
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 6,842,997 – 6,843,066 |
| Length | 69 |
| Max. P | 0.892616 |

| Location | 6,842,997 – 6,843,066 |
|---|---|
| Length | 69 |
| Sequences | 11 |
| Columns | 82 |
| Reading direction | reverse |
| Mean pairwise identity | 86.91 |
| Shannon entropy | 0.24631 |
| G+C content | 0.33871 |
| Mean single sequence MFE | -17.62 |
| Consensus MFE | -11.82 |
| Energy contribution | -12.29 |
| Covariance contribution | 0.47 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.43 |
| Structure conservation index | 0.67 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.11 |
| SVM RNA-class probability | 0.892616 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
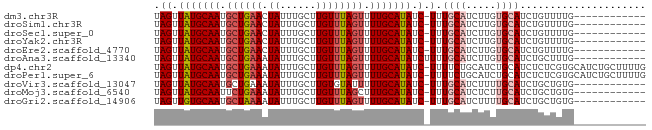
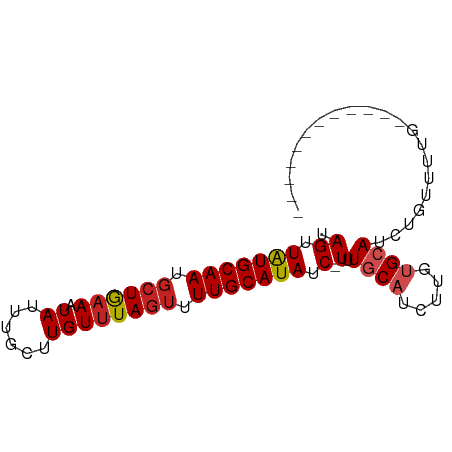
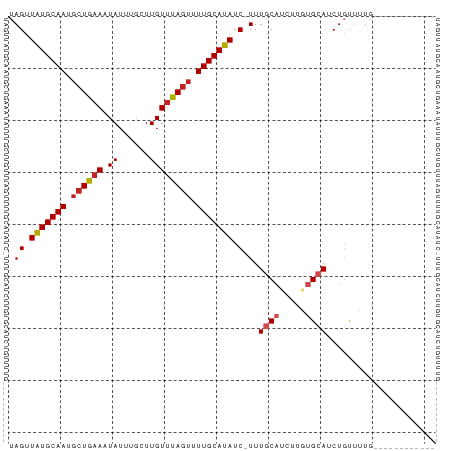
>dm3.chr3R 6842997 69 - 27905053 UAGUUAUGCAAUGCUGAACUAUUUGCUUGUUUAGUUUUGCAUAUC-UUUGCAUCUUGUGCAUCUGUUUUG------------ .((.(((((((.(((((((.........))))))).))))))).)-).(((((...))))).........------------ ( -18.80, z-score = -3.25, R) >droSim1.chr3R 12997981 69 + 27517382 UAGUUAUGCAAUGCUGAACUAUUUGCUUGUUUAGUUUUGCAUAUC-UUUGCAUCUUGUGCAUCUGUUUUG------------ .((.(((((((.(((((((.........))))))).))))))).)-).(((((...))))).........------------ ( -18.80, z-score = -3.25, R) >droSec1.super_0 6005534 69 + 21120651 UAGUUAUGCAAUGCUGAACUAUUUGCUUGUUUAGUUUUGCAUAUC-UUUGCAUCUUGUGCAUCUGUUUUG------------ .((.(((((((.(((((((.........))))))).))))))).)-).(((((...))))).........------------ ( -18.80, z-score = -3.25, R) >droYak2.chr3R 10877536 69 - 28832112 UAGUUAUGCAAUGCUGAACUAUUUGCUUGUUUAGUUUUGCAUAUC-UUUGCAUCUUGUGCAUCUGUUUUG------------ .((.(((((((.(((((((.........))))))).))))))).)-).(((((...))))).........------------ ( -18.80, z-score = -3.25, R) >droEre2.scaffold_4770 2977190 69 + 17746568 UAGUUAUGCAAUGCUGAACUAUUUGCUUGUUUAGUUUUGCAUAUC-UUUGCAUCUUGUGCAUCUGUUUUG------------ .((.(((((((.(((((((.........))))))).))))))).)-).(((((...))))).........------------ ( -18.80, z-score = -3.25, R) >droAna3.scaffold_13340 11411893 70 + 23697760 UAGUUAUGCAAUGCUGAAAUAUUUGCUUGUUUAGUUUUGCAUAUCUUUUGCAUCUUGUGCAUCUGCUUUG------------ .((.(((((((.((((((.((......)))))))).))))))).))..(((((...))))).........------------ ( -15.30, z-score = -1.24, R) >dp4.chr2 28306054 81 - 30794189 UAGUUAUGCAAUGCUGAAAUAUUUGCUUGUUUAGUUUUGCAUAUC-UUUUCUGCAUCUGCAUCUCUCGUGCAUCUGCUUUUG .((.(((((((.((((((.((......)))))))).))))))).)-).....(((..(((((.....)))))..)))..... ( -20.20, z-score = -2.85, R) >droPer1.super_6 3642863 81 - 6141320 UAGUUAUGCAAUGCUGAAAUAUUUGCUUGUUUAGUUUUGCAUAUC-UUUUCUGCAUCUGCAUCUCUCGUGCAUCUGCUUUUG .((.(((((((.((((((.((......)))))))).))))))).)-).....(((..(((((.....)))))..)))..... ( -20.20, z-score = -2.85, R) >droVir3.scaffold_13047 14169420 69 + 19223366 UAGUUAUGCAAUGCUGAAAUAUUUGCUUGUGUAUUUUUGCAUAUC-UUUGCAUCUUUUGCAUCUGCUGUG------------ .......(((((((.(((((((........))))))).))))...-..((((.....))))..)))....------------ ( -14.00, z-score = -0.73, R) >droMoj3.scaffold_6540 21372990 69 - 34148556 UAGUUAUGCAAUUCUGAAAUAUUUGCUUGUUUAGCUUUGCAUAUC-UUUGCAUCUCUUGCAUCUGCUGUG------------ (((((((((((..(((((.((......)))))))..)))))))..-..((((.....))))...))))..------------ ( -14.30, z-score = -1.17, R) >droGri2.scaffold_14906 3712058 69 + 14172833 UAGUUGUGCAAUGCUAAAAUAUUUGCUUGUUUAGUUUUGCAUAUC-UUUGCAUCUUUUGCAUCUGCUGUG------------ (((((((((((.((((((.((......)))))))).)))))))..-..((((.....))))...))))..------------ ( -15.80, z-score = -1.58, R) >consensus UAGUUAUGCAAUGCUGAAAUAUUUGCUUGUUUAGUUUUGCAUAUC_UUUGCAUCUUGUGCAUCUGUUUUG____________ ....(((((((.((((((.((......)))))))).))))))).....((((.....))))..................... (-11.82 = -12.29 + 0.47)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:04:55 2011