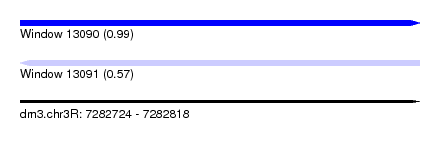

| Sequence ID | dm3.chr3R |
|---|---|
| Location | 7,282,724 – 7,282,818 |
| Length | 94 |
| Max. P | 0.992038 |

| Location | 7,282,724 – 7,282,818 |
|---|---|
| Length | 94 |
| Sequences | 4 |
| Columns | 94 |
| Reading direction | forward |
| Mean pairwise identity | 79.04 |
| Shannon entropy | 0.34212 |
| G+C content | 0.31234 |
| Mean single sequence MFE | -16.30 |
| Consensus MFE | -9.01 |
| Energy contribution | -10.57 |
| Covariance contribution | 1.56 |
| Combinations/Pair | 1.15 |
| Mean z-score | -3.23 |
| Structure conservation index | 0.55 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.51 |
| SVM RNA-class probability | 0.992038 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
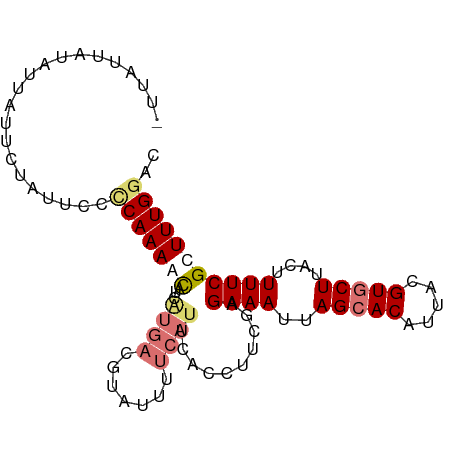
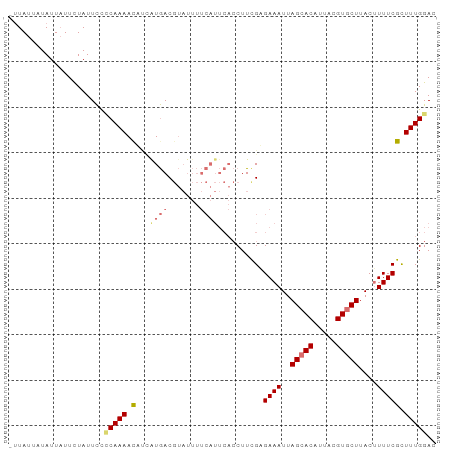
>dm3.chr3R 7282724 94 + 27905053 AUUAUUCUAUUAUUUUAUUCACCAAAACAUAGUGACGUAUUUUCAUUCACCUUCGUGAAAUUAGAACAUUACGUGCUUACAUUUCGUUUUGGAC .....................(((((((..(((((((((..(((.(((((....)))))....)))...))))).....))))..))))))).. ( -17.30, z-score = -2.69, R) >droEre2.scaffold_4770 3408799 93 - 17746568 -UUAUAAUAUUUUUCUAUUCUUCAAAAUUGCAUGACGUAUUUUCAUUCACCUUCGAGAAAUUAGCACAUUACGUGCUUACUUUUCGCUUUGGAC -...................((((((..((.((((.......)))).))....((((((...(((((.....)))))...)))))).)))))). ( -14.80, z-score = -2.14, R) >droYak2.chr3R 11317127 78 + 28832112 --UAUUAUAUUAUUCUAUUUCACAAAACUCCAUAAC--------------CUUCGAGAAAUUAGCACAUUACGUGCUUACUUUUCGCUUUGGAC --..........................((((....--------------...((((((...(((((.....)))))...))))))...)))). ( -14.00, z-score = -3.95, R) >droSim1.chr3R_random 448175 94 + 1307089 AUUAUUCUAUUAUUUUAUUCCCCAAAACAUUGUGACGUAUUUUCAUUCACCUUUGCGAAAUUAGCACAUUACGUGCUUACUUUUCGUUUUGGAC .....................(((((((...((((...........))))......((((..(((((.....)))))....))))))))))).. ( -19.10, z-score = -4.15, R) >consensus _UUAUUAUAUUAUUCUAUUCCCCAAAACAUCAUGACGUAUUUUCAUUCACCUUCGAGAAAUUAGCACAUUACGUGCUUACUUUUCGCUUUGGAC .....................(((((((...((((.......))))..........((((..(((((.....)))))....))))))))))).. ( -9.01 = -10.57 + 1.56)

