
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 6,333,825 – 6,333,930 |
| Length | 105 |
| Max. P | 0.977600 |

| Location | 6,333,825 – 6,333,930 |
|---|---|
| Length | 105 |
| Sequences | 5 |
| Columns | 109 |
| Reading direction | forward |
| Mean pairwise identity | 91.11 |
| Shannon entropy | 0.14802 |
| G+C content | 0.39089 |
| Mean single sequence MFE | -20.78 |
| Consensus MFE | -19.08 |
| Energy contribution | -19.12 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.28 |
| Structure conservation index | 0.92 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.98 |
| SVM RNA-class probability | 0.977600 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
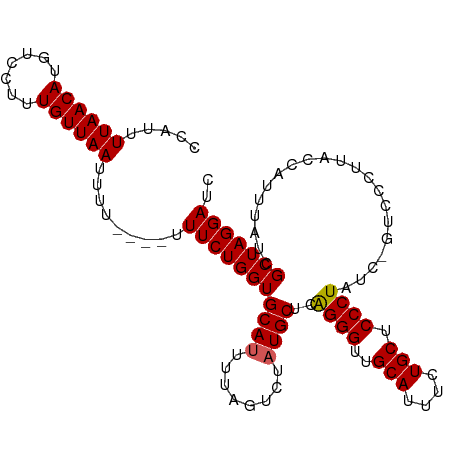
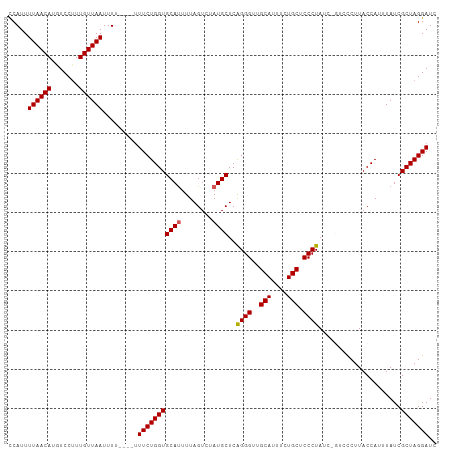
>dm3.chr3R 6333825 105 + 27905053 CCAUUUUAACAAGUCCUUUGUUAAUUUU----UUUCUGGUGCAUUUUAGUCUAUGCUCAGGGUUGCAUUUCUGCUCCCUAUCCGUCUCUUACCAUUUAUCGCUAGGAUC .....((((((((...))))))))....----.(((((((((((........))))..((((..(((....))).)))).....................))))))).. ( -20.90, z-score = -2.41, R) >droEre2.scaffold_4770 2471063 103 - 17746568 UCAUUUUAACAUGUCCUUUGUUAAUUU-----UUUCUGGUGCACUUUAGUCUAUGCUCAGGGUUGCAUUUCUGCGCCCUAUC-GUCCCUUACCAUUUAUCGCUAGGAUC .....((((((.......))))))...-----.((((((((((..........)))..(((((.(((....))))))))...-.................))))))).. ( -19.40, z-score = -1.36, R) >droYak2.chr3R 10362001 104 + 28832112 UCAUUUUAACAUGUCCUUUGUUAAUUUA----CUUCUGGUGCAUUUUAGUCUAUGCUCAGGGUUGCAUUUCUGCGCCCUAUC-GUCCCUUACCAUUUAUCGCUAGGAUC .....((((((.......))))))....----.(((((((((((........))))..(((((.(((....))))))))...-.................))))))).. ( -22.00, z-score = -2.21, R) >droSec1.super_0 5512915 105 - 21120651 CCAUUUUAACAUUUCCUUUGUUAAUUUUCUUUUUUCUGGUGCAUUUUAGUCUAUGCUCAGGGUUGCAUUUCUGCUCCCUAUC----CCUUACCAUUUAUCGCUAGGAUC .....((((((.......)))))).........(((((((((((........))))..((((..(((....))).))))...----..............))))))).. ( -20.80, z-score = -2.97, R) >droSim1.chr3R 12470778 105 - 27517382 CCAUUUUAACAUUUCCAUUGUUAAUUUUCUUUUUUCUGGUGCAUUUUAGUCUAUGCUCGGGGUUGCAUUUCUGCUCCCUAUC----CCUUACCAUUUAUCGCUAGGAUC .....((((((.......)))))).........(((((((((((........))))..((((..(((....))).))))...----..............))))))).. ( -20.80, z-score = -2.45, R) >consensus CCAUUUUAACAUGUCCUUUGUUAAUUUU____UUUCUGGUGCAUUUUAGUCUAUGCUCAGGGUUGCAUUUCUGCUCCCUAUC_GUCCCUUACCAUUUAUCGCUAGGAUC .....((((((.......)))))).........(((((((((((........))))..((((..(((....))).)))).....................))))))).. (-19.08 = -19.12 + 0.04)

