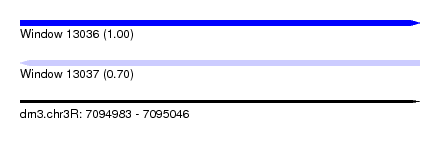

| Sequence ID | dm3.chr3R |
|---|---|
| Location | 7,094,983 – 7,095,046 |
| Length | 63 |
| Max. P | 0.998875 |

| Location | 7,094,983 – 7,095,046 |
|---|---|
| Length | 63 |
| Sequences | 6 |
| Columns | 76 |
| Reading direction | forward |
| Mean pairwise identity | 70.20 |
| Shannon entropy | 0.51851 |
| G+C content | 0.35906 |
| Mean single sequence MFE | -15.23 |
| Consensus MFE | -10.97 |
| Energy contribution | -11.12 |
| Covariance contribution | 0.15 |
| Combinations/Pair | 1.47 |
| Mean z-score | -2.59 |
| Structure conservation index | 0.72 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 3.53 |
| SVM RNA-class probability | 0.998875 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
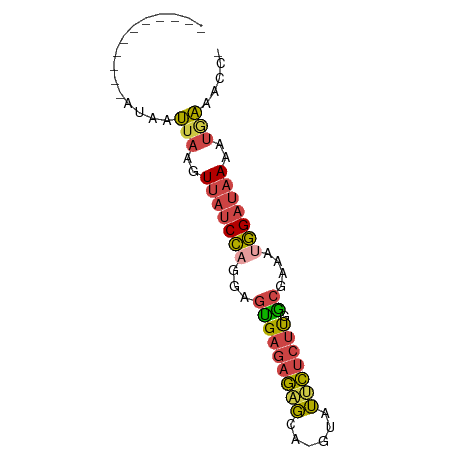
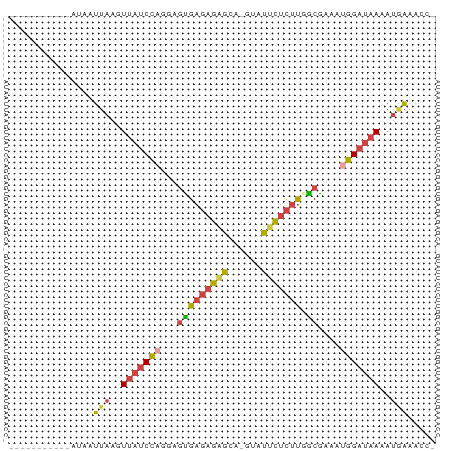
>dm3.chr3R 7094983 63 + 27905053 -----------AUAAUUAAGUUAUCCAGGAGUGAGAGAGCA-GUAUUCUCUUCGCGAAAUGGAUAAAAUGAUACC- -----------...((((..(((((((...(((((((((..-...))))).))))....)))))))..))))...- ( -18.90, z-score = -4.13, R) >droEre2.scaffold_4770 3221962 63 - 17746568 -----------AUAAUUAAGUUAUCCAGAAGCGAGAGCGCA-CUGUUCUCUUGGCGAAGAGGAUAAAAUGAAAAC- -----------....(((..((((((....((((((((...-..))))))...)).....))))))..)))....- ( -14.30, z-score = -1.78, R) >droYak2.chr3R 11129505 63 + 28832112 -----------AUAAUUAAGUUAUCCAGAAGUGAGAGAGCA-CUGUUCUCUUGGCGAAAUGGAUAAAAUAAAACC- -----------....(((..(((((((...((((((((((.-..)))))))).))....)))))))..)))....- ( -17.30, z-score = -3.95, R) >droSec1.super_0 6261037 63 - 21120651 -----------AUAAUUAAGUUAUCCAGGAGUGAGAGAGCA-GUAUUCUCUUUGCGAAAUGGAUAAAAUGAAACC- -----------....(((..(((((((...(..((((((..-...))))).)..)....)))))))..)))....- ( -15.50, z-score = -3.16, R) >droSim1.chr3R 13248104 63 - 27517382 -----------AUAAUUAAGUUAUCCAGGAGUGAGAGAGCA-GUAUUCUCUUGGCGAUAUGGAUAAAAUGAAACC- -----------....(((..(((((((...(((((((((..-...))))))).))....)))))))..)))....- ( -15.10, z-score = -2.53, R) >triCas2.ChLG6 11705509 76 - 13544221 CUAAUAGAAUUACAAAUGAGUGUCCGAUUAAUGAUAUGUUACGUAACGAUCCGACAUAUCUGACAAGAAUUAAUCG ........................((((((((((((((((............))))))))........)))))))) ( -10.30, z-score = 0.01, R) >consensus ___________AUAAUUAAGUUAUCCAGGAGUGAGAGAGCA_GUAUUCUCUUGGCGAAAUGGAUAAAAUGAAACC_ ...............(((..(((((((...(((((((((......))))))).))....)))))))..)))..... (-10.97 = -11.12 + 0.15)


