
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 7,081,571 – 7,081,674 |
| Length | 103 |
| Max. P | 0.598912 |

| Location | 7,081,571 – 7,081,674 |
|---|---|
| Length | 103 |
| Sequences | 6 |
| Columns | 115 |
| Reading direction | reverse |
| Mean pairwise identity | 68.00 |
| Shannon entropy | 0.59744 |
| G+C content | 0.43478 |
| Mean single sequence MFE | -25.41 |
| Consensus MFE | -8.66 |
| Energy contribution | -10.17 |
| Covariance contribution | 1.50 |
| Combinations/Pair | 1.23 |
| Mean z-score | -1.73 |
| Structure conservation index | 0.34 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.22 |
| SVM RNA-class probability | 0.598912 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 7081571 103 - 23011544 --------UUUCCCCAAAGCUGACGACAAUCAUUUUUAGAUUCUGCGAUUAAUCGCUUAUCAGCACAAUGC----UGUCGCUUUUUAUUCCCCGUCGCAAAGUCUGGUUUGGAUU --------.....(((((.(.((((((...........(((...((((....))))..)))(((.....))----))))((...............))...))).).)))))... ( -23.56, z-score = -1.68, R) >droWil1.scaffold_180772 3119289 102 + 8906247 CUCAAUUGUGACUCGAAAAAUGUUUUGAAUUCUCUAU---UUCUACUGUCAAUUACUUCUCA-UGUAAUGCGAAUUAACACCCUUAAUUAAAUUUCAUUUUAUAUG--------- .......(((..((((((....)))))).........---........((.(((((......-.)))))..)).....))).........................--------- ( -7.20, z-score = 0.51, R) >droEre2.scaffold_4929 15995998 104 - 26641161 --------UUUCCCCAA--CCGGCGACAAUCACUUUUAGAUUCUGCGAUUAAUCGCUUAUCAGCCCAAGGUUCAGCCUGGGUUUUGAUUCCCCGUCGCAGAGUCUGCCUUGAUU- --------..((.((..--..)).)).(((((....(((((((((((((.........(((((((((.((.....)))))))..)))).....)))))))))))))...)))))- ( -32.84, z-score = -3.44, R) >droYak2.chr2L 16497284 103 + 22324452 --------UUUCCCCAAAGCGGGCGACAAUCACUUUUAGAUUCUGCGUUGAAUCGCUUAUCAGUCCAAGGC----CUGCGUUCCUUAUUCCCCGUCGCAGAGUCUGGACGGGAUU --------..((((..((((.(....)...........(((((......)))))))))....(((((.(((----(((((...............))))).)))))))))))).. ( -33.66, z-score = -2.12, R) >droSec1.super_3 2605607 102 - 7220098 --------UUUCCCCAAAGCUGGCGACAAUCAUUUUUAGAUUCUGCGAUUAAUCGCUUAUCAGCCCAAGGC----CGUCGC-UUUUAUUCCUCGACGCAGAGUCUGUGUUGGAUU --------.............((((((...........(((...((((....))))..))).(((...)))----.)))))-)........(((((((((...)))))))))... ( -27.60, z-score = -1.76, R) >droSim1.chr2L 6868918 102 - 22036055 --------UUUCCCCAAAGCUGGCGACAAUCAUUUUUAGAUUCUGCGAUUAAUCGCUUAUCAGCCCAAGGC----CGUCGC-UUUUAUUCAUCGUCGCAGAGUCUGUGUUGGAUU --------.....((((.((((....)).........((((((((((((.....(((....)))..(((((----....))-)))........)))))))))))))).))))... ( -27.60, z-score = -1.91, R) >consensus ________UUUCCCCAAAGCUGGCGACAAUCAUUUUUAGAUUCUGCGAUUAAUCGCUUAUCAGCCCAAGGC____CGUCGCUUUUUAUUCCCCGUCGCAGAGUCUGGGUUGGAUU ....................................(((((((((((((............................................)))))))))))))......... ( -8.66 = -10.17 + 1.50)

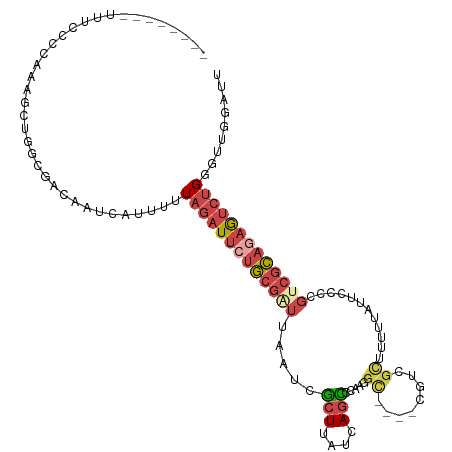

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:22:50 2011