
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 7,371,535 – 7,371,646 |
| Length | 111 |
| Max. P | 0.861728 |

| Location | 7,371,535 – 7,371,646 |
|---|---|
| Length | 111 |
| Sequences | 7 |
| Columns | 113 |
| Reading direction | reverse |
| Mean pairwise identity | 61.83 |
| Shannon entropy | 0.78470 |
| G+C content | 0.29499 |
| Mean single sequence MFE | -14.94 |
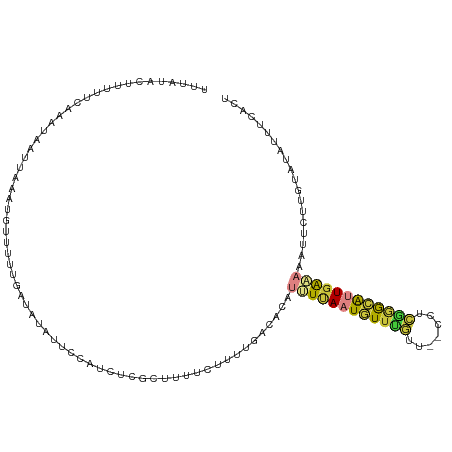
| Consensus MFE | -6.19 |
| Energy contribution | -6.30 |
| Covariance contribution | 0.11 |
| Combinations/Pair | 1.83 |
| Mean z-score | -0.99 |
| Structure conservation index | 0.41 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.96 |
| SVM RNA-class probability | 0.861728 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
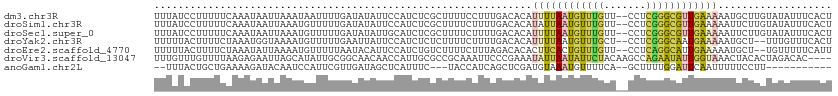
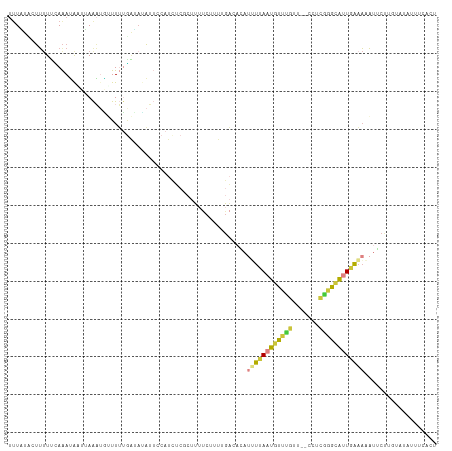
>dm3.chr3R 7371535 111 - 27905053 UUUAUCCUUUUUCAAAUAAUUAAAUAAUUUUGAUAUAUUCCAUCUCGCUUUUCCUUUGACACAUUUUAAUGUUUGUU--CCUCGGGCGUUGAAAAAUGCUUGUAUAUUUCACU ............((((((((((((....)))))).....................((((......))))))))))..--...((((((((....))))))))........... ( -11.60, z-score = -0.10, R) >droSim1.chr3R 13493274 111 + 27517382 UUUAUCCUUUUUCAAAUAAUUAAAUGUUUUUGAUAUAUUCCAUCUCGCUUUUCUUUUGACACAUAUUAAUGUUUGUU--CCUCGGGCGUUGAAAAAUUCUUGUAUAUUUCACU .......((((((((...((((((....))))))...........(((((.......((((........))))....--....)))))))))))))................. ( -11.36, z-score = -0.14, R) >droSec1.super_0 6527331 111 + 21120651 UUUAUCCUUUUUCAAAUAAUUAAAUGUUUUUGAUAUAUUGCAUCUCGCUUUUCUUUUGACACAUUUUAAUGUUUGUU--CCUCGGGCGUUGAAAAAUUCUUGUAUAUUUCACU ...........(((((..((((((....)))))).....((.....))......)))))....((((((((((((..--...))))))))))))................... ( -13.10, z-score = -0.52, R) >droYak2.chr3R 11411568 109 - 28832112 UUUUUACUUUUCUAAAUGGUAAAAUGUUUUUGAAUUAUUCCAUCUCUCUUUUCUUUUGACACAUUUUAAUGUUUGCU--CCUCGGGCAAUGAAAAAUGCU--UUUGUUUCACU .(((((((.........)))))))((((...(((................)))....))))((((((.....(((((--.....)))))...))))))..--........... ( -11.69, z-score = 0.22, R) >droEre2.scaffold_4770 3497348 109 + 17746568 UUUUUACUUUUCUAAAUAUUAAAAUGUUUUUAAUACAUUCCAUCUGUCUUUUCUUUAGACACACUUCACUGUUUGUU--CCUCAGGCAUUGAAAAAUGCU--UGUUUUUCAUU ............((((((....(((((.......))))).....(((((.......)))))........))))))..--...((((((((....))))))--))......... ( -16.20, z-score = -2.41, R) >droVir3.scaffold_13047 331270 109 + 19223366 UUUGUUUGUUUUAAGAGAAUUAGCAUAUUGCGGCAACAACCAUUGCGCCGCAAAUUCCCGAAAUAUUAAUAUUCUACAAGCCAGAAUAUUGGUAAACUACACUAGACAC---- ......(((((....((..........(((((((............)))))))..........((((((((((((.......))))))))))))..)).....))))).---- ( -22.90, z-score = -2.37, R) >anoGam1.chr2L 39972629 95 + 48795086 --UUUACUGCUGAAAAGAUACAAUCCAUUCGUUGAUAGCUCAUUUC---UACCAUCAGCUCGAUGUAAAUGUUUUCA--GCUUUUGGAUUCAAUUUUUCCUU----------- --.........((((((....((((((...(((((.....(((((.---...((((.....)))).)))))...)))--))...))))))...))))))...----------- ( -17.70, z-score = -1.60, R) >consensus UUUAUACUUUUUCAAAUAAUUAAAUGUUUUUGAUAUAUUCCAUCUCGCUUUUCUUUUGACACAUUUUAAUGUUUGUU__CCUCGGGCAUUGAAAAAUUCUUGUAUAUUUCACU ...............................................................((((((((((((.......))))))))))))................... ( -6.19 = -6.30 + 0.11)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:06:48 2011