
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 6,100,078 – 6,100,176 |
| Length | 98 |
| Max. P | 0.914525 |

| Location | 6,100,078 – 6,100,176 |
|---|---|
| Length | 98 |
| Sequences | 10 |
| Columns | 108 |
| Reading direction | reverse |
| Mean pairwise identity | 66.46 |
| Shannon entropy | 0.68295 |
| G+C content | 0.46023 |
| Mean single sequence MFE | -26.25 |
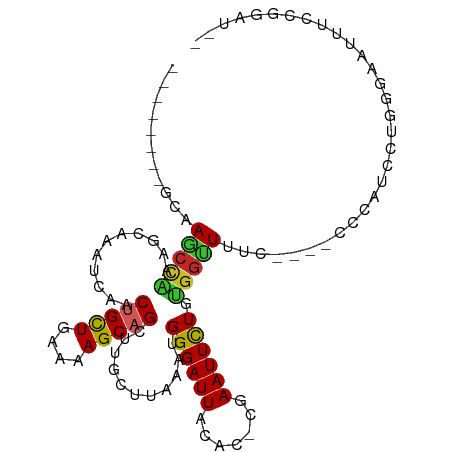
| Consensus MFE | -10.55 |
| Energy contribution | -10.14 |
| Covariance contribution | -0.41 |
| Combinations/Pair | 1.44 |
| Mean z-score | -1.53 |
| Structure conservation index | 0.40 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.24 |
| SVM RNA-class probability | 0.914525 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
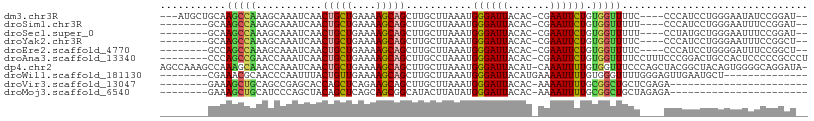
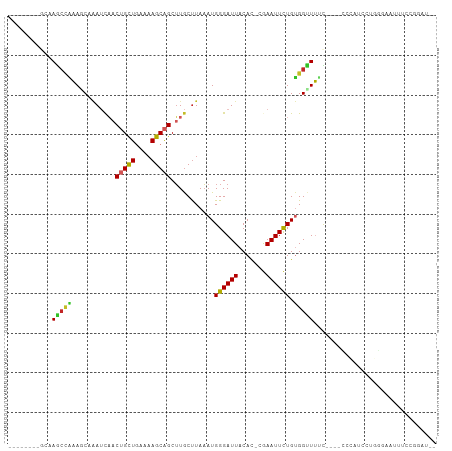
>dm3.chr3R 6100078 98 - 27905053 ---AUGCUGCAAGCCAAAGCAAAUCAACUGCUGAAAAGCAGCUUGCUUAAAUGGGAUUACAC-CGAAUUCUGUGGUUUUC----CCCAUCCUGGGAAUAUCCGGAU-- ---..........(((((((((.....(((((....))))).))))))...)))........-.....((((.(((.(((----((......))))).))))))).-- ( -26.80, z-score = -1.08, R) >droSim1.chr3R 12229483 93 + 27517382 --------GCAAGCCAAAGCAAAUCAACUGCUGAAAAGCAGCUUGCUUAAAUGGGAUUACAC-CGAAUUCUGUGGUUUUU----CCCAUCCUGGGAAUUUCCGGAU-- --------.....((.((((((.....(((((....))))).))))))..((((((((....-..))))))))((...((----(((.....)))))...))))..-- ( -27.30, z-score = -1.83, R) >droSec1.super_0 5288705 93 + 21120651 --------GCAAGCCAAAGCAAAUCAACUGCUGAAAAGCAGCUUGCUUAAAUGGGAUUACAC-CGAAUUCUGUGGUUUUU----CCUAUGCUGGGAAUUUCCGGAU-- --------((((((..............((((....))))))))))....((((((...(((-.(....).))).....)----))))).(((((....)))))..-- ( -25.13, z-score = -1.16, R) >droYak2.chr3R 10135675 93 - 28832112 --------GCAAGCCAAAGCAAAUCAACUGCUGAAAAGCAGCUUGCUUAAAUGGGAUUACAC-CGAAUUCUGUGGUUUUC----CCCAUCCUGGGAAUUUCCGGCU-- --------...((((.((((((.....(((((....))))).))))))..((((((((....-..))))))))((..(((----((......)))))...))))))-- ( -30.20, z-score = -2.48, R) >droEre2.scaffold_4770 2239564 93 + 17746568 --------GCCAGCCAAAGCAAAUCAACUGCUGAAAAGCAGCUUGCUUAAAUGGGAUUACAC-CGAAUUCUGUGGUUUUC----CCCAUCCUGGGGAUUUCCGGCU-- --------...((((.((((((.....(((((....))))).))))))..((((((((....-..))))))))((...((----(((.....)))))...))))))-- ( -33.70, z-score = -2.96, R) >droAna3.scaffold_13340 22713833 99 + 23697760 --------CCCAGCCGAACCAAAUCAACUGCUGAAAAGCAGCUUGCCUAAAUGGGAUUACAC-CGAAUUCUGUGGUUUUCCUUUCCCGGACUGCCACUCCCCCGCCCU --------....((..........((((((((....))))).))).......((((......-.(....).(((((..(((......)))..)))))))))..))... ( -24.60, z-score = -2.08, R) >dp4.chr2 23410666 106 + 30794189 AGCCAAAGCCAAAGCAAACCAAAUCAACUGCUGAAAAGCAGCUUGCUUAAAUGGGAUUACAU-CAAAUUUUGUGGUUUCCCAGCUACGGCUACAGUGGGGCAGGAUA- .(((..((((..(((.........((((((((....))))).)))......((((((((((.-.......)))))..))))))))..)))).......)))......- ( -31.40, z-score = -1.16, R) >droWil1.scaffold_181130 1650842 86 + 16660200 --------CGAAACGCAACCCAAUUUACUGUUGAAAAGCAGCUUGCUUAAAUGGGAUUACAUGAAAAUUUUGUGGGUUUUGGGAGUUGAAUGCU-------------- --------((((((.(..((((.(((((((((....)))))......))))))))...(((.........)))).)))))).............-------------- ( -16.60, z-score = 0.03, R) >droVir3.scaffold_13047 18177220 76 + 19223366 --------GAAAGCUGCAGCCGAGCACCAGCUCAGAAGCAGCUUGCUUAAAUGGGAUUACAC-AAAAUUUUGCGGCUGCUCGAGA----------------------- --------((.((((((((((((((....))))..((((.....))))....))((((....-..))))))))))))..))....----------------------- ( -22.00, z-score = -0.25, R) >droMoj3.scaffold_6540 29084193 76 + 34148556 --------GAAAGCUGCAUCCCAGCUACAGCUCAGCAGCGGCAUACUUAUAUGGGAUUACAC-AAAAUUUUGCGGCUGCUAGAGA----------------------- --------...(((((.....)))))....(((((((((.(((........((......)).-.......))).)))))).))).----------------------- ( -24.79, z-score = -2.29, R) >consensus ________GCAAGCCAAAGCAAAUCAACUGCUGAAAAGCAGCUUGCUUAAAUGGGAUUACAC_CGAAUUCUGUGGUUUUC____CCCAUCCUGGGAAUUUCCGGAU__ ...........(((((...........(((((....)))))...........((((((.......)))))).)))))............................... (-10.55 = -10.14 + -0.41)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:02:37 2011