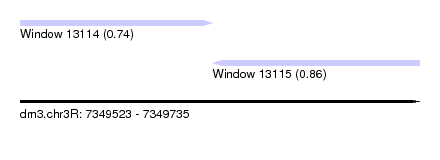
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 7,349,523 – 7,349,735 |
| Length | 212 |
| Max. P | 0.856503 |

| Location | 7,349,523 – 7,349,625 |
|---|---|
| Length | 102 |
| Sequences | 8 |
| Columns | 107 |
| Reading direction | forward |
| Mean pairwise identity | 75.18 |
| Shannon entropy | 0.49237 |
| G+C content | 0.53712 |
| Mean single sequence MFE | -32.90 |
| Consensus MFE | -16.55 |
| Energy contribution | -17.96 |
| Covariance contribution | 1.41 |
| Combinations/Pair | 1.35 |
| Mean z-score | -1.57 |
| Structure conservation index | 0.50 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.56 |
| SVM RNA-class probability | 0.742365 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 7349523 102 + 27905053 --UUUUUUUUCAUUGUUUUUUUGACUGCUCAUUUCCAUGACGCGCAGUUGGCAUUGUGUAGGGGCGUGGUCGAUG-GGGGCGGUAGAUGGGCGGCAAGAAGGCAC-- --...........((((((((((.((((((((((((....(((((.....((.....))....)))))(((....-..))))).)))))))))))))))))))).-- ( -36.90, z-score = -3.50, R) >droSim1.chr3R 13473183 103 - 27517382 -CUUUUUUUUCAUUGUUUUUUUGACUGCUCAUUUCCAUGACGCGCAGUUGGCAUUGUGUAGGGGCGUGGUCGAAG-GGGGCGGUAGAUGGGCGGCAAGGAGGCGC-- -.............(((((((((.((((((((((((....(((((.....((.....))....)))))(((....-..))))).)))))))))))))))))))..-- ( -35.90, z-score = -2.55, R) >droSec1.super_0 6505555 104 - 21120651 CUUUUUUUUUCAUUGUUUUUUUGACUGCUCAUUUCCAUGACGCGCAGUUGGCAUUGUGUAGGGGCGUGGUCGAAG-GGGGCGGUAGAUGGGCGGCAAGGAGGCGC-- ..............(((((((((.((((((((((((....(((((.....((.....))....)))))(((....-..))))).)))))))))))))))))))..-- ( -35.90, z-score = -2.50, R) >droYak2.chr3R 11385102 103 + 28832112 --CUUUUUUUCAUUGUUUUUUUGACUGCUCAUUUCCAUGACGCGCAGUUGGCAUUGUGUAGGGGCGUGGUCGAAUGGGGGCGGCAGAUGGGCGGCAAGGAGGCGC-- --............(((((((((.((((((((((((....(((((.....((.....))....)))))(((.......))))).)))))))))))))))))))..-- ( -35.40, z-score = -1.86, R) >droEre2.scaffold_4770 3475783 103 - 17746568 --CUUUUUUUCAUUGUUUUUUUGACUGCUCAUUUCCAUGACGCGCAGUUGGCAUUGUGUAGAGGCGUGGUCGAAUUGGGGCGGUAGAUGGGCGACAAGGAGGCGG-- --...........((((((((((..(((((((((((....(((((.....((.....))....)))))(((.......))))).))))))))).)))))))))).-- ( -30.90, z-score = -1.18, R) >droAna3.scaffold_13340 13793275 88 + 23697760 -------UCAUUUUUUUUUUUUGACUGCUCAUUUCCCUGACGCUCAACUGGCAUUGUAAGGAGGCGUGGCCGGGG-GAGGCGG-GGACAGUAGGCGG---------- -------.................((((..(((((((((.(.(((..(((((..(((......)))..)))))..-)))))))-))).)))..))))---------- ( -32.40, z-score = -2.19, R) >dp4.Unknown_group_353 13459 82 + 17550 --CUCUUUUUUUGUGUUUUUUUGACUGCUCAUUUCCAUGACGCGC-------AUUGUGCAGAGGCGUGCCCCAGA-GCGGUGG-GGCAGGGCA-------------- --.......