| Location | 7,282,724 – 7,282,818 |
|---|---|
| Length | 94 |
| Sequences | 4 |
| Columns | 94 |
| Reading direction | reverse |
| Mean pairwise identity | 79.04 |
| Shannon entropy | 0.34212 |
| G+C content | 0.31234 |
| Mean single sequence MFE | -17.73 |
| Consensus MFE | -8.79 |
| Energy contribution | -11.10 |
| Covariance contribution | 2.31 |
| Combinations/Pair | 1.17 |
| Mean z-score | -2.13 |
| Structure conservation index | 0.50 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.15 |
| SVM RNA-class probability | 0.565533 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
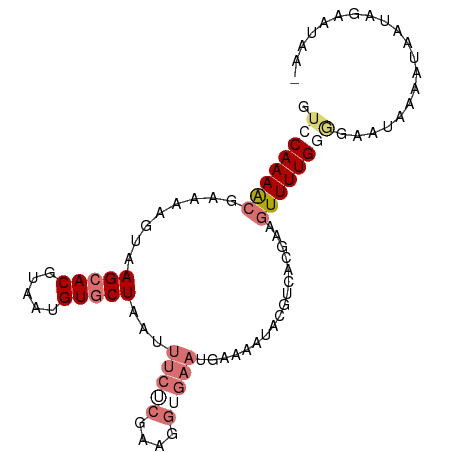
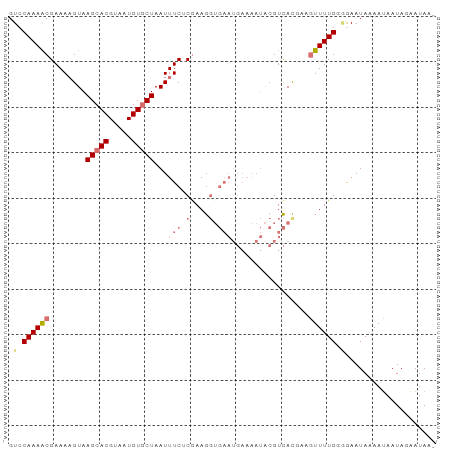
>dm3.chr3R 7282724 94 - 27905053 GUCCAAAACGAAAUGUAAGCACGUAAUGUUCUAAUUUCACGAAGGUGAAUGAAAAUACGUCACUAUGUUUUGGUGAAUAAAAUAAUAGAAUAAU ...((...(((((((((...(((((...(((....(((((....))))).)))..)))))...))))))))).))................... ( -19.50, z-score = -2.43, R) >droEre2.scaffold_4770 3408799 93 + 17746568 GUCCAAAGCGAAAAGUAAGCACGUAAUGUGCUAAUUUCUCGAAGGUGAAUGAAAAUACGUCAUGCAAUUUUGAAGAAUAGAAAAAUAUUAUAA- .......((.........))..(((((((.(((.((..((((((.((.((((.......)))).)).))))))..)))))....)))))))..- ( -14.60, z-score = -0.70, R) >droYak2.chr3R 11317127 78 - 28832112 GUCCAAAGCGAAAAGUAAGCACGUAAUGUGCUAAUUUCUCGAAG--------------GUUAUGGAGUUUUGUGAAAUAGAAUAAUAUAAUA-- ........(((.((((.(((((.....))))).)))).)))...--------------((((((..(((((((...)))))))..)))))).-- ( -16.10, z-score = -2.22, R) >droSim1.chr3R_random 448175 94 - 1307089 GUCCAAAACGAAAAGUAAGCACGUAAUGUGCUAAUUUCGCAAAGGUGAAUGAAAAUACGUCACAAUGUUUUGGGGAAUAAAAUAAUAGAAUAAU .(((((((((..((((.(((((.....))))).))))..)....((((.((......))))))...)))))))).................... ( -20.70, z-score = -3.16, R) >consensus GUCCAAAACGAAAAGUAAGCACGUAAUGUGCUAAUUUCUCGAAGGUGAAUGAAAAUACGUCACGAAGUUUUGGGGAAUAAAAUAAUAGAAUAA_ .((((((((........(((((.....)))))...(((((....))))).................)))))))).................... ( -8.79 = -11.10 + 2.31)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:06:26 2011