
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 6,348,541 – 6,348,635 |
| Length | 94 |
| Max. P | 0.819074 |

| Location | 6,348,541 – 6,348,635 |
|---|---|
| Length | 94 |
| Sequences | 9 |
| Columns | 115 |
| Reading direction | reverse |
| Mean pairwise identity | 62.55 |
| Shannon entropy | 0.68314 |
| G+C content | 0.41765 |
| Mean single sequence MFE | -19.88 |
| Consensus MFE | -7.15 |
| Energy contribution | -7.89 |
| Covariance contribution | 0.74 |
| Combinations/Pair | 1.23 |
| Mean z-score | -1.54 |
| Structure conservation index | 0.36 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.79 |
| SVM RNA-class probability | 0.819074 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 6348541 94 - 27905053 --UUUCAUUUGGCCAGAUUCCGUUGGCC-------AGCUCGGAAUGUACU-----AUAUG------UACA-CAUACACUAUAGGCCAACGCAUUUCCAUAAUUGCUAAAUUAUGC --......((((((..((((((..((..-------..))))))))((((.-----....)------))).-...........))))))........(((((((....))))))). ( -24.50, z-score = -2.44, R) >droSim1.chr3R 12485447 93 + 27517382 --UUUCGUUUGGCCAGAUUCCGUUGGCC-------AACUCGGAAUGUACU-----AUA--------CACCACAUACACUAUAGGCCAACGCAUUUCCAUAAUUGCUAAAUUAUGC --(((((.((((((((......))))))-------))..)))))((..((-----(((--------............)))))..)).........(((((((....))))))). ( -21.40, z-score = -2.03, R) >droSec1.super_0 5527514 93 + 21120651 --UUUCAUUUGGCCAGAUUCCGUUGGCC-------AACUCGGAAUGUACU-----AUA--------CACCACAUACACUAUAGGCCAACGCAUUUCCAUAAUUGCUAAAUUAUGC --......((((((((......))))))-------))...(((((((.((-----(((--------............)))))......)))))))(((((((....))))))). ( -20.20, z-score = -2.00, R) >droYak2.chr3R 10377148 113 - 28832112 --UUCCAUUUGGCCAGAUUCCGUUGGCCAACUCGGAAUGUACUAUAUACUACUGCAUAUAUGUGUAUACUGCAUACACUAUAGGCCAACGCAUUUCCAUAAUUGCUAAAUUAUGC --..................((((((((......(.(((((.(((((((............))))))).))))).)......))))))))......(((((((....))))))). ( -28.10, z-score = -2.00, R) >droEre2.scaffold_4770 2485679 98 + 17746568 --UUUCAUUUGGCCAGAUUCCGCUGACC--------AUGUACCUUAUACU-----AUAUGCUUUU-UAUAGUAUAUAGUAUA-GCCAGCGCAUUUCCAUAAUUGCUAAAUUAUGC --......((((((((......)))...--------........((((((-----(((((((...-...)))))))))))))-)))))........(((((((....))))))). ( -25.70, z-score = -3.61, R) >dp4.chr2 23675708 82 + 30794189 CCUCCGAUUUGGCCAGAUUCGGUUGGUC----------AUUCCAGCUCC-----------------CAUCUCAU------UGGGGCGACGCAUUUCCAUAAUUGCUAAAUUAUGC .....((..(((((((......))))))----------).))..((.((-----------------((......------))))))..........(((((((....))))))). ( -20.90, z-score = -0.61, R) >droPer1.super_0 10850309 81 + 11822988 CCUCCGAUUUGGCCAGAUUCGGUUGGUC----------AUU-CAGCUCC-----------------CAUCUCAU------UGGGGCGACGCAUUUCCAUAAUUGCUAAAUUAUGC .....((..(((((((......))))))----------).)-).((.((-----------------((......------))))))..........(((((((....))))))). ( -21.80, z-score = -1.12, R) >droWil1.scaffold_181108 134567 77 - 4707319 -----CAUUUGGUCAAAGACGACAGAGA----------GAAAGAGCAACUA---------------CAUUUC--------CACAUUGACGCAUUUCCAUAAUUGCUAAAUUAUGC -----....(((((((...(....).(.----------((((.........---------------..))))--------)...))))).))....(((((((....))))))). ( -7.70, z-score = 0.80, R) >droAna3.scaffold_13340 22945772 81 + 23697760 --------CCAGUCCCAUUCUAUUCCCAA-----UCCCAUUCGAGCAGU-----------------CAUUCUGUGCA----AUCCCGACGCAUUUCCAUAAUUGCUAAAUUAUGC --------.....................-----...(((...((((((-----------------......((((.----........)))).......)))))).....))). ( -8.62, z-score = -0.82, R) >consensus __UUUCAUUUGGCCAGAUUCCGUUGGCC________ACAUACAAUCUACU_____AUA________CAUCUCAUACA_UAUAGGCCAACGCAUUUCCAUAAUUGCUAAAUUAUGC ..........((((((......))))))....................................................................(((((((....))))))). ( -7.15 = -7.89 + 0.74)


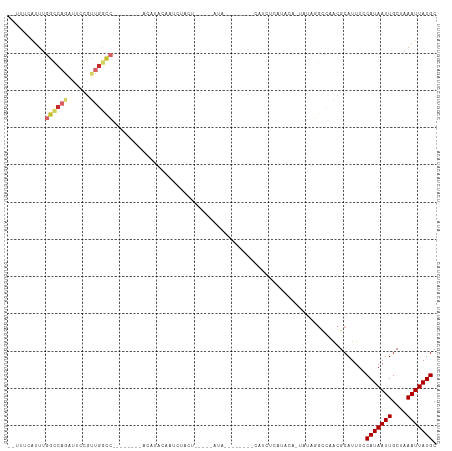
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:03:25 2011