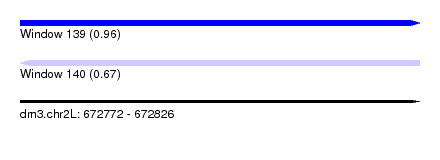

| Sequence ID | dm3.chr2L |
|---|---|
| Location | 672,772 – 672,826 |
| Length | 54 |
| Max. P | 0.960341 |

| Location | 672,772 – 672,826 |
|---|---|
| Length | 54 |
| Sequences | 5 |
| Columns | 56 |
| Reading direction | forward |
| Mean pairwise identity | 75.05 |
| Shannon entropy | 0.44415 |
| G+C content | 0.45993 |
| Mean single sequence MFE | -15.18 |
| Consensus MFE | -12.04 |
| Energy contribution | -10.60 |
| Covariance contribution | -1.44 |
| Combinations/Pair | 1.80 |
| Mean z-score | -1.50 |
| Structure conservation index | 0.79 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.68 |
| SVM RNA-class probability | 0.960341 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
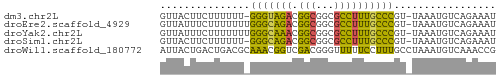
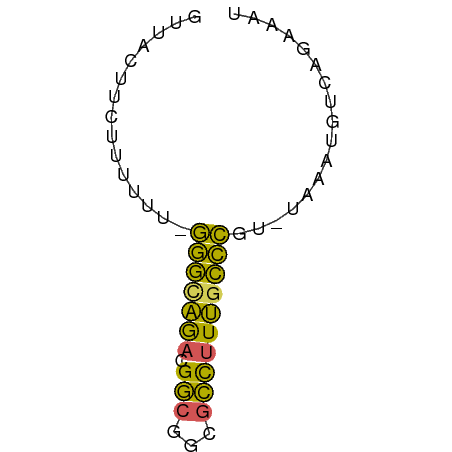
>dm3.chr2L 672772 54 + 23011544 GUUACUUCUUUUUU-GGGUAGACGGCGGCGCCUUUGCCCGU-UAAAUGUCAGAAAU ........(((..(-(((((((.(((...))))))))))).-.))).......... ( -13.90, z-score = -0.97, R) >droEre2.scaffold_4929 735038 55 + 26641161 GUUAUUUCUUUUUUUGGGCAGACGGCGGCGCCUUUGCCCGU-UAAAUGUCAGAAAU ...(((((((((..((((((((.(((...))))))))))).-.)))....)))))) ( -18.20, z-score = -2.40, R) >droYak2.chr2L 663773 55 + 22324452 GUUAUUUCUUUUUUUGGGCAAACGGCGGCGCCUUUGCCCGU-UAAAUGUCAGAAAU ...(((((((((..((((((((.(((...))))))))))).-.)))....)))))) ( -18.10, z-score = -2.54, R) >droSim1.chr2L 683483 54 + 22036055 GUUACUUCUUUUUU-GGGCAGACGGCGGCGCCUUUGCCCGU-UAAAUGUCAGAAAU ........(((..(-(((((((.(((...))))))))))).-.))).......... ( -15.90, z-score = -1.43, R) >droWil1.scaffold_180772 6347701 56 - 8906247 AUUACUGACUGACGCAAACGGUCGACGGGUUUUUCCUUUGCCUAAAUGUCAAACCG ......(((((.......)))))(((((((.........))))....)))...... ( -9.80, z-score = -0.14, R) >consensus GUUACUUCUUUUUU_GGGCAGACGGCGGCGCCUUUGCCCGU_UAAAUGUCAGAAAU ...............(((((((.(((...))))))))))................. (-12.04 = -10.60 + -1.44)

| Location | 672,772 – 672,826 |
|---|---|
| Length | 54 |
| Sequences | 5 |
| Columns | 56 |
| Reading direction | reverse |
| Mean pairwise identity | 75.05 |
| Shannon entropy | 0.44415 |
| G+C content | 0.45993 |
| Mean single sequence MFE | -12.58 |
| Consensus MFE | -8.38 |
| Energy contribution | -7.58 |
| Covariance contribution | -0.80 |
| Combinations/Pair | 1.62 |
| Mean z-score | -1.10 |
| Structure conservation index | 0.67 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.39 |
| SVM RNA-class probability | 0.671497 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
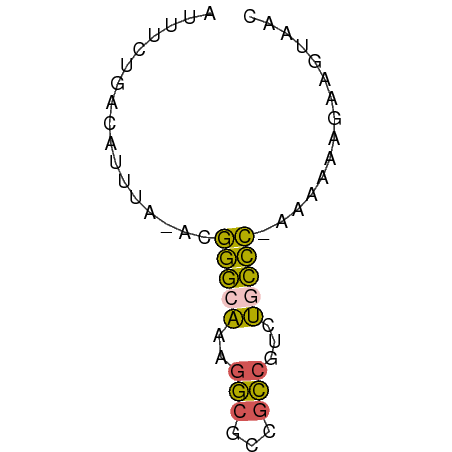
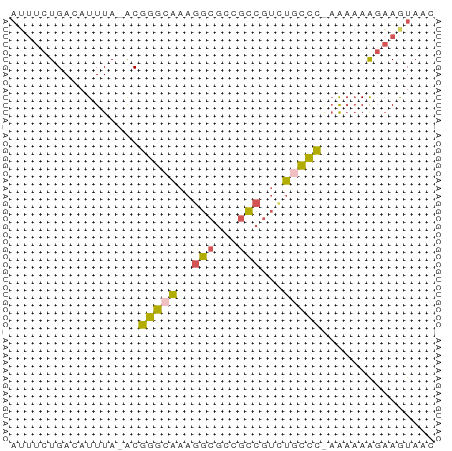
>dm3.chr2L 672772 54 - 23011544 AUUUCUGACAUUUA-ACGGGCAAAGGCGCCGCCGUCUACCC-AAAAAAGAAGUAAC (((((((.......-.)(((...(((((....))))).)))-.....))))))... ( -9.50, z-score = -0.21, R) >droEre2.scaffold_4929 735038 55 - 26641161 AUUUCUGACAUUUA-ACGGGCAAAGGCGCCGCCGUCUGCCCAAAAAAAGAAAUAAC ((((((........-..(((((..(((...)))...)))))......))))))... ( -14.19, z-score = -1.85, R) >droYak2.chr2L 663773 55 - 22324452 AUUUCUGACAUUUA-ACGGGCAAAGGCGCCGCCGUUUGCCCAAAAAAAGAAAUAAC ((((((........-..((((((((((...))).)))))))......))))))... ( -16.19, z-score = -2.52, R) >droSim1.chr2L 683483 54 - 22036055 AUUUCUGACAUUUA-ACGGGCAAAGGCGCCGCCGUCUGCCC-AAAAAAGAAGUAAC (((((((.......-.)(((((..(((...)))...)))))-.....))))))... ( -13.90, z-score = -1.40, R) >droWil1.scaffold_180772 6347701 56 + 8906247 CGGUUUGACAUUUAGGCAAAGGAAAAACCCGUCGACCGUUUGCGUCAGUCAGUAAU (((((.(((.(((....)))((......)))))))))).((((........)))). ( -9.10, z-score = 0.46, R) >consensus AUUUCUGACAUUUA_ACGGGCAAAGGCGCCGCCGUCUGCCC_AAAAAAGAAGUAAC .................(((((..(((...)))...)))))............... ( -8.38 = -7.58 + -0.80)


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:06:35 2011