
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 6,228,804 – 6,228,919 |
| Length | 115 |
| Max. P | 0.869209 |

| Location | 6,228,804 – 6,228,894 |
|---|---|
| Length | 90 |
| Sequences | 7 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 76.20 |
| Shannon entropy | 0.44199 |
| G+C content | 0.37109 |
| Mean single sequence MFE | -21.59 |
| Consensus MFE | -11.90 |
| Energy contribution | -11.89 |
| Covariance contribution | -0.02 |
| Combinations/Pair | 1.46 |
| Mean z-score | -1.84 |
| Structure conservation index | 0.55 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.99 |
| SVM RNA-class probability | 0.869209 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 6228804 90 - 27905053 UUGAACUGCGGUACGAGUAAUUUACGUACCGAUUGAUUUAG----UUCCAUGCUUCACUG-AAGGAGCAUGUGGAAAAUAUACAUAUGUAAUCAA ........(((((((.........))))))).(((((....----(((((((((((....-..)))))))).))).....(((....)))))))) ( -24.20, z-score = -2.62, R) >droSim1.chr3R 12352944 90 + 27517382 UUGAACUGCGGUACGAGUAAUUUACGUACCGAUUUAAUUAG----UUCCAUGCUUCACUG-AAGGAGCAUGUGGAAAAUAUACAUAUGUAAUCAU ..(((((((((((((.........))))))).......)))----)))((((((((....-..)))))))).........(((....)))..... ( -23.71, z-score = -2.72, R) >droSec1.super_0 5406097 90 + 21120651 UUGAACUGCGGUACAAGUAAUUUACGUACCGAUUUAAUUAG----UUCAAUGCUUCACUG-AAGGAGCAUGUGGAAAAUAUACAUAUGUAGUAAA ....((((((((((...........)))))...........----(((.(((((((....-..)))))))...)))...........)))))... ( -21.50, z-score = -2.14, R) >droYak2.chr3R 10257356 90 - 28832112 UUGAACUGCGGUACAAGUAAUUAACGUACCGAUUUAAUUAG----UCUCAUGCUUCACAG-AAGAAGCAUGUGGAAAAUAUACACAUGUAAUCAA .(((....((((((...........)))))).....(((..----((.((((((((....-..)))))))).))..)))............))). ( -21.60, z-score = -2.61, R) >droEre2.scaffold_4770 2357311 90 + 17746568 UUGAACUGCGGUACAAGUAAUUAACGUACCGAUUUAAUUAG----UCUCGUGCUUCACUG-AAGGAGCAUGUUACAAAUAUACACAUGUAAUCAA .(((....((((((...........)))))).........(----(..((((((((....-..))))))))..))................))). ( -21.20, z-score = -1.98, R) >dp4.chr2 23555693 95 + 30794189 ACGAACUACGGUACUAACGAUGCCAGUACUAAUUUAGUUAGCGGCCGCCCUACUCCUCUGCAGGGUGUGCAUAAAAAAUAUACAACUUUAAUCAA ..((.....(((((....).)))).((((((((...)))))..((((((((.(......).)))))).))..........)))........)).. ( -19.60, z-score = -0.47, R) >droPer1.super_0 10723794 95 + 11822988 ACGAACUACGGUACUAACGAUGCCAGUACUAAUUUAGUUAGCGGCCGCCCUUCUCCUCUGCAGGGUGUGCAUAAAAAAUAUACAACUUUAAUCUA .((.((((.((((((.........))))))....))))...))((((((((.(......).)))))).))......................... ( -19.30, z-score = -0.36, R) >consensus UUGAACUGCGGUACAAGUAAUUUACGUACCGAUUUAAUUAG____UUCCAUGCUUCACUG_AAGGAGCAUGUGGAAAAUAUACAUAUGUAAUCAA ........((((((...........))))))..................(((((((.......)))))))......................... (-11.90 = -11.89 + -0.02)