((((..(.....((...((((((....)))).)).)-------)..)..))))(.(.(((((((..-....)))-)))).).).-------------- ( -23.90, z-score = 0.49, R) >droPer1.super_6 970654 96 + 6141320 --CUCUUUUUUUGUGUUUUUUUGACUGCUCAUUUCCAUGACGCGC-------AUUGUGCAGAGGCGUGCCCCAGA-GCGGUGG-GGCAGGGCGGUGGAGGGCCGGGC --........................((((.(((((((...((((-------...))))...(.(.(((((((..-....)))-)))).).).)))))))...)))) ( -31.90, z-score = 0.75, R) >consensus __CUUUUUUUCAUUGUUUUUUUGACUGCUCAUUUCCAUGACGCGCAGUUGGCAUUGUGUAGAGGCGUGGUCGAAG_GGGGCGGUAGAUGGGCGGCAAGGAGGCGC__ ..............(..((((((.((((((.(((((....(((((..................)))))(((.......))))).))).))))))))))))..).... (-16.55 = -17.96 + 1.41)



| Location | 7,349,625 – 7,349,735 |
|---|---|
| Length | 110 |
| Sequences | 11 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 86.92 |
| Shannon entropy | 0.27281 |
| G+C content | 0.48610 |
| Mean single sequence MFE | -21.85 |
| Consensus MFE | -15.02 |
| Energy contribution | -14.86 |
| Covariance contribution | -0.15 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.14 |
| Structure conservation index | 0.69 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.94 |
| SVM RNA-class probability | 0.856503 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 7349625 110 - 27905053 ----CUCGUUCAGCCGCCCCACGUGUGCAG-AAAGUUUUGCCAACAGUUUUAAUUUCCCAUAAUUACACAAUUACAAAGCGCAAAGUUUU-CAGCACACGCCACGCCUC----GCGCUCC ----........((.((....(((((((.(-(((..((((((...((((.(((((......)))))...)))).....).)))))..)))-).)))))))....))...----))..... ( -22.50, z-score = -2.14, R) >droSim1.chr3R 13473286 110 + 27517382 ----CUCGUUCAGCCGCCCCACGUGUGCAG-AAAGUUUUGCCAACAGUUUUAAUUUCCCAUAAUUACACAAUUACAAAGCGCAAAGUUUU-CAGCACACGCCACGCCCC----GCGCUCC ----........((.((....(((((((.(-(((..((((((...((((.(((((......)))))...)))).....).)))))..)))-).)))))))....))...----))..... ( -22.50, z-score = -2.40, R) >droSec1.super_0 6505659 110 + 21120651 ----CUCGUUCAGCCGCCCCACGUGUGCAG-AAAGUUUUGCCAACAGUUUUAAUUUCCCAUAAUUACACAAUUACAAAGCGCAAAGUUUU-CAGCACACGCCACGCCCC----GCGCUCC ----........((.((....(((((((.(-(((..((((((...((((.(((((......)))))...)))).....).)))))..)))-).)))))))....))...----))..... ( -22.50, z-score = -2.40, R) >droYak2.chr3R 11385205 110 - 28832112 ----CUCGUUCAGCCGCCCCACGUGUGCAG-AAAGUUUUGCCAACAGUUUUAAUUUCCCAUAAUUACACAAUUACAAAGCGCAAAGUUUU-CAGCACACGCCACGCCCC----ACGCCCC ----...........((....(((((((.(-(((..((((((...((((.(((((......)))))...)))).....).)))))..)))-).)))))))....))...