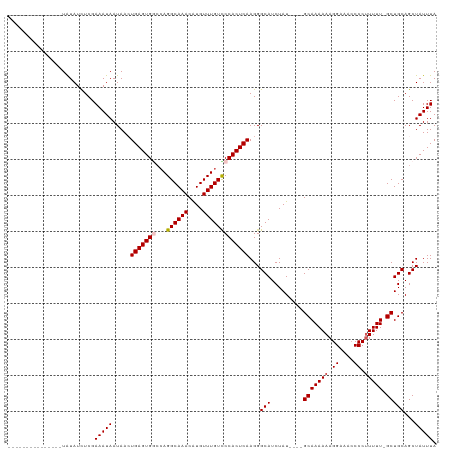
>dm3.chr3R 7539034 99 + 27905053 ---------------UAUAUCUCGAAAUAAUAACUGAGUGGCCAGGCAAACAUGUUUGUUCCACUCAAGGGCAUCUAU----GCAAAAAAGGAAACCCCUUUUU-GCUGCAGCUAUUUA ---------------...................(((((((.((((((....))))))..)))))))..(((......----(((((((.((....)).)))))-))....)))..... ( -25.40, z-score = -1.80, R) >droEre2.scaffold_4770 3668541 104 - 17746568 ---------------UGUAUUUCGAAAUAAUAACUGAGUGCCCAGGCAAACGUGUUUGUCCCACUCAAGGGCAUCUAUCUAUGCAAAAAAGGAAACCCCUUUUUUGCUGCAGCUAUUUA ---------------.............((((.(((.((((((.((((((....))))))........))))))........((((((((((.....))))))))))..))).)))).. ( -27.20, z-score = -2.14, R) >droYak2.chr3R 11949164 114 - 28832112 CUCGAAAUAAUAACGUAUAUCUCGAAAUAAUAACUGAGUGAAGAGGCAAACAUGUUUGUUCCACUCAAGGGCAUCUAU----GCAAAAAAGGAUAACCCAUUUU-GCUGCAGCUAUUUA .((((.(((........))).)))).........((((((((.(((((....))))).)).))))))..(((......----((((((..((....))..))))-))....)))..... ( -23.00, z-score = -1.04, R) >droSec1.super_0 6693877 99 - 21120651 ---------------UAUAUCUCGAAAUAAUAAGUGAGUGGCCAGGCAAACAUGUUUGUCCCACUCAAGGGCAUCUAU----GCAAAAAAGGAAACCCCUUUUU-GCUGCAGCUAUUUA ---------------...................(((((((.((((((....))))))..)))))))..(((......----(((((((.((....)).)))))-))....)))..... ( -25.80, z-score = -1.61, R) >droSim1.chr3R 13660350 99 - 27517382 ---------------UAUAUCUCGAAAUAAUAAGUGAGUGGCCAGGCAAACAUGUUUGUCCCACUCAAGGGCAUCUAU----GCAAAAAAGGAAACCCCUUUUU-GCUGCAGCUAUUUA ---------------...................(((((((.((((((....))))))..)))))))..(((......----(((((((.((....)).)))))-))....)))..... ( -25.80, z-score = -1.61, R) >consensus _______________UAUAUCUCGAAAUAAUAACUGAGUGGCCAGGCAAACAUGUUUGUCCCACUCAAGGGCAUCUAU____GCAAAAAAGGAAACCCCUUUUU_GCUGCAGCUAUUUA ........................(((((.....(((((((...((((((....)))))))))))))...(((.........((.(((((((.....))))))).)))))...))))). (-21.86 = -22.22 + 0.36)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:07:22 2011