| Location | 6,333,825 – 6,333,930 |
|---|---|
| Length | 105 |
| Sequences | 5 |
| Columns | 109 |
| Reading direction | reverse |
| Mean pairwise identity | 91.11 |
| Shannon entropy | 0.14802 |
| G+C content | 0.39089 |
| Mean single sequence MFE | -21.20 |
| Consensus MFE | -20.18 |
| Energy contribution | -20.22 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.08 |
| Structure conservation index | 0.95 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.94 |
| SVM RNA-class probability | 0.975951 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
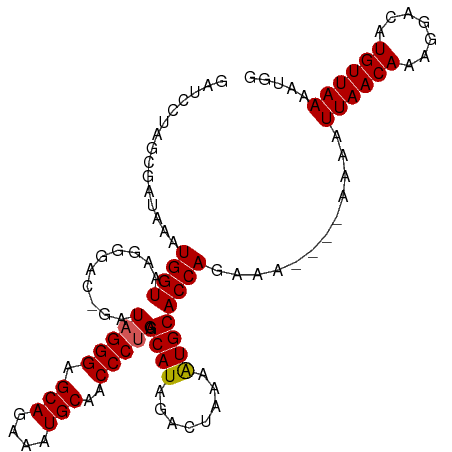
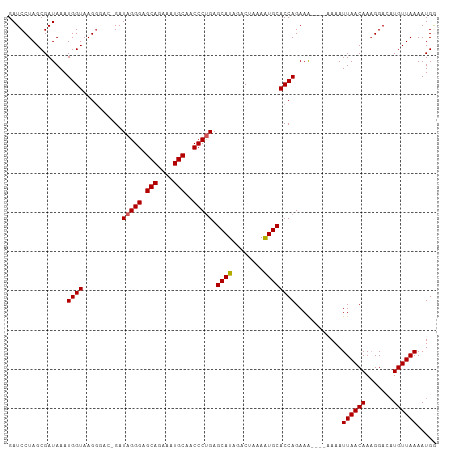
>dm3.chr3R 6333825 105 - 27905053 GAUCCUAGCGAUAAAUGGUAAGAGACGGAUAGGGAGCAGAAAUGCAACCCUGAGCAUAGACUAAAAUGCACCAGAAA----AAAAUUAACAAAGGACUUGUUAAAAUGG ...............((((..........(((((.(((....)))..))))).((((........))))))))....----....(((((((.....)))))))..... ( -22.10, z-score = -2.62, R) >droEre2.scaffold_4770 2471063 103 + 17746568 GAUCCUAGCGAUAAAUGGUAAGGGAC-GAUAGGGCGCAGAAAUGCAACCCUGAGCAUAGACUAAAGUGCACCAGAAA-----AAAUUAACAAAGGACAUGUUAAAAUGA ..((((.((........)).))))..-..(((((.(((....)))..))))).((((........))))........-----...((((((.......))))))..... ( -21.40, z-score = -1.77, R) >droYak2.chr3R 10362001 104 - 28832112 GAUCCUAGCGAUAAAUGGUAAGGGAC-GAUAGGGCGCAGAAAUGCAACCCUGAGCAUAGACUAAAAUGCACCAGAAG----UAAAUUAACAAAGGACAUGUUAAAAUGA ..((((.........((((.......-..(((((.(((....)))..))))).((((........))))))))...(----(......))..))))(((......))). ( -22.50, z-score = -2.06, R) >droSec1.super_0 5512915 105 + 21120651 GAUCCUAGCGAUAAAUGGUAAGG----GAUAGGGAGCAGAAAUGCAACCCUGAGCAUAGACUAAAAUGCACCAGAAAAAAGAAAAUUAACAAAGGAAAUGUUAAAAUGG ..((((.((........)).)))----).(((((.(((....)))..))))).((((........))))................((((((.......))))))..... ( -20.90, z-score = -2.41, R) >droSim1.chr3R 12470778 105 + 27517382 GAUCCUAGCGAUAAAUGGUAAGG----GAUAGGGAGCAGAAAUGCAACCCCGAGCAUAGACUAAAAUGCACCAGAAAAAAGAAAAUUAACAAUGGAAAUGUUAAAAUGG ..((((.((........)).)))----)...(((.(((....)))...)))..((((........))))................((((((.(....)))))))..... ( -19.10, z-score = -1.56, R) >consensus GAUCCUAGCGAUAAAUGGUAAGGGAC_GAUAGGGAGCAGAAAUGCAACCCUGAGCAUAGACUAAAAUGCACCAGAAA____AAAAUUAACAAAGGACAUGUUAAAAUGG ...............((((..........(((((.(((....)))..))))).((((........))))))))............((((((.......))))))..... (-20.18 = -20.22 + 0.04)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:03:22 2011