----....... ( -21.00, z-score = -2.73, R) >droEre2.scaffold_4770 3475886 110 + 17746568 ----CUCGUUCAGCCGCCCCACGUGUGCAG-AAAGUUUUGCCAACAGUUUUAAUUUCCCAUAAUUACACAAUUACAAAGCGCAAAGUUUU-CAGCACACGCCACGCCCC----GCGCCCA ----........((.((....(((((((.(-(((..((((((...((((.(((((......)))))...)))).....).)))))..)))-).)))))))....))...----))..... ( -22.50, z-score = -2.49, R) >droAna3.scaffold_13340 13793363 110 - 23697760 ----CUCGC-AGGCCGCCCCACGUGUGCAGAAAAGUUUUGCCAACAGUUUUAAUUUCCCAUAAUUACACAAUUACAAAGCGAAAAGUUUU-CAGCACACGCCACGCCCU----UCGCCUG ----....(-((((.((....(((((((.(((((.((((((....((((.(((((......)))))...)))).....))))))..))))-).)))))))....))...----..))))) ( -27.00, z-score = -3.26, R) >dp4.Unknown_group_353 13541 109 - 17550 ----CGUGC--CGCCACCGUGUGUGUGCAGAAAAGUUUUGCCAACAGUUUUAAUUUCCCAUAAUUACACAAUUACAAAGCGCAAAGUUUU-CAGCACACGCCACGCCCC----CCGCCCG ----((.((--.(....(((((((((((.(((((.(((((((...((((.(((((......)))))...)))).....).))))))))))-).))))))).))))....----).)).)) ( -24.80, z-score = -2.34, R) >droWil1.scaffold_181089 8238208 108 + 12369635 -----------CCUGUUCCCACGUGUGCAGAAAAGUUUUGCCAACAGUUUUAAUUUCCCAUAAUUACACAAUUACAAAGCGCAAAGUUUU-CAGCACACGCCACGCCUUCUUACAGCCCC -----------.((((.....(((((((.(((((.(((((((...((((.(((((......)))))...)))).....).))))))))))-).)))))))............)))).... ( -18.83, z-score = -2.19, R) >droVir3.scaffold_12855 9846404 114 - 10161210 -UCGUUGCGCCCACGGACCCACGUGUGCAG-AAAGUUUUGCCAACAGUUUUAAUUUCCCAUAAUUACACAAUUACAAAGCGUAAAGUUUUCCAGCACAUGCCACGCCUC----UAGCCCC -.......((....((.....(((((((.(-(((..((((((...((((.(((((......)))))...)))).....).)))))..))))..))))))).....))..----..))... ( -18.10, z-score = -0.05, R) >droMoj3.scaffold_6540 24846483 112 - 34148556 ---UUUGCGCCCAUGGACCCACGUGUGCAG-AAAGUUUCGCCAACAGUUUUAAUUUCCCAUAAUUACACAAUUACAAAGCGUAAAGUUUUCCAGCACACGCCACGCCUC----CAGCCCC ---.....((....((.....(((((((.(-((((((((((....((((.(((((......)))))...)))).....))).))).)))))..))))))).....))..----..))... ( -20.50, z-score = -1.70, R) >droGri2.scaffold_14830 2187359 115 - 6267026 UUCGUUUCGUCCACGGACCCACGUGUGCAG-AAAGUUUUGCCAACAGUUUUAAUUUCCCAUAAUUACACAAUUACAAAGCGUAAAGUUUUCCAGCACACGCCACGCCUC----CAGCCCC ..(((...(((....)))...(((((((.(-(((..((((((...((((.(((((......)))))...)))).....).)))))..))))..)))))))..)))....----....... ( -20.10, z-score = -1.88, R) >consensus ____CUCGUUCAGCCGCCCCACGUGUGCAG_AAAGUUUUGCCAACAGUUUUAAUUUCCCAUAAUUACACAAUUACAAAGCGCAAAGUUUU_CAGCACACGCCACGCCCC____CCGCCCC .....................(((((((...(((..((((((...((((.(((((......)))))...)))).....).)))))..)))...))))))).................... (-15.02 = -14.86 + -0.15)
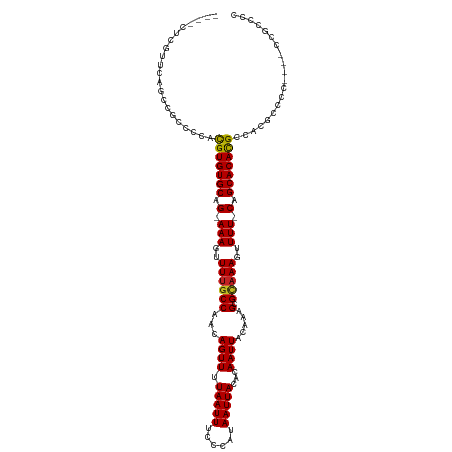



Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:06:45 2011