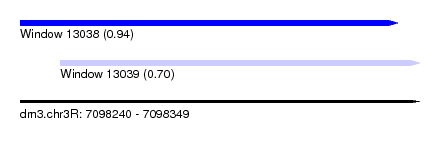

| Sequence ID | dm3.chr3R |
|---|---|
| Location | 7,098,240 – 7,098,349 |
| Length | 109 |
| Max. P | 0.939628 |

| Location | 7,098,240 – 7,098,343 |
|---|---|
| Length | 103 |
| Sequences | 5 |
| Columns | 118 |
| Reading direction | forward |
| Mean pairwise identity | 69.38 |
| Shannon entropy | 0.53646 |
| G+C content | 0.50139 |
| Mean single sequence MFE | -35.82 |
| Consensus MFE | -22.46 |
| Energy contribution | -23.10 |
| Covariance contribution | 0.64 |
| Combinations/Pair | 1.45 |
| Mean z-score | -1.31 |
| Structure conservation index | 0.63 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.43 |
| SVM RNA-class probability | 0.939628 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
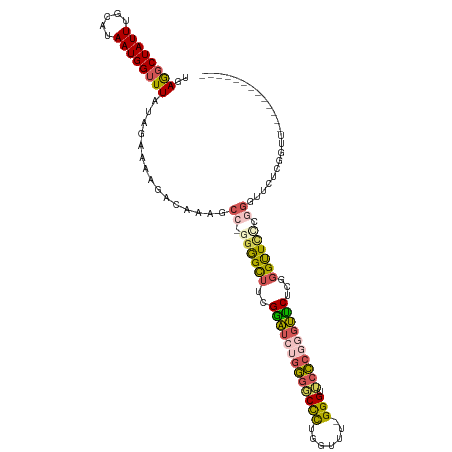
>dm3.chr3R 7098240 103 + 27905053 UGAGGCUAUUUGCAUAAUGGUUUAUAGAAAAGACAAAGCC-GGGGCUUCGGAUCUGGGUCCUUGGUUU-GGGUUCCCGGGUUCUCGGGUUCCUGGGUUCUCGGUU------------- ..((((((((.....)))))))).............((((-.(((((..((((....))))..)))))-.)))).(((((..(((((....)))))..)))))..------------- ( -34.50, z-score = -0.71, R) >droSim1.chr3R 13252340 111 - 27517382 UGAGGCUAUUUGCAUAAUGGUUUAUAGAAAAGACAAAGCC-GGGGCUUCGGAUCUGGGGCUCUGGUUU-GGGUUCCCGGGUUCUCCGGUUCUCGGGUUCCUGGGUUCUCGGUU----- ..((((((((.....))))))))........(((.(((((-(((((((((....))))))))))))))-(((..((((((..(.(((.....))))..))))))..))).)))----- ( -44.70, z-score = -2.56, R) >droSec1.super_0 6264235 103 - 21120651 UGAGGCUAUUUGCAUAAUGGUUUAUAGAAAAGACAAAGCC-GGGGCUUCGGAUCUGGGGCUCUGUUUU-GGGUUCCCGGGUUCUCGGGUUCCUGGGUUCUCGGUU------------- ..((((((((.....))))))))...(((....((((((.-(((((((((....))))))))))))))-)..)))(((((..(((((....)))))..)))))..------------- ( -37.10, z-score = -1.69, R) >droYak2.chr3R 11133124 89 + 28832112 UGAGGCUAUUUGCAUAAUGGUUUAUAGAAAAGACAAAGCC-GGGGCUUGGGAUCUGGGGCCCAGGUUUUGGGUUGCCUAGUCCUCGGGUU---------------------------- .(((((((((.....)))))....(((..((..(((((((-.((((((.(....).)))))).)))))))..))..)))..)))).....---------------------------- ( -29.60, z-score = -0.67, R) >droWil1.scaffold_181089 7982202 118 - 12369635 CCAACCUAUUUACAUAAUGGUUUAUAGAAAAGACAAAGCCCAGAGUGUGGUCACAGUGUCCCUUGGUCCGGGUACAAAAGGACAAUGGCUUCCUGCUUGGGCUACUUUCUUUUUUGUG ..((((.(((.....)))))))(((((((((((...((((((.((((.(((((...(((((.(((.........)))..))))).)))))...))))))))))....))))))))))) ( -33.20, z-score = -0.90, R) >consensus UGAGGCUAUUUGCAUAAUGGUUUAUAGAAAAGACAAAGCC_GGGGCUUCGGAUCUGGGGCCCUGGUUU_GGGUUCCCGGGUUCUCGGGUUCCCGGGUUCUCGGUU_____________ ..((((((((.....))))))))...............((.((((((..(((((((((((((.......))).))))))))))...)))))).))....................... (-22.46 = -23.10 + 0.64)