| Location | 7,094,983 – 7,095,046 |
|---|---|
| Length | 63 |
| Sequences | 6 |
| Columns | 76 |
| Reading direction | reverse |
| Mean pairwise identity | 70.20 |
| Shannon entropy | 0.51851 |
| G+C content | 0.35906 |
| Mean single sequence MFE | -10.66 |
| Consensus MFE | -5.47 |
| Energy contribution | -6.19 |
| Covariance contribution | 0.72 |
| Combinations/Pair | 1.33 |
| Mean z-score | -1.34 |
| Structure conservation index | 0.51 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.45 |
| SVM RNA-class probability | 0.699092 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
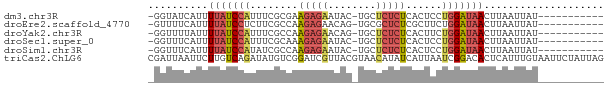
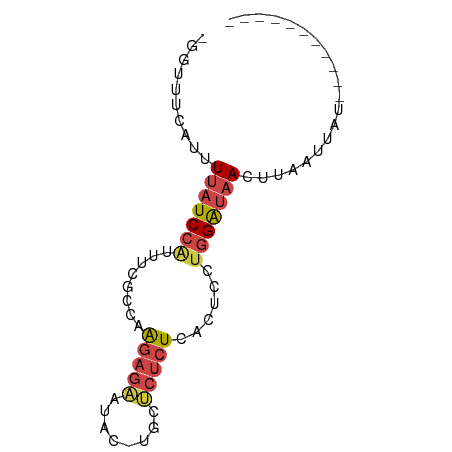
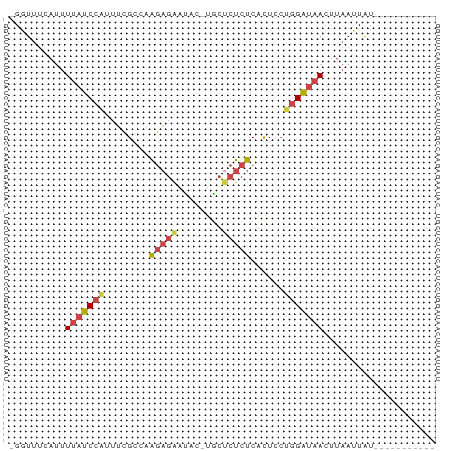
>dm3.chr3R 7094983 63 - 27905053 -GGUAUCAUUUUAUCCAUUUCGCGAAGAGAAUAC-UGCUCUCUCACUCCUGGAUAACUUAAUUAU----------- -.........(((((((....(.((.((((....-...)))))).)...))))))).........----------- ( -10.50, z-score = -1.43, R) >droEre2.scaffold_4770 3221962 63 + 17746568 -GUUUUCAUUUUAUCCUCUUCGCCAAGAGAACAG-UGCGCUCUCGCUUCUGGAUAACUUAAUUAU----------- -(((.((........(((((....)))))..(((-.(((....)))..))))).)))........----------- ( -11.20, z-score = -1.34, R) >droYak2.chr3R 11129505 63 - 28832112 -GGUUUUAUUUUAUCCAUUUCGCCAAGAGAACAG-UGCUCUCUCACUUCUGGAUAACUUAAUUAU----------- -....(((..(((((((((((.....))))..((-((......))))..)))))))..)))....----------- ( -10.70, z-score = -1.65, R) >droSec1.super_0 6261037 63 + 21120651 -GGUUUCAUUUUAUCCAUUUCGCAAAGAGAAUAC-UGCUCUCUCACUCCUGGAUAACUUAAUUAU----------- -.........(((((((........(((((....-...)))))......))))))).........----------- ( -9.64, z-score = -1.65, R) >droSim1.chr3R 13248104 63 + 27517382 -GGUUUCAUUUUAUCCAUAUCGCCAAGAGAAUAC-UGCUCUCUCACUCCUGGAUAACUUAAUUAU----------- -.........(((((((........(((((....-...)))))......))))))).........----------- ( -9.64, z-score = -1.47, R) >triCas2.ChLG6 11705509 76 + 13544221 CGAUUAAUUCUUGUCAGAUAUGUCGGAUCGUUACGUAACAUAUCAUUAAUCGGACACUCAUUUGUAAUUCUAUUAG ((((((((........(((((((((........))..)))))))))))))))........................ ( -12.30, z-score = -0.51, R) >consensus _GGUUUCAUUUUAUCCAUUUCGCCAAGAGAAUAC_UGCUCUCUCACUCCUGGAUAACUUAAUUAU___________ ..........(((((((........(((((........)))))......))))))).................... ( -5.47 = -6.19 + 0.72)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:05:42 2011