| Location | 6,228,824 – 6,228,919 |
|---|---|
| Length | 95 |
| Sequences | 9 |
| Columns | 106 |
| Reading direction | reverse |
| Mean pairwise identity | 60.43 |
| Shannon entropy | 0.80707 |
| G+C content | 0.36174 |
| Mean single sequence MFE | -21.00 |
| Consensus MFE | -5.99 |
| Energy contribution | -6.96 |
| Covariance contribution | 0.97 |
| Combinations/Pair | 1.58 |
| Mean z-score | -1.33 |
| Structure conservation index | 0.29 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.29 |
| SVM RNA-class probability | 0.631453 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 6228824 95 - 27905053 AUUUAAUAUUAGUUAGCAA------UAUCUUUUGAACUGCGGUACGAGUAAUUUACGUACCGAUUGAUUUAGU-UCCAUGCUUCACUGAAGGAGCAUGUGGA---- ..((((.(((((((.(((.------..((....))..)))((((((.........)))))))))))))))))(-((((((((((......)))))))).)))---- ( -25.10, z-score = -2.55, R) >droSim1.chr3R 12352964 95 + 27517382 AUCUAAUAUUAGUUAGCAA------UAUCUUUUGAACUGCGGUACGAGUAAUUUACGUACCGAUUUAAUUAGU-UCCAUGCUUCACUGAAGGAGCAUGUGGA---- .((((..(((((((((((.------..((....))..)))((((((.........))))))....))))))))-..((((((((......))))))))))))---- ( -25.60, z-score = -2.69, R) >droSec1.super_0 5406117 95 + 21120651 AUCUAAUAUUAGUUAGCAA------UGUCUUUUGAACUGCGGUACAAGUAAUUUACGUACCGAUUUAAUUAGU-UCAAUGCUUCACUGAAGGAGCAUGUGGA---- .((((..(((((((((((.------..((....))..)))(((((...........)))))....))))))))-...(((((((......))))))).))))---- ( -22.00, z-score = -1.45, R) >droYak2.chr3R 10257376 95 - 28832112 CUCUAAUAUUAGUUAGCAA------UGUCUUUUGAACUGCGGUACAAGUAAUUAACGUACCGAUUUAAUUAGU-CUCAUGCUUCACAGAAGAAGCAUGUGGA---- .......(((((((((((.------..((....))..)))(((((...........)))))....))))))))-(.((((((((......)))))))).)..---- ( -24.60, z-score = -2.45, R) >droEre2.scaffold_4770 2357331 101 + 17746568 CUGUAAUAUUAGUUAGUUAGUUAGUUAUCUUUUGAACUGCGGUACAAGUAAUUAACGUACCGAUUUAAUUAGU-CUCGUGCUUCACUGAAGGAGCAUGUUAC---- ..((((.(((((((((.....(((((........)))))((((((...........))))))..)))))))))-..((((((((......))))))))))))---- ( -22.70, z-score = -1.10, R) >dp4.chr2 23555713 98 + 30794189 CUUUAAUAUUAGUUAGUAA--------CUCUACGAACUACGGUACUAACGAUGCCAGUACUAAUUUAGUUAGCGGCCGCCCUACUCCUCUGCAGGGUGUGCAUAAA ..(((((...((((((((.--------......(.....)(((((....).))))..))))))))..)))))..((((((((.(......).)))))).))..... ( -22.30, z-score = -0.08, R) >droPer1.super_0 10723814 98 + 11822988 CUUUAAUAUUAGUUAGUAA--------CUCUACGAACUACGGUACUAACGAUGCCAGUACUAAUUUAGUUAGCGGCCGCCCUUCUCCUCUGCAGGGUGUGCAUAAA ..(((((...((((((((.--------......(.....)(((((....).))))..))))))))..)))))..((((((((.(......).)))))).))..... ( -22.30, z-score = -0.26, R) >droWil1.scaffold_181089 12288922 84 + 12369635 CUUUAGUCUUAGUUAGUGG-------GUAUAAUUAAUUACA--------AAUUUACUUAC--AUCUAGUUAGCGGUU-CUCCGAAAUAUACAAGUCUAUUCA---- ...(((.(((.....((((-------(((.((((.......--------)))))))))))--..........(((..-..)))........))).)))....---- ( -12.00, z-score = -0.54, R) >droVir3.scaffold_13047 18301029 72 + 19223366 -------AUAAAUAAGUAA------AAGAAUAUGAA-UGUGGGCUUAGAAAUAUGUUGCC--ACAUAGCUAG--AUAAGCCUGUCCUAUG---------------- -------............------...........-.(..((((((....(((((....--))))).....--.))))))..)......---------------- ( -12.40, z-score = -0.82, R) >consensus CUUUAAUAUUAGUUAGUAA______UAUCUUUUGAACUGCGGUACAAGUAAUUUACGUACCGAUUUAAUUAGU_GCCAUGCUUCACUGAAGGAGCAUGUGGA____ ........................................(((((...........)))))...............((((((.(......).))))))........ ( -5.99 = -6.96 + 0.97)


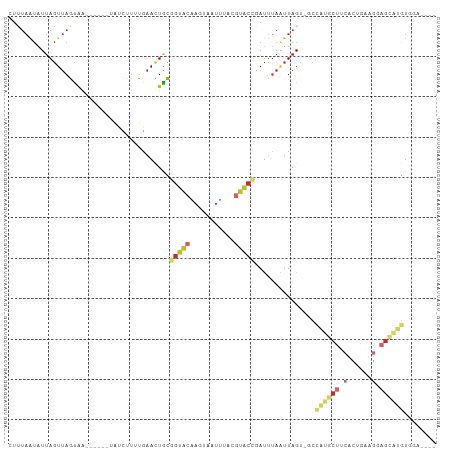
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:02:48 2011