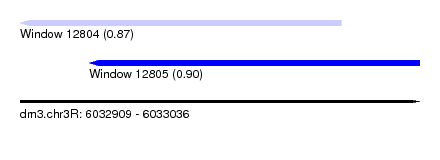

| Sequence ID | dm3.chr3R |
|---|---|
| Location | 6,032,909 – 6,033,036 |
| Length | 127 |
| Max. P | 0.904684 |

| Location | 6,032,909 – 6,033,011 |
|---|---|
| Length | 102 |
| Sequences | 4 |
| Columns | 103 |
| Reading direction | reverse |
| Mean pairwise identity | 76.96 |
| Shannon entropy | 0.34502 |
| G+C content | 0.43186 |
| Mean single sequence MFE | -22.32 |
| Consensus MFE | -14.06 |
| Energy contribution | -13.88 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.04 |
| Structure conservation index | 0.63 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.99 |
| SVM RNA-class probability | 0.869282 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
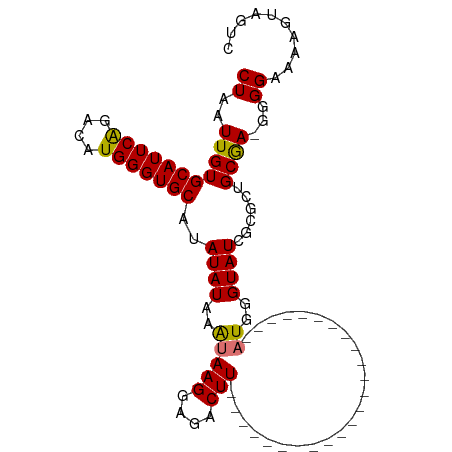
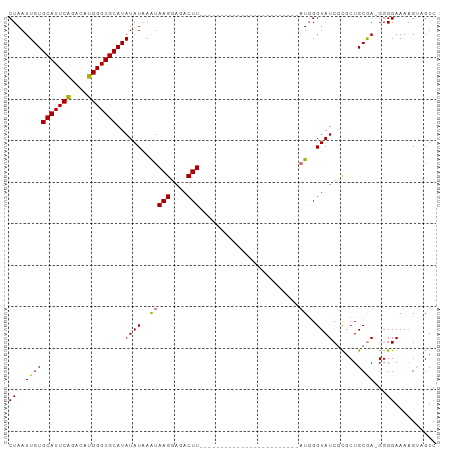
>dm3.chr3R 6032909 102 - 27905053 CUAAUUGUGCAUUCAGACAUGGGUGCAUAUAUUAGUAAGGAGACUUUUUUAUGUAUACACACACACACAGAUGGGUAUCGCUCUGCGA-GGGGAAAAGUAGUC (((((((((((((((....)))))))))).)))))...((..((((((((.....(((...((........)).)))((((...))))-..))))))))..)) ( -23.20, z-score = -0.99, R) >droSim1.chr3R 12169985 79 + 27517382 CUAAUUGUGCAUUCAGACAUGGGUGCAUAUAUAAAUAAGGAGACUU------------------------AUGGGUAUCGCGCUGCAAGGGGGAAAAGUCGUC .....((((((((((....))))))))))(((..(((((....)))------------------------))..)))((.(.(.....).).))......... ( -21.40, z-score = -2.80, R) >droSec1.super_0 5232368 78 + 21120651 CUAAUUGUGCAUUCAGACAUGGGUGCAUAUAUAAAUAAGGAGACUU------------------------AUGGGUAUCGCGCUGCGA-GGGGAAAAGUCGUC .....((((((((((....))))))))))(((..(((((....)))------------------------))..)))((((...))))-.............. ( -22.00, z-score = -2.74, R) >droYak2.chr3R 10077304 92 - 28832112 CUAAUUGUGCAUUCGGACAUGGGUGCAUAUAUAAAUAAGGAGACUUAUCUGCAUAUACACA---------UUUGGUAUC-UGCUGCGA-GGGGAAACGGAGUC .....((((((((((....))))))))))......((((....))))((((((.((((.(.---------...))))).-))).....-..(....))))... ( -22.70, z-score = -1.63, R) >consensus CUAAUUGUGCAUUCAGACAUGGGUGCAUAUAUAAAUAAGGAGACUU________________________AUGGGUAUCGCGCUGCGA_GGGGAAAAGUAGUC .....((((((((((....)))))))))).......(((....)))............................(((......)))................. (-14.06 = -13.88 + -0.19)