| Location | 7,098,251 – 7,098,349 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 113 |
| Reading direction | forward |
| Mean pairwise identity | 67.95 |
| Shannon entropy | 0.55903 |
| G+C content | 0.52190 |
| Mean single sequence MFE | -35.12 |
| Consensus MFE | -17.54 |
| Energy contribution | -17.10 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.40 |
| Mean z-score | -1.18 |
| Structure conservation index | 0.50 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.44 |
| SVM RNA-class probability | 0.696043 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 7098251 98 + 27905053 GCAUAAUGGUUUAUAGAAAAGACAAAGCC-GGGGCUUCGGAUCUGGGUCCUUGGUUU-GGGUUCCCGGGUUCU--------CGGGUUCCUGGGUUCUCGGUUGCUAGA----- ........((((.......)))).(((((-((((.((((....)))).)))))))))-.(((..(((((..((--------(((....)))))..)))))..)))...----- ( -32.70, z-score = -0.40, R) >droSim1.chr3R 13252351 106 - 27517382 GCAUAAUGGUUUAUAGAAAAGACAAAGCC-GGGGCUUCGGAUCUGGGGCUCUGGUUU-GGGUUCCCGGGUUCUCCGGUUCUCGGGUUCCUGGGUUCUCGGUUGCUGGA----- .............(((....(((.(((((-(((((((((....))))))))))))))-(((..((((((..(.(((.....))))..))))))..))).))).)))..----- ( -42.30, z-score = -1.91, R) >droSec1.super_0 6264246 98 - 21120651 GCAUAAUGGUUUAUAGAAAAGACAAAGCC-GGGGCUUCGGAUCUGGGGCUCUGUUUU-GGGUUCCCGGGUUCU--------CGGGUUCCUGGGUUCUCGGUUGCUGGA----- ......................((((((.-(((((((((....))))))))))))))-)(((..(((((..((--------(((....)))))..)))))..)))...----- ( -35.90, z-score = -1.37, R) >droYak2.chr3R 11133135 83 + 28832112 GCAUAAUGGUUUAUAGAAAAGACAAAGCC-GGGGCUUGGGAUCUGGGGCCCAGGUUUUGGGUUGCCUAGUCCU--------CGGGUUCUGGA--------------------- ........((((.......))))....((-(((((((((((.(((((((((((...))))))..)))))))).--------)))))))))).--------------------- ( -31.40, z-score = -1.52, R) >droWil1.scaffold_181089 7982213 113 - 12369635 ACAUAAUGGUUUAUAGAAAAGACAAAGCCCAGAGUGUGGUCACAGUGUCCCUUGGUCCGGGUACAAAAGGACAAUGGCUUCCUGCUUGGGCUACUUUCUUUUUUGUGGCUCUU .......(..((((((((((((...((((((.((((.(((((...(((((.(((.........)))..))))).)))))...))))))))))....))))))))))))..).. ( -33.30, z-score = -0.70, R) >consensus GCAUAAUGGUUUAUAGAAAAGACAAAGCC_GGGGCUUCGGAUCUGGGGCCCUGGUUU_GGGUUCCCGGGUUCU________CGGGUUCCUGGGUUCUCGGUUGCUGGA_____ ((.....(((((............)))))....))...(((((((((((((.......))).))))))))))......................................... (-17.54 = -17.10 + -0.44)

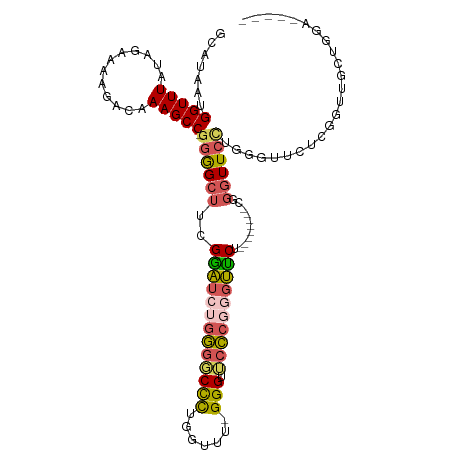

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:05:44 2011