| Location | 6,032,931 – 6,033,036 |
|---|---|
| Length | 105 |
| Sequences | 5 |
| Columns | 105 |
| Reading direction | reverse |
| Mean pairwise identity | 78.06 |
| Shannon entropy | 0.36105 |
| G+C content | 0.40125 |
| Mean single sequence MFE | -25.76 |
| Consensus MFE | -18.88 |
| Energy contribution | -19.04 |
| Covariance contribution | 0.16 |
| Combinations/Pair | 1.15 |
| Mean z-score | -1.73 |
| Structure conservation index | 0.73 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.18 |
| SVM RNA-class probability | 0.904684 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
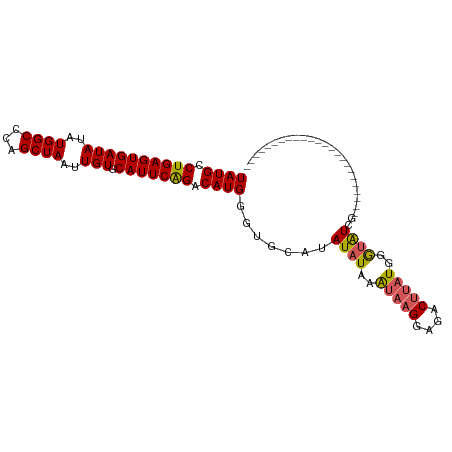
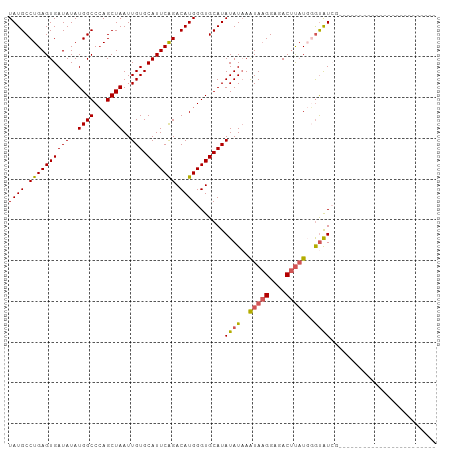
>dm3.chr3R 6032931 105 - 27905053 UAUGCCUGAGUGAUAUAUGGCCCAGCUAAUUGUGCAUUCAGACAUGGGUGCAUAUAUUAGUAAGGAGACUUUUUUAUGUAUACACACACACACAGAUGGGUAUCG .((((((..(((((((((((....((((((((((((((((....)))))))))).))))))(((....)))..)))))))).)))...(.....)..)))))).. ( -30.00, z-score = -1.87, R) >droSim1.chr3R 12170008 81 + 27517382 UAUGCCUGAGUGAUAUAUGGCCCAGCUAAUUGUGCAUUCAGACAUGGGUGCAUAUAUAAAUAAGGAGACUUAUGGGUAUCG------------------------ .((((((......((((((((...))))..((((((((((....)))))))))))))).(((((....)))))))))))..------------------------ ( -24.10, z-score = -1.83, R) >droSec1.super_0 5232390 81 + 21120651 UAUGCCUGAGUGAUAUAUGGCCCAGCUAAUUGUGCAUUCAGACAUGGGUGCAUAUAUAAAUAAGGAGACUUAUGGGUAUCG------------------------ .((((((......((((((((...))))..((((((((((....)))))))))))))).(((((....)))))))))))..------------------------ ( -24.10, z-score = -1.83, R) >droYak2.chr3R 10077326 95 - 28832112 UAUGCCUGAGUGAUAUAUGGCCAAGCUAAUUGUGCAUUCGGACAUGGGUGCAUAUAUAAAUAAGGAGACUUAUCUGCAUAUACACAUUUGGUAUC---------- .(((((.(((((.((((((...........((((((((((....)))))))))).....(((((....)))))...))))))))).)).))))).---------- ( -28.20, z-score = -2.22, R) >droEre2.scaffold_4770 2180990 80 + 17746568 UAUGCCUGAGUGAUAUAUGGCCCAGCUAAUUGUGCAUUCAGACAUGGGUGCAUAUAUAAAUACGGAUACACAUGGGUAUC------------------------- .(((((((.(((.....((((...))))..((((((((((....))))))))))..............))).))))))).------------------------- ( -22.40, z-score = -0.90, R) >consensus UAUGCCUGAGUGAUAUAUGGCCCAGCUAAUUGUGCAUUCAGACAUGGGUGCAUAUAUAAAUAAGGAGACUUAUGGGUAUCG________________________ ((((.((((((((((..((((...))))..))).))))))).)))).......((((..(((((....)))))..)))).......................... (-18.88 = -19.04 + 0.16)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:02